Abstract
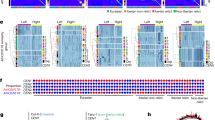
Despite the conserved roles and conserved protein machineries of centromeres, their nucleotide sequences can be highly diverse even among related species. The diversity reflects rapid evolution, but the underlying mechanism is largely unknown. One approach to monitor rapid evolution is examination of intra-specific variation. Here we report variant centromeric satellites of Arabidopsis thaliana found through survey of 103 natural accessions (ecotypes). Among them, a cluster of variant centromeric satellites was detected in one ecotype, Cape Verde Islands (Cvi). Recombinant inbred mapping revealed that the variant satellites are distributed in centromeric region of the chromosome 5 (CEN5) of this ecotype. This apparently recent variant accumulation is associated with large deletion of a pericentromeric region and the expansion of satellite region. The variant satellite was bound to HTR12 (centromeric variant histone H3), although expansion of the satellite was not associated with comparable increase in the HTR12 binding. The results suggest that variant satellites with centromere function can rapidly accumulate in one centromere, supporting the model that the satellite repeats in the array are homogenized by occasional unequal crossing-over, which has a potential to generate an expansion of local sequence variants within a centromere cluster.



Similar content being viewed by others
References
Alonso-Blanco C, Peeters AJ, Koornneef M, Lister C, Dean C, van den Bosch N, Pot L, Kuiper MT (1998) Development of an AFLP based linkage map of Ler, Col and Cvi Arabidopsis thaliana ecotypes and construction of a Ler/Cvi recombinant inbred line population. Plant J 14:259–271
Amor D, Bentley K, Ryan J, Perry J, Wong L, Slater H, Choo K (2004) Human centromere repositioning “in progress”. Proc Natl Acad Sci USA 101:6542–6547
Choo KHA (2001) Domain organization at the centromere and neocentromere. Dev Cell 1:165–177
Copenhaver GP, Nickel K, Kuromori T, Benito M, Kaul S, Lin X, Bevan M, Murphy G, Harris B, Parnell LD, McCombie WR, Martienssen RA, Marra M, Preuss D (1999) Genetic definition and sequence analysis of Arabidopsis centromeres. Science 286:2468–2474
Fransz P, Armstrong S, Alonso-Blanco C, Fischer TC, Torres-Ruiz RA, Jones G (1998) Cytogenetics for the model system Arabidopsis thaliana. Plant J 13:867–876
Gray KM, White JW, Costanzi C, Gillespie D, Schroeder WT, Calabretta B, Saunders GF (1985) Recent amplification of an alpha satellite DNA in humans. Nucleic Acids Res 13:521–535
Hall SE, Kettler G, Preuss D (2003) Centromere satellites from Arabidopsis populations: maintenance of conserved and variable domains. Genome Res 13:195–205
Haupt W, Fischer TC, Winderl S, Fransz P, Torres-Ruiz RA (2001) The centromere1 (CEN1) region of Arabidopsis thaliana: architecture and functional impact of chromatin. Plant J 27:285–296
Henikoff S, Ahmad K, Malik HS (2001) The centromere paradox stable inheritance with rapidly evolving DNA. Science 293:1098–1102
Heslop-Harrison JS, Murata M, Ogura Y, Schwarzacher T, Motoyoshi F (1999) Polymorphisms and genomic organization of repetitive DNA from centromeric regions of Arabidopsis chromosomes. Plant Cell 11:31–42
Hosouchi T, Kumekawa N, Tsuruoka H, Kotani H (2002) Physical map-based sizes of the centromeric regions of Arabidopsis thaliana chromosomes 1, 2, and 3. DNA Res 9:117–121
Jackson SA, Wang ML, Goodman HM, Jiang J (1998) Application of fiber-FISH in physical mapping of Arabidopsis thaliana. Genome 41:566–572
Jasencakova Z, Meister A, Schubert I (2001) Chromatin organization and its relation to replication and histone acetylation during the cell cycle in barley. Chromosoma 110:83–92
Jeddeloh JA, Stokes TL, Richards EJ (1999) Maintenance of genomic methylation requires a SWI2/SNF2-like protein. Nat Genet 22:94–97
Jiang J, Birchler JA, Parrott WA, Dawe RK (2003) A molecular view of plant centromres. Trends Plant Sci 8:570–575
Johnson L, Cao X, Jacobsen S (2002) Interplay between two epigenetic marks DNA methylation and histone H3 lysine 9 methylation. Curr Biol 12:1360–1367
Karpen G, Allshire R (1997) The case for epigenetic effects on centromere identity and function. Trends Genet 13:486–496
Kawabe A, Nasuda S (2006) Polymorphic chromosomal specificity of centromeric satellite families in Arabidopsis halleri spp. gemmifera. Genetica 126:335–342
Kumekawa N, Hosouchi T, Tsuruoka H, Kotani H (2000) The size and sequence organization of the centromeric region of arabidopsis thaliana chromosome 5. DNA Res 7:315–321
Kumekawa N, Hosouchi T, Tsuruoka H, Kotani H (2001) The size and sequence organization of the centromeric region of arabidopsis thaliana chromosome 4. DNA Res 8:285–290
Miura A, Kato M, Watanabe K, Kawabe A, Kotani H, Kakutani T (2004) Genomic localization of endogenous mobile CACTA family transposons in natural variants of Arabidopsis thaliana. Mol Genet Genomics 270:524–532
Nagaki K, Talbert P, Zhong CX, Henikoff S, Jiang J (2003) Choromatin immunoprecipitation reveales that the 180-bp satellite repeat is the key functional DNA element of Arabidopsis thaliana centromeres. Genetics 163:759–770
Nagaki K, Cheng Z, Ouyang S, Talbert PB, Kim M, Jones KM, Henikoff S, Buell CR, Jiang J (2004) Sequencing of a rice centromere uncovers active genes. Nat Genet 36:138–145
Nordborg M, Hy TT, Ishino Y, Jhaveri J, Toomajian C, Zheng H, Bakker E, Calabrese P, Gladstone J, Goyal R, Jakobsson M, Kim S, Morozov Y, Padhukasakasram B, Plagnol V, Rosenberg NA, Shah C, Wall JD, Wang J, Zhao K, Kalbfleisch T, Schulz V, Kreitman M, Bergelson J (2005) The pattern of polymorphism in Arabidopsis thaliana. PLos Biol 3:e196. Epub 2005 May 24
Round EK, Flowers SK, Richards EJ (1997) Arabidopsis thaliana centromere regions: genetic map positions and repetitive DNA structure. Genome Res 7:1045–1053
du Sart D, Cancilla MR, Earle E, Mao JI, Saffery R, Tainton KM, Kalitsis P, Martyn J, Barry AE, Choo KH (1997) A functional neo-centromere formed through activation of a latent human centromere and consisting of non-alpha-satellite DNA. Nat Genet 16:144–153
Smith GP (1976) Evolution of repeated DNA sequences by unequal crossover. Science 191:528–535
Talbert PB, Masuelli R, Tyagi AP, Comai L, Henikoff S (2002) Centromeric localization and adaptive evolution of an Arabidopsis histone H3 variant. Plant Cell 14:1053–1066
Vongs A, Kakutani T, Martienssen RA, Richards EJ (1993) Arabidopsis thaliana DNA methylation mutants. Science 260:1926–1928
Warburton PE (2004) Chromosomal dynamics of human neocentromere formation. Chromosome Res 12:617–626
Willard HF (1991) Evolution of alpha satellite. Curr Opin Genet Dev 1:509–514
Wolfe J, Darling SM, Erickson RP, Craig IW, Buckle VJ, Rigby PW, Willard HF, Goodfellow PN (1985) Isolation and characterization of an alphoid centromeric repeat family from the human Y chromosome. J Mol Biol 182:477–485
Acknowledgments
We thank Akiko Terui for technical assistance. Special thanks to Paul Talbert and Steve Henikoff for the anti-HTR12 antibody, Ales Pecinka and Ingo Schubert for the BAC clone information for the chromosomal painting, Federico Tessadori and Paul Fransz for technical advice, and Eric Richards for critical comments on the manuscript. We acknowledge Arabidopsis Biological Stock Center at Ohio State University for the seed stocks. Supported by Grant-in-Aid for Creative Scientific Research 14GS0321.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Aguilera.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Ito, H., Miura, A., Takashima, K. et al. Ecotype-specific and chromosome-specific expansion of variant centromeric satellites in Arabidopsis thaliana . Mol Genet Genomics 277, 23–30 (2007). https://doi.org/10.1007/s00438-006-0172-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-006-0172-2