Abstract
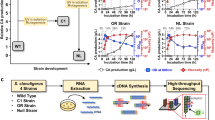
Mechanisms for high l-lactic acid production remain unclear in many bacteria. Lactobacillus rhamnosus SCT-10-10-60 was previously obtained from L. rhamnosus ATCC 11443 via mutagenesis and showed improved l-lactic acid production. In this study, the genomes of strains SCT-10-10-60 and ATCC 11443 were sequenced. Both genomes are a circular chromosome, 2.99 Mb in length with a GC content of approximately 46.8%. Eight split genes were identified in strain SCT-10-10-60, including two LytR family transcriptional regulators, two Rex redox-sensing transcriptional repressors, and four ABC transporters. In total, 60 significantly up-regulated genes (log2fold-change ≥ 2) and 39 significantly down-regulated genes (log2fold-change ≤ − 2) were identified by a transcriptome comparison between strains SCT-10-10-60 and ATCC 11443. KEGG pathway enrichment analysis revealed that “pyruvate metabolism” was significantly different (P < 0.05) between the two strains. The split genes and the differentially expressed genes involved in the “pyruvate metabolism” pathway are probably responsible for the increased l-lactic acid production by SCT-10-10-60. The genome and transcriptome sequencing information and comparison of SCT-10-10-60 with ATCC 11443 provide insights into the anabolism of l-lactic acid and a reference for improving l-lactic acid production using genetic engineering.



Similar content being viewed by others
References
Anders S, Huber W (2010) Differential expression analysis for sequence count data. Genome Biol 11:R106
Anders S, Pyl PT, Huber W (2015) HTSeq-a python framework to work with high-throughput sequencing data. Bioinformatics 31:166–169
Bao T, Zhang X, Zhao X, Rao Z, Yang T, Yang S (2015) Regulation of the NADH pool and NADH/NADPH ratio redistributes acetoin and 2,3-butanediol proportion in Bacillus subtilis. Biotechnol J 10:1298–1306
Berlin K, Koren S, Chin CS, Drake JP, Landolin JM, Phillippy AM (2015) Assembling large genomes with single-molecule sequencing and locality-sensitive hashing. Nat Biotechnol 33:623–630
Besemer J, Lomsadze A, Borodovsky M (2001) GeneMarkS: a self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions. Nucleic Acids Res 29:2607–2618
Bolotin A, Wincker P, Mauger S, Jaillon O, Malarme K, Weissenbach J, Ehrlich SD, Sorokin A (2001) The complete genome sequence of lactic acid bacterium Lactococcus lactis ssp lactis IL1403. Genome Res 11:731–753
Chatfield CH, Koo H, Jr QR (2005) The putative autolysin regulator LytR in Streptococcus mutans plays a role in cell division and is growth-phase regulated. Microbiology 151:625–631
Chiaromonte F, Yap VB, Miller W (2001) Scoring pairwise genomic sequence alignments. Pac Symp Biocomput 7:115–126
Cui X, Lu Z, Wang S, Jing-Yan Wang J, Gao X (2016) CMsearch: simultaneous exploration of protein sequence space and structure space improves not only protein homology detection but also protein structure prediction. Bioinformatics 32:i332–i340
Djukić-Vuković AP, Jokić BM, Kocić-Tanackov SD, Pejin JD, Mojović LV (2015) Mg-modified zeolite as a carrier for Lactobacillus rhamnosus in l(+)-lactic acid production on distillery wastewater. J Taiwan Inst Chem E 59:262–266
Douillard FP, de Vos WM (2014) Functional genomics of lactic acid bacteria: from food to health. Microb Cell Fact 13:S8
Douillard FP, Ribbera A, Kant R, Pietilä TE, Järvinen HM, Messing M, Randazzo CL, Paulin L, Laine P, Ritari J, Caggia C, Lähteinen T, Brouns SJ, Satokari R, von Ossowski I, Reunanen J, Palva A, de Vosm WM (2013) Comparative genomic and functional analysis of 100 Lactobacillus rhamnosus strains and their comparison with strain GG. PLoS Genet 9:e1003683
Eid J, Fehr A, Gray J, Luong K, Lyle J, Otto G, Peluso P, Rank D, Baybayan P, Bettman B, Bibillo A, Bjornson K, Chaudhuri B, Christians F, Cicero R, Clark S, Dalal R, deWinter A, Dixon J, Foquet M, Gaertner A, Hardenbol P, Heiner C, Hester K, Holden D, Kearns G, Kong X, Kuse R, Lacroix Y, Lin S, Lundquist P, Ma C, Marks P, Maxham M, Murphy D, Park I, Pham T, Phillips M, Roy J, Sebra R, Shen G, Sorenson J, Tomaney A, Travers K, Trulson M, Vieceli J, Wegener J, Wu D, Yang A, Zaccarin D, Zhao P, Zhong F, Korlach J, Turner S (2009) Real-time DNA sequencing from single polymerase molecules. Science 323:133–138
Even S, Lindley ND, Cocaign-Bousquet M (2001) Molecular physiology of sugar catabolism in Lactococcus lactis IL1403. J Bacteriol 183:3817–3824
Gao C, Ma C, Xu P (2011) Biotechnological routes based on lactic acid production from biomass. Biotechnol Adv 29:930–939
Gardner PP, Daub J, Tate JG, Nawrocki EP, Kolbe DL, Lindgreen S, Wilkinson AC, Finn RD, Griffiths-Jones S, Eddy SR, Bateman A (2009) Rfam: updates to the RNA families database. Nucleic Acids Res 37:D136–D140
Harris RS (2007) Improved pairwise alignment of genomic DNA. Dissertations & Theses - Gradworks
Hujanen M, Linko S, Linko YY, Leisola M (2001) Optimisation of media and cultivation conditions for L(+)(S)-lactic acid production by Lactobacillus casei NRRL B-441. Appl Microbiol Biotechnol 56:126–130
John RP, Nampoothiri KM, Pandey A (2006) Solid-state fermentation for l-lactic acid production from agro wastes using Lactobacillus delbrueckii. Proc Biochem 41:759–763
Juturu V, Wu JC (2015) Microbial production of lactic acid: the latest development. Crit Rev Biotechnol 11:1–11
Kadam SR, Patil SS, Bastawde KB, Khire JM, Gokhale DV (2006) Strain improvement of Lactobacillus delbrueckii NCIM 2365 for lactic acid production. Proc Biochem 41:120–126
Kankainen M, Paulin L, Tynkkynen S, von Ossowski I, Reunanen J, Partanen P, Satokari R, Vesterlund S, Hendrickx AP, Lebeer S, De Keersmaecker SC, Vanderleyden J, Hämäläinen T, Laukkanen S, Salovuori N, Ritari J, Alatalo E, Korpela R, Mattila-Sandholm T, Lassig A, Hatakka K, Kinnunen KT, Karjalainen H, Saxelin M, Laakso K, Surakka A, Palva A, Salusjärvi T, Auvinen P, de Vos WM (2009) Comparative genomic analysis of Lactobacillus rhamnosus GG reveals pili containing a human-mucus binding protein. Proc Natl Acad Sci USA 106:17193–17198
Klotz S, Kaufmann N, Kuenz A, Prüße U (2016) Biotechnological production of enantiomerically pure d-lactic acid. Appl Microbiol Biotechnol 100:9423–9437
Kurtz S, Phillippy A, Delcher AL, Smoot M, Shumway M, Antonescu C, Salzberg SL (2004) Versatile and open software for comparing large genomes. Genome Biol 5:R12
Lagesen K, Hallin P, Rødland EA, Stærfeldt H, Rognes T, Ussery DW (2007) RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res 35:3100–3108
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat methods 9:357–359
Li Z, Lu J, Zhao L, Xiao K, Tan T (2010) Improvement of l-lactic acid production under glucose feedback controlled culture by Lactobacillus rhamnosus. Appl Biochem Biotechnol 162:1762–1767
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25:955–964
Makarova K, Slesarev A, Wolf Y, Sorokin A, Mirkin B, Koonin E, Pavlov A, Pavlova N, Karamychev V, Polouchine N, Shakhova V, Grigoriev I, Lou Y, Rohksar D, Lucas S, Huang K, Goodstein DM, Hawkins T, Plengvidhya V, Welker D, Hughes J, Goh Y, Benson A, Baldwin K, Lee JH, Díaz-Muñiz I, Dosti B, Smeianov V, Wechter W, Barabote R, Lorca G, Altermann E, Barrangou R, Ganesan B, Xie Y, Rawsthorne H, Tamir D, Parker C, Breidt F, Broadbent J, Hutkins R, O’Sullivan D, Steele J, Unlu G, Saier M, Klaenhammer T, Richardson P, Kozyavkin S, Weimer B, Mills D (2006) Comparative genomics of the lactic acid bacteria. Proc Natl Acad Sci USA 103:15611–15616
Mao X, Cai T, Olyarchuk JG, Wei L (2005) Automated genome annotation and pathway identification using the KEGG Orthology (KO) as a controlled vocabulary. Bioinformatics 21:3787–3793
Melchiorsen CR, Jokumsen KV, Villadsen J, Israelsen H, Arnau J (2002) The level of pyruvate-formate lyase controls the shift from homolactic to mixed-acid product formation in Lactococcus lactis. Appl Microbiol Biotechnol 58:338–344
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-SEq. Nat Methods 5:621–628
Narayanan N, Roychoudhury PK, Srivastava A (2004) l-Lactic acid fermentation and its product polymerization. Electron J Biotechn 7:167–179
Naveena BJ, Altaf M, Bhadriah K, Reddy G (2005) Selection of medium components by Placket Burman design for the production of l(+)-lactic acid by Lactobacillus amylophilus GV6 in SSF using wheat bran. Bioresour Technol 96:485–490
Nawrocki EP, Kolbe DL (2009) Eddy SR: Infernal 1.0: inference of RNA alignments. Bioinformatics 25:1335–1337
Ohkouchi Y, Inoue Y (2006) Direct production of l(+)-lactic acid from starch and food wastes using Lactobacillus manihotivorans LMG18011. Bioresour Technol 97:1554–1562
Ravcheev DA, Li X, Latif H, Zengler K, Leyn SA, Korostelev YD, Kazakov AE, Novichkov PS, Osterman AL, Rodionov DA (2012) Transcriptional regulation of central carbon and energy metabolism in bacteria by redox-responsive repressor Rex. J Bacteriol 194:1145–1157
Reunanen J, von Ossowski I, Hendrickx APA, Palva A, de Vos WM (2012) Characterization of the SpaCBA pilus fibers in the probiotic Lactobacillus rhamnosus GG. Appl Environ Microbiol 78:2337–2344
Rojan PJ, Nampoothiri KM, Nair AS, Pandey A (2005) l(+)-Lactic acid production using Lactobacillus casei in solid-state fermentation. Biotechnol Lett 27:1685–1688
Senedese ALC, Maciel Filho R, Maciel MR (2015) l-Lactic acid production by Lactobacillus rhamnosus ATCC 10863. Sci World J 2015:501029
Sergey K, Adam MP (2015) One chromosome, one contig: complete microbial genomes from long-read sequencing and assembly. Curr Opin Microbiol 23:110–120
Stefanovic E, Fitzgerald G, Mcauliffe O (2017) Advances in the genomics and metabolomics of dairy lactobacilli: a review. Food Microbiol 61:33–49
Sun L, Sun F, Huang Y, Lu F, Huang R (2009) Optimization of Fermentation conditions in producing l-lactic acid by cassava starch. China Brewing 7:33–37
Sun L, Li J, Sun F, Huang Y, Guo L, Huang R (2013) Breeding of a Lactobacillus rhamnosus strain with the high yield fermentation of l-lactic acid. Guangxi Sciences 20:152–157
Sun Z, Harris HMB, McCann A, Guo C, Argimón S, Zhang W, Yang X, Jeffery IB, Cooney JC, Kagawa TF, Liu W, Song Y, Salvetti E, Wrobel A, Rasinkangas P, Parkhill J, Rea MC, O’Sullivan O, Ritari J, Douillard FP, PaulRoss R, Yang R, Briner AE, Felis GE, de Vos WM, Barrangou R, Klaenhammer TR, Caufield PW, Cui Y, Zhang H, O’Toole PW (2015) Expanding the biotechnology potential of lactobacilli through comparative genomics of 213 strains and associated genera. Nat Commun 6:8322
Tango MS, Ghaly AE (2002) A continuous lactic acid production system using an immobilized packed bed of Lactobacillus helveticus. Appl Microbiol Biotechnol 58:712–720
von Ossowski I, Reunanen J, Satokari R, Vesterlund S, Kankainen M, Huhtinen H, Tynkkynen S, Salminen S, de Vos WM, Palva A (2010) Mucosal adhesion properties of the probiotic Lactobacillus rhamnosus GG SpaCBA and SpaFED pilin subunits. Appl Environ Microbiol 76:2049–2057
Wang Y, Li Y, Pei X, Yu L, Feng Y (2007) Genome-shuffling improved acid tolerance and l-lactic acid volumetric productivity in Lactobacillus rhamnosus. J Biotechnol 129:510–515
Wang L, Zhao B, Liu B, Yang C, Yu B, Li Q, Ma C, Xu P, Ma Y (2010) Efficient production of l-lactic acid from cassava powder by Lactobacillus rhamnosus. Bioresour Technol 101:7895–7901
Wilkens S (2015) Structure and mechanism of ABC transporters. F1000 prime Reports 7:14
Yang M, Xu L, Liu Y, Yang P (2015) RNA-Seq uncovers SNPs and alternative splicing events in Asia lotus (Nelumbo nucifera). PLoS One 10:e0125702
Yu L, Pei X, Lei T, Wang Y, Feng Y (2008) Genome shuffling enhanced l-lactic acid production by improving glucose tolerance of Lactobacillus rhamnosus. J Biotechnol 134:154–159
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This research was supported jointly by the National High Technology Research and Development Program (“863” Program) of China (No. 2013AA050701) and Basic Science and Research Funding Program of Guangxi Academy of Science (No. 12YJ25SW06).
Conflict of interest
Liang Sun declares that he/she has no conflict of interest. Zhilong Lu declares that he/she has no conflict of interest. Jianxiu Li declares that he/she has no conflict of interest. Feifei Sun declares that he/she has no conflict of interest. Ribo Huang declares that he/she has no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by S. Hohmann.
Electronic supplementary material
Below is the link to the electronic supplementary material.


Rights and permissions
About this article
Cite this article
Sun, L., Lu, Z., Li, J. et al. Comparative genomics and transcriptome analysis of Lactobacillus rhamnosus ATCC 11443 and the mutant strain SCT-10-10-60 with enhanced l-lactic acid production capacity. Mol Genet Genomics 293, 265–276 (2018). https://doi.org/10.1007/s00438-017-1379-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-017-1379-0