Abstract
Osmotin is an important multifunctional protein related to plant stress responses and is classified into the thaumatin-like protein (TLP) family. Using genome-wide and phylogenetic approaches, we investigated osmotin origin and diversification across plant TLP evolution. Genomic and protein in silico analysis tools were also accessed and considered for the study conclusions. Phylogenetic analysis including a total of 722 sequences from 32 Viridiplantae species allowed the identification of an osmotin group that includes all previously characterized osmotins. Based on the phylogenetic tree results, it is evident that the osmotin group emerged from spermatophytes. Phylogenetic separation and gene expansion could be accounted for by an exclusive motif composition and organization that emerged and was maintained following tandem and block duplications as well as natural selection. The TLP family conserved residues and structures that were also identified in the sequences of the osmotin group, thus suggesting their maintenance for defense responses. The gene expression of Arabidopsis and rice putative osmotins reinforces its roles during stress response.









Similar content being viewed by others
Availability of data and materials
The data analyzed in this study are available on Phytozome v.12.0, Congenie, and the NCBI database. See Table S1 for accession numbers. Phytozome v.12.0 Database. http://www.phytozome.net/. Accessed 8 May 2017. Congenie Database. http://congenie.org/. Accessed 16 May 2017. NCBI Database. http://www.ncbi.nlm.nih.gov. Accessed 16 May 2017. The Arabidopsis Information Resource (TAIR10). http://www.arabidopsis.org/. Accessed 9 July 2017. The software used in the present study is available at the links below: Molecular Evolutionary Genetics Analysis (MEGA)—https://www.megasoftware.net/. Simple Modular Architecture Research Tool (SMART)—http://smart.embl-heidelberg.de/. ProtTest 3.4—http://darwin.uvigo.es/software/prottest2_server.html. Bayesian Evolutionary Analysis Sampling Trees (BEAST)—http://beast.community/. Figtree—http://tree.bio.ed.ac.uk/software/figtree/. Gene Structure Display Server (GSDS)—http://gsds.cbi.pku.edu.cn/. TargetP 1.1 Server—http://www.cbs.dtu.dk/services/TargetP/. WebLogo—https://weblogo.berkeley.edu/logo.cgi. WGmapping available in Plaza 3.0—http://bioinformatics.psb.ugent.be/plaza/. MCScanX software—http://chibba.pgml.uga.edu/mcscan2/. Swiss-model—https://swissmodel.expasy.org/.
Abbreviations
- TLP:
-
Thaumatin-like proteins
- OLPs:
-
Osmotin-like proteins
- PR-5:
-
Protein family 5
- AA:
-
Amino acids
- CDS:
-
Coding sequences
- WGD:
-
Whole-genome duplication
References
Abascal F, Zardoya R, Posada D (2005) ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21:2104–2105
Abdin MZ, Kiran U, Alam A (2011) Analysis of osmotin, a PR protein as metabolic modulator in plants. Bioinformation 5:336
Ahmed NU, Park JI, Jung HJ, Chung MY, Cho YG, Nou IS (2013) Characterization of thaumatin-like gene family and identification of Pectobacterium carotovorum subsp. carotovorum inducible genes in Brassica oleracea. Plant Breed Biotech 1:111–121
Campos MA, Ribeiro SG, Rigden DJ, Monte DC, Grossi de Sa MF (2002) Putative pathogenesis-related genes within Solanum nigrum var. Americanum genome: isolation of two genes coding for PR5-like proteins, phylogenetic and sequence analysis. Physiol Mol Plant Pathol 61:205–216
Cannon SB, Mitra A, Baumgarten A, Young ND, May G (2004) The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol 4:10
Cao J, Lv Y, Hou Z, Li X, Ding L (2016) Expansion and evolution of thaumatin-like protein (TLP) gene family in six plants. Plant Growth Regul 79:299–307
Capelli N, Diogon N, Greppin H, Simon P (1997) Isolation and characterization of a cDNA clone encoding an osmotin-like protein from Arabidopsis thaliana. Gene 191:51–56
Castillo RA, Herrera C, Ghislain M, Gebhardt C (2005) Organization of phenylalanine ammonia lyase (PAL), acidic PR-5 and osmotin-like (OSM) defence-response gene families in the potato genome. Mol Genet Genom 274:168–179
Chen WP, Chen PD, Liu DJ, Kynast R, Friebe B, Velazhahan R, Muthukrishnan S, Gill BS (1999) Development of wheat scab symptoms is delayed in transgenic wheat plants that constitutively express a rice thaumatin-like protein gene. Theor Appl Genet 99:755–760
Dong W, Vannozzi A, Chen F, Hu Y, Chen Z, Zhang L (2018) MORC domain definition and evolutionary analysis of the MORC gene family in green plants. Genome Biol Evol 10:1730–1744
Drummond AJ, Suchard MA, Xie D, Rambaut A (2012) Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol Biol Evol 29:1969–1973
Emanuelsson O, Nielsen H, Brunak S, Von Herijne G (2000) Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J Mol Biol 300:1005–1016
Evers D, Overney S, Simon P, Greppin H, Hausman JF (1999) Salt tolerance of Solanum tuberosum L.: overexpressing an heterologous osmotin-like protein. Biol Plant 42:105–112
Gao ZS, Weg WE, Schaart JG, Arkel G, Breiteneder H, Hoffmann-Sommergruber K, Gilissen LJ (2005) Genomic characterization and linkage mapping of the apple allergen genes Mal d 2 (thaumatin-like protein) and Mal d 4 (profilin). Theor Appl Genet 111:1087–1097
Hakim Ullah A, Hussain A, Shaban M, Khan AH, Alariqi M, Gul S, Jun Z, Lin S, Li J, Jin S, Munis MFH (2017) Osmotin: a plant defense tool against biotic and abiotic stresses. Plant Physiol Biochem 123:149–159
Hu B, Jin J, Guo AY, Zhang H, Luo J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31:1296–1297
Inschlag C, Hoffmann-Sommergruber K, O’Riordain G, Ahorn H, Ebner C, Scheiner O, Breiteneder H (1998) Biochemical characterization of Pru a 2, a 23-kD thaumatin-like protein representing a potential major allergen in cherry (Prunus avium). Int Arch Allergy Immunol 116:22–28
Jiang Y, Yang B, Harris NS, Deyholos MK (2007) Comparative proteomic analysis of NaCl stress-responsive proteins in Arabidopsis roots. J Exp Bot 58:3591–3607
Kim H, Mun JH, Byun BH, Hwang HJ, Kwon YM, Kim SG (2002) Molecular cloning and characterization of the gene encoding osmotin protein in Petunia hybrid. Plant Sci 162:745–752
Koonin EV (2006) The origin of introns and their role in eukaryogenesis: a compromise solution to the introns-early versus introns-late debate. Biol Direct 1:22
Kozlowski LP (2016) IPC–isoelectric point calculator. Biol Direct 11:55
Kumar V, Spencer ME (1992) Nucleotide sequence of an osmotin cDNA from the Nicotiana tabacum cv. white burley generated by the polymerase chain reaction. Plant Mol Biol 18:621–622
Kumar SA, Kumari PH, Kumar GS, Mohanalatha C, Kishor PK (2015) Osmotin: a plant sentinel and a possible agonist of mammalian adiponectin. Front Plant Sci 6:163
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Lehtonen MT, Takikawa Y, Rönnholm G, Akita M, Kalkkinen N, Ahola-Iivarinen E et al (2014) Protein secretome of moss plants (Physcomitrella patens) with emphasis on changes induced by a fungal elicitor. J Proteome Res 13:447–459
León IP, Montesano M (2017) Adaptation mechanisms in the evolution of moss defenses to microbes. Front Plant Sci 8:366
Letunic I, Bork P (2017) 20 years of the SMART protein domain annotation resource. Nucl Acids Res 46:D493–D496
Liu JJ, Sturrock R, Ekramoddoullah AKM (2010) The superfamily of thaumatin-like proteins: its origin, evolution, and expression towards biological function. Plant Cell Rep 29:419–436
Mani T, Sivakumar KC, Manjula S (2012) Expression and functional analysis of two osmotin (PR5) isoforms with differential antifungal activity from Piper colubrinum: prediction of structure–function relationship by bioinformatics approach. Mol Biotechnol 52:251–261
Medina J, Quatrano RS (1996) Characterization of a rice cDNA (Accession No. L76377) encoding an osmotin-like protein (PGR96-058). Plant Physiol 111:1354
Mohr PG, Cahill DM (2007) Suppression by ABA of salicylic acid and lignin accumulation and the expression of multiple genes, in Arabidopsis infected with Pseudomonas syringae pv. tomato. Funct Integr Genom 7:181–191
Mukherjee AK, Carp MJ, Zuchman R, Ziv T, Horwitz BA, Gepstein S (2010) Proteomics of the response of Arabidopsis thaliana to infection with Alternaria brassicicola. J Proteom 73:709–720
Onishi M, Tachi H, Kojima T, Shiraiwa M, Takahara H (2006) Molecular cloning and characterization of a novel salt-inducible gene encoding an acidic isoform of PR-5 protein in soybean. Plant Physiol Biochem 44:574–580
Petre B, Major I, Rouhier N, Duplessis S (2011) Genome-wide analysis of eukaryote thaumatin-like proteins (TLPs) with an emphasis on poplar. BMC Plant Biol 11:33
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarisation in Bayesian phylogenetics using Tracer 1.7. Syst Biol 67(5):901–904. https://doi.org/10.1093/sysbio/syy032
Salzman RA, Koiwa H, Ibeas JI, Pardo JM, Hasegawa PM, Bressan RA (2004) Inorganic cations mediate plant PR5 protein antifungal activity through fungal Mnn1-and Mnn4-regulated cell surface glycans. Mol Plant Microbe Interact 17:780–788
Shatters RG, Boykin LM, Lapointe SL, Hunter WB, Weathersbee AA III (2006) Phylogenetic and structural relationships of the PR5 gene family reveal an ancient multigene family conserved in plants and select animal taxa. J Mol Evol 63:12–29
Tachi H, Yamada KF, Kojima T, Shiraiwa M, Takahara H (2009) Molecular characterization of a novel soybean gene encoding a neutral PR-5 protein induced by high-salt stress. Plant Physiol Biochem 47:73–79
Vigers AJ, Roberts WK, Selitrennikoff CP (1991) A new family of plant antifungal proteins. Mol Plant Microbe Interact 4:315–323
Wang X, Zafian P, Choudhary M, Lawton M (1996) The PR5K receptor protein kinase from Arabidopsis thaliana is structurally related to a family of plant defense proteins. Proc Natl Acad Sci USA 93:2598–2602
Wang Y, Wang X, Tang H, Tan X, Ficklin SP, Feltus FA, Paterson AH (2011) Modes of gene duplication contribute differently to genetic novelty and redundancy, but show parallels across divergent angiosperms. PLoS One 6:e28150
Wang Y, Tang H, DeBarry JD, Xu T, Li J, Wang X, Lee T, Jin H, Marler B, Kissinger HGJC, Paterson AH (2012) MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucl Acids Res 40:e49–e49
Wang H, Devos KM, Bennetzen JL (2014) Recurrent loss of specific introns during angiosperm evolution. PLoS Genet 10:e1004843
Zong W, Zhong X, You J, Xiong L (2013) Genome-wide profiling of histone H3K4-tri-methylation and gene expression in rice under drought stress. Plant Mol Biol 81:175–188
Acknowledgements
We thank the two anonymous reviewers for their helpful comments on the manuscript.
Funding
This study was funded by Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), and INCT—MCTI/CNPq/CAPES/FAPs nº 16/2014, Ativos Biotecnológicos Aplicados a Seca e Pragas em Culturas Relevantes para o Agronegócio (INCT Biotec Seca-Pragas) [88887.136360/2017-00–465480/2014-4]. The funds provided by these funding institutions have been used to pay the stipends of GRF and LAO-B, respectively.
Author information
Authors and Affiliations
Contributions
Participated in the study design: GRF, LAO-B, ACT-Z, and MHB-Z. Performed the in silico analyses: GRF. Performed gene duplication analysis: FLG. Performed the phylogenetic analysis: GRF and ACT-Z. Wrote the paper: GRF. Revised the paper: LAO-B, ACT-Z, and MHB-Z. Supervised and coordinated the study: MHB-Z. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
GRF, ACT-Z, FLG, LAO-B and MHB-Z declares that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by S. Hohmann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
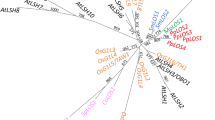
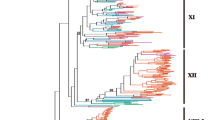
Phylogenetic tree comprising 722 thaumatin domains from 32 plant species with labels and posterior probability values. Branch lengths are proportional to phylogenetic distances. Pink, blue, yellow, and orange stars indicate the previously characterized permatin, osmotin, allergenic, and kinase proteins, respectively. Blue arrow indicates the osmotin group. The small blue triangle and dark gray circle next to the labels indicate the presence of a transmembrane domain and different domains beyond THN, respectively. (JPEG 7169 kb)
Fig. S2
Isoelectric points vs molecular weight for protein sequences (a) after osmotin group emergence, (b) for the osmotin group, and (c) before osmotin group emergence. (JPEG 124 kb)
Fig. S3
Architecture of conserved protein motifs in protein sequences localized (a) after osmotin group emergence, (b) for the osmotin group, and (c) before osmotin group emergence according to the ordering of the phylogenetic tree. Each motif is represented by a colored block. The lengths and positions of the blocks correspond to the lengths and positions of motifs in the individual protein sequences. The height of each block is proportional to its −log (p value), truncated at the height corresponding to a motif with a p value of 1e −10. The gene names and combined p values are shown on the left side of the figure. (JPEG 5367 kb)
Fig. S4
Motif localization in homologous modeling of Arabidopsis protein for (a) non-putative osmotin localized after osmotin group emergence, (b) osmotin, and (c) non-putative osmotin localized before osmotin group emergence. The green, purple, and red background in the sequence model-template alignment represents the stranded β-sheet, ⍺-helix, and motif sequences, respectively. The red circles in the 3D proteins represent the region of each motif. (d) Domain localization in TLP 3D protein. (JPEG 1113 kb)
File S1
Amino acid sequence alignment in FASTA format. (FAS 125 kb)
File S2
Phylogenetic tree in Newick format. (FAS 28 kb)
Table S1
Summary of all plant, gene, and protein sequences employed in this study. (XLSX 57 kb)
Table S2
TLP structural information. (XLSX 35 kb)
Table S3
Transposable elements that surround the Arabidopsis thaliana osmotin gene. Data accessed in TAIR10 (Wang et al. 2012). (XLSX 11 kb)
Table S4
Ontology annotation for putative Arabidopsis thaliana and Oryza sativa osmotin genes. (XLSX 11 kb)
Rights and permissions
About this article
Cite this article
Faillace, G.R., Turchetto-Zolet, A.C., Guzman, F.L. et al. Genome-wide analysis and evolution of plant thaumatin-like proteins: a focus on the origin and diversification of osmotins. Mol Genet Genomics 294, 1137–1157 (2019). https://doi.org/10.1007/s00438-019-01554-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-019-01554-y