Abstract
The rapid emergence of drug resistance against the mainstream antimalarial drugs has increased the need for development of novel drugs. Recent approaches have embarked on the repurposing of existing drugs to induce cell death via programmed cell death pathways. However, little is known about the ER stress response and programmed cell death pathways of the malaria parasite. In this study, we treated ex vivo Plasmodium berghei cultures with tunicamycin, 5-fluorouracil, and chloroquine as known stress inducer drugs to probe the transcriptional changes of autophagy and apoptosis-related genes (PbATG5, PbATG8, PbATG12, and PbMCA2). Treatments with 5-fluorouracil and chloroquine resulted in the upregulation of all analyzed markers, yet the levels of PbATG5 and PbATG12 were dramatically higher in chloroquine-treated ex vivo cultures. In contrast, tunicamycin treatment resulted in the downregulation of both PbATG8 and PbATG12, and upregulation of PbMCA2. Our results indicate that the malaria parasite responds to various ER stressors by inducing autophagy- and/or apoptosis-like pathways.
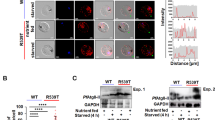
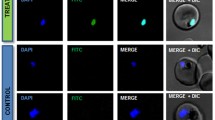
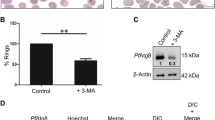
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Malaria is a vector-borne infectious disease that causes almost a quarter billion cases and over half a million deaths each year (WMR 2023). The disease is caused by Plasmodium species protozoan parasites and spread by Anopheline mosquitoes. One of the main obstacles in the fight against malaria is the increasing drug resistance. Repurposing of existing drugs is a useful strategy to meet the urgent need for new drugs (Laudisi et al. 2019). Studies show that using severe and sustained ER stress to induce programmed cell death (PCD) pathways is an efficient method against cancer and parasitic protozoan infections (Peng et al. 2021; Zhang et al. 2019).
Endoplasmic reticulum (ER) homeostasis is maintained through well-conserved mechanisms in eukaryotes. It is a vital balance between protein load and capacity of ER folding machinery. When it is broken by either intracellular or extracellular stress conditions, unfolded protein response (UPR) is activated to reduce the ER load or to increase the folding capacity of ER (Walter and Ron 2011). There are three transmembrane proteins, responsible from three separate signaling pathways that mediates unfolded protein response in mammalian cells: activating transcription factor 6 (ATF6), inositol-requiring kinase/endoribonuclease 1 (IRE1), and protein kinase RNA-like ER kinase (PERK) (Ma and Hendershot 2001). During ER homeostasis, these proteins interact with a chaperone protein commonly known as glucose-regulated proteins 78 (GRP78) or immunoglobulin heavy chain binding protein (BiP) (Kozutsumi et al. 1988). When UPR is activated, BiP dissociates from the transmembrane proteins and binds to misfolded proteins. While dissociation of BiP activates IRE1 and ATF6 for transcriptional regulation of UPR by increasing secretion and chaperone capacity of ER, it also activates PERK for translational attenuation. Failure of these mechanisms to eliminate ER stress results in activation of various cell death pathways (Peng et al. 2021).
Malaria parasites have a secretome of at least 320 identified proteins (Hiller et al. 2004). Excessive amounts of protein trafficking are necessary for the establishment of infection in the host cell, which alone could overload ER to activate UPR. Malaria parasites harbor a highly reduced UPR, in which transcriptional regulators of the UPR are absent, but few homologues of UPR components such as BiP and PERK have been identified (Kaiser et al. 2016).
Although not fully understood, there are many studies reporting that Plasmodium parasites harbor apoptosis-like PCD pathways exhibiting common signs of apoptosis such as chromatin condensation and DNA fragmentation (Pollitt et al. 2010; Meslin et al. 2007; Le Chat et al. 2007; Al-Olayan et al. 2002; Picot et al. 1997). Malaria parasites do not possess classical caspase family genes; instead, they have homologues of metacaspase genes MCA1, MCA2, and MCA3. (Kumari et al. 2022; Kumar et al. 2019; Le Chat et al. 2007). MCA1 and MCA2 are thought to be involved in these unique cell death pathways (Timothy and Zininga 2023).
On the other hand, Plasmodium parasites contain homologs of almost half of the yeast autophagy proteins. Erythrocytic stages of the parasite were shown to undergo autophagy-like cell death mechanism in which autophagosome-like vacuoles were observed under certain drug stresses (Navale et al. 2014; Hain and Bosch 2013; Kamil et al. 2022).
In our study, we aimed to analyze the expression profiles of autophagy-related genes (PbATG5, PbATG8, and PbATG12) and a metacaspase gene, PbMCA2, to probe the responses of the rodent malaria parasite Plasmodium berghei against well-known ER stress inducers. We reasoned that the ex vivo Plasmodium berghei culture would be a more appropriate model for measuring the effect of drug-induced ER stress for mimicking the mammalian infection due to the possibility of alterations in transcriptomics, virulence factors, and drug sensitivity of the extensively in vitro cultured, lab-adapted strains of P. falciparum (Brown and Guler 2020).
Materials and methods
Ethics statement
All animal experiments described here were performed in accordance with the Experimental Animals Ethical Committee of Bezmialem Vakif University, Istanbul, Turkey (no. 2020/235).
Parasite cultures and drug treatment
Six to eight-week-old female CD1 mice were purchased from Bezmialem Vakif University, Experimental Animal Research Center. Donor mice were infected intraperitoneally with cryopreserved stocks of wild type-like eGFP expressing P. berghei Pbp230p(-) strain. This strain was recently shown to have similar growth characteristics as wild-type parasites (Kamil et al. 2024). Parasitemia of the donor mice was monitored by flow cytometry. Infected mice blood was collected by cardiac puncture when parasitemia reached 5%. Parasite life cycle stages were synchronized by culturing overnight in RPMI 1640 supplemented with 20% heat-inactivated FBS and 20 µg/mL gentamicin. Synchronized schizonts were collected by density gradient centrifugation. Briefly, 10 mL 60% (v/w) Accudenz solution was added gently into the bottom of a 35 mL parasite culture mix in a 50-mL centrifuge tube. Mix was carefully centrifuged at 200 g for 20 min with no brakes. Schizonts formed brown-gray ring in between two phases, and they were collected by using long, glass Pasteur pipettes. Synchronized schizonts were injected into naïve mice intravenously.
Mice infected with synchronized parasites were bled after 48 h by cardiac puncture, and collected blood was mixed with parasite culture medium. Culture was aliquoted into six-well plates to create ex vivo experiment groups (three wells per group). Parasitemia of cultures was monitored at the beginning and every 2.5 h at a dilution of 1:200 in PBS.
Non-lethal concentration of drugs was used that do not cause parasite death but induce cell stress (Unal 2022). The concentrations of drugs were 0.25 µg/mL chloroquine (CQ), 5 µg/mL tunicamycin (TN), and 2.5 µg/mL 5-fluorouracil (5-FU).
RT-qPCR analysis
Three milliliters of cultures were collected at 5th hour, pelleted at 300 g for 5 min, and washed with 500 µL PBS. RNA isolation was performed by using RNeasy Kit (Qiagen), and cDNA synthesis was performed by using High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems).
Synthesized cDNAs were used in qPCR analysis to probe ER stress-related genes (PbBiP), autophagy genes (PbATG5, PbATG8, and PbATG12), and a metacaspase gene (PbMCA2). 18S rRNA housekeeping gene was used as reference. Reactions were performed in BioRad CFx96 real-time PCR system by using iTaq SYBR Green Master Mix (BioRad) according to the manufacturer’s instructions. Primer sequences are given in Table 1.
Statistical analysis
Statistically significant differences between the median values of treatment groups were evaluated by the analysis of variance (ANOVA) method in all experiments. GraphPad Prism 9 software was used for all analyses. p values of 0.05 were considered statistically significant.
Results
Treatment with sub-lethal drug doses does not affect parasite growth
Parasite cultures were treated with sub-lethal doses of CQ, TN, and 5-FU that do not cause death but generate stress response in the parasite (Unal 2022). Parasite survival was measured by flow cytometry every 2.5 h. As expected, applied drugs had no significant effect on parasite survival (Fig. 1).
Drug treatments do not affect parasite growth. Parasite cultures were treated with 0.25 µg/mL CQ, 5 µg/mL TN, and 2.5 µg/mL 5-FU. Parasite growth was recorded every 2.5 h by flow cytometry. Experiments were performed in three replicates. Results were shown as percent parasitemia for each time point. Bars represent mean ± SD values for each treatment
Drug treatments induce elevated levels of ER stress
To evaluate whether the drugs induced ER stress, we checked the transcription profile of ER resident chaperone protein GRP78 (BiP) gene (Fig. 2). TN treatment upregulated PbBiP transcription ~ 40 folds, while CQ and 5-FU treatments upregulated ~ 150 and ~ 190 folds, respectively, indicating a strong ER stress response under all three drug treatments.
Drug treatments significantly upregulate PbBiP transcription. Parasite cultures were treated with 0.25 µg/mL CQ, 5 µg/mL TN, and 2.5 µg/mL 5-FU. Five hours later, total RNA was isolated for cDNA synthesis from 3 mL of culture wells. All qPCR reaction mixtures contained equal amount of cDNA from the treated or untreated cultures. Chloroquine (CQ), tunicamycin (TN), and 5-fluorouracil (5-FU) treatments caused ~ 150, ~ 40, and ~ 190-folds upregulation of PbBiP, respectively. Experiments were performed in three replicates. Results were shown as fold changes relative to the untreated control. Each bar shows mean ± SD values for each gene. *p < 0.05, and ****p < 0.0001
Drug-induced ER stress induces upregulation of autophagy-related genes
Prolonged effects of ER stress are known to induce autophagy and apoptosis pathways (Galluzzi et al. 2017; Kamil et al. 2017). Therefore, we further analyzed the transcription levels of autophagy-related genes (PbATG5, PbATG8, and PbATG12), and the metacaspase, PbMCA2, gene under drug-induced ER stress conditions to see the effect of ER stress on common markers of PCD pathways. All four genes showed elevated levels of transcription under CQ (Fig. 3A) and 5-FU (Fig. 3B) treatments. In contrast, ATG5 and MCA2 were upregulated, but ATG8 and ATG12 were downregulated under TN treatment (Fig. 3C).
Drug-induced ER-stress alters the transcription levels of autophagy and apoptosis-related genes. Parasite cultures were treated with 0.25 µg/mL CQ, 5 µg/mL TN, and 2.5 µg/mL 5-FU. Five hours later, total RNA was isolated for cDNA synthesis from 3 mL of culture wells. All qPCR reaction mixtures contained equal amount of cDNA from the treated or untreated cultures. A Chloroquine (CQ) treatment upregulated all genes, but PbATG5 and PbATG12 transcription levels were significantly higher than PbATG8 and PbMCA2. B Tunicamycin (TN) treatment resulted in downregulation of PbATG8 and PbATG12, and upregulation of PbATG5 and PbMCA2. C 5-Fluorouracil (5-FU) treatments upregulated all genes similar to CQ treatment. Experiments were performed in three replicates. Results were shown as fold changes relative to the untreated control. The positive fold changes indicate the upregulation as a result of the drug treatments, and the negative fold changes indicate the downregulation. Each bar shows mean ± SD values for each gene. *p < 0.05, ***p < 0.001, and ****p < 0.0001
Discussion
The life cycle of Plasmodium parasites involves two distinct host organisms, multiple tissues, various host cells, and very different environments. Parasites adapt to changing conditions by constantly altering their morphology and physiology, as well as altering their host cells. These alterations naturally require heavy trafficking of large repertoire of intracellular and secreted proteins that leads to heavy protein loads in ER, consequently triggering stress responses (Kaiser et al. 2016). Exploiting ER stress is a common strategy in drug development, but repurposing of existing drugs is also attracting attention. Therefore, it is important to understand parasitic stress responses and PCD pathways of parasitic protozoans in response to chemical compounds.
In this study, the effects of three ER stressor drugs, with various effector mechanisms, on the transcriptions of three autophagy genes and one metacaspase gene were investigated. Ex vivo P. berghei cultures were treated with the selected drugs: chloroquine, tunicamycin, and 5-fluorouracil.
Chloroquine is the most common quinoline derivate antimalarial which inhibits detoxification of hematin, a byproduct of host erythrocyte hemoglobin, causing high toxicity. In addition, quinoline derivatives have been postulated to have various antimalarial and anticancer roles such as inhibition of tyrosine kinase (Kaur et al. 2010) and inhibition of autophagosome-lysosome fusion (Mauthe et al. 2018; Kamil et al. 2022).
Tunicamycin is known for its ability to induce ER stress by inhibiting N-glycosylation of proteins, resulting in accumulation of unfolded proteins in the ER lumen. It has antiviral, antibiotic, antifungal, and antitumor activities (Myers et al. 1993), but is not suitable for clinical use due to its toxicity in mammalian cells.
5-Fluorouracil is used as an anticancer drug, and it was shown to induce high levels of ER stress (Yao et al. 2020). It also has an indirect role in negative drug selection in genetic manipulation of Plasmodium species due to its high toxicity (Manzoni et al. 2014). 5-FU is a uracil analogue that has a fluorine atom at its fifth carbon. It is transported and converted to fluorodeoxyuridine monophosphate (FdUMP) which binds and inhibits thymidine synthase, preventing the synthesis of deoxythymidine monophosphate (dTMP). Depletion of dTMP leads to severe disruption of DNA synthesis and repair (Longley et al. 2003). 5-FU might create more severe effects in Plasmodium species due to the extremely AT-rich genome structure.
Our results showed that treatments of parasite cultures with all three drugs induced significant levels of ER stress and altered transcription profiles of autophagy and apoptosis-related genes. Tunicamycin-induced ER stress usually leads to apoptosis (Guha et al. 2017). TN treatment upregulated MCA2 as expected, but it also upregulated ATG5 together with the downregulation of ATG8 and ATG12. Atg5 had previously been shown to have pro-apoptotic roles in mammalian cells (Codogno and Meijer 2006). This expression profile suggests induction of an autophagy-induced apoptosis-like cell death mechanism. Same line of thinking might also be applied to expression profiles of CQ and 5-FU treatments; induction of all four genes could be explained by a similar autophagy-induced apoptosis-like pathway. Prolonged or excessive autophagy is known to lead to cell death. Other likely explanation is the excessive levels of ER stress could induce more than one response pathways simultaneously. Similarly, DNA damage can induce apoptosis, but it has also been shown to induce autophagy (Rodriguez-Rocha et al. 2011). Therefore, DNA replication and repair inhibitions by 5-FU treatments might lead to activation of both autophagy and apoptosis-like pathways.
CQ and 5-FU treatments induced significantly elevated levels of BiP upregulation, but as a noticeable difference, CQ treatment caused drastically upregulated transcription levels of ATG5 and ATG12 genes. Atg5 and Atg12 are required to form autophagopore, and Atg8 is required for the elongation and closure of autophagosome membrane (Hain and Bosch 2013). In previous studies, CQ treatment resulted with the cytosolic accumulation of autophagic vesicles, and CQ treatment was reported to inhibit autophagosome-lysosome fusion in an autophagy-independent manner (Mauthe et al. 2018). Thus, noticeable upregulation of the ATG5 and ATG12 transcripts in response to CQ treatment could be the result of autophagosome-lysosome fusion inhibition.
Although this is a limited study, our results indicate that Plasmodium parasites respond to various ER stressors by inducing autophagy- and/or apoptosis-like pathways. Atg8 is generally accepted as a classical autophagy marker, but as an important limitation of this study, there are no established markers for apoptosis in Plasmodium. More work is required to overcome this limitation and uncover PCD pathways of malaria parasites. Further studies are warranted to explore relationship between autophagic and apoptotic pathways, especially at the translational level, which is likely to provide interesting results.
Data availability
Data will be made publicly available upon publication and upon request for peer review.
References
Al-Olayan EM, Williams GT, Hurd H (2002) Apoptosis in the malaria protozoan, Plasmodium berghei: a possible mechanism for limiting intensity of infection in the mosquito. Int J Parasitol 32:1133–1143. https://doi.org/10.1016/S0020-7519(02)00087-5
Brown AC, Guler JL (2020) From circulation to cultivation: Plasmodium in vivo versus in vitro. Trends Parasitol 36:914–926. https://doi.org/10.1016/j.pt.2020.08.008
Codogno P, Meijer AJ (2006) Atg5: more than an autophagy factor. Nat Cell Biol 8:1045–1047. https://doi.org/10.1038/ncb1006-1045
Galluzzi L, Diotallevi A, Magnani M (2017) Endoplasmic reticulum stress and unfolded protein response in infection by intracellular parasites. Future Sci OA 3:Fso198. https://doi.org/10.4155/fsoa-2017-0020
Guha P, Kaptan E, Gade P et al (2017) Tunicamycin induced endoplasmic reticulum stress promotes apoptosis of prostate cancer cells by activating mTORC1. Oncotarget 8:68191–68207. https://doi.org/10.18632/oncotarget.19277
Hain AU, Bosch J (2013) Autophagy in Plasmodium, a multifunctional pathway? Comput Struct Biotechnol J 8:e201308002. https://doi.org/10.5936/csbj.201308002
Hiller NL, Bhattacharjee S, van Ooij C et al (2004) A host-targeting signal in virulence proteins reveals a secretome in malarial infection. Science 306:1934–1937. https://doi.org/10.1126/science.1102737
Kaiser G, De Niz M, Zuber B et al (2016) High resolution microscopy reveals an unusual architecture of the Plasmodium berghei endoplasmic reticulum. Mol Microbiol 102:775–791. https://doi.org/10.1111/mmi.13490
Kamil M, Atmaca HN, Unal S et al (2022) An alternative autophagy-related mechanism of chloroquine drug resistance in the malaria parasite. Antimicrob Agents and Chemother 66:e00269-00222.https://doi.org/10.1128/aac.00269-22
Kamil M, Haque E, Irfan S et al (2017) ER chaperone GRP78 regulates autophagy by modulation of p53 localization. Front Biosci (Elite Ed) 9:54–66. https://doi.org/10.2741/e785
Kamil M, Kina UY, Atmaca HN et al (2024) Endoplasmic reticulum localized TMEM33 domain-containing protein is crucial for all life cycle stages of the malaria parasite. Mol Microbiol. https://doi.org/10.1111/mmi.15228
Kaur K, Jain M, Reddy RP et al (2010) Quinolines and structurally related heterocycles as antimalarials. Eur J Med Chem 45:3245–3264. https://doi.org/10.1016/j.ejmech.2010.04.011
Kozutsumi Y, Segal M, Normington K et al (1988) The presence of malfolded proteins in the endoplasmic reticulum signals the induction of glucose-regulated proteins. Nature 332:462–464. https://doi.org/10.1038/332462a0
Kumar B, Verma S, Kashif M et al (2019) Metacaspase-3 of Plasmodium falciparum: an atypical trypsin-like serine protease. Int J Biol Macromol 138:309–320. https://doi.org/10.1016/j.ijbiomac.2019.07.067
Kumari V, Prasad KM, Kalia I et al (2022) Dissecting the role of Plasmodium metacaspase-2 in malaria gametogenesis and sporogony. Emerg Microbes Infect 11:938–955. https://doi.org/10.1080/22221751.2022.2052357
Laudisi F, Di Grazia A, De Simone V et al (2019) Induction of endoplasmic reticulum stress and inhibition of colon carcinogenesis by the anti-helmintic drug rafoxanide. Cancer Lett 462:1–11. https://doi.org/10.1016/j.canlet.2019.07.014
Le Chat L, Sinden RE, Dessens JT (2007) The role of metacaspase 1 in Plasmodium berghei development and apoptosis. Mol Biochem Parasitol 153:41–47. https://doi.org/10.1016/j.molbiopara.2007.01.016
Longley DB, Harkin DP, Johnston PG (2003) 5-Fluorouracil: mechanisms of action and clinical strategies. Nat Rev Cancer 3:330–338. https://doi.org/10.1038/nrc1074
Ma Y, Hendershot LM (2001) The unfolding tale of the unfolded protein response. Cell 107:827–830. https://doi.org/10.1016/S0092-8674(01)00623-7
Manzoni G, Briquet S, Risco-Castillo V et al (2014) A rapid and robust selection procedure for generating drug-selectable marker-free recombinant malaria parasites. Sci Rep 4:4760. https://doi.org/10.1038/srep04760
Mauthe M, Orhon I, Rocchi C et al (2018) Chloroquine inhibits autophagic flux by decreasing autophagosome-lysosome fusion. Autophagy 14:1435–1455. https://doi.org/10.1080/15548627.2018.1474314
Meslin B, Barnadas C, Boni V et al (2007) Features of apoptosis in Plasmodium falciparum Erythrocytic stage through a putative role of PfMCA1 metacaspase-like protein. J Infect Dis 195:1852–1859. https://doi.org/10.1086/518253
Myers AG, Gin DY, Rogers DH (1993) A convergent synthetic route to the tunicamycin antibiotics. Synthesis of (+)-tunicamycin V. J Am Chem Soc 115:2036–2038. https://doi.org/10.1021/ja00058a060
Navale R, Atul AAD et al (2014) Characterization of the autophagy marker protein Atg8 reveals atypical features of autophagy in Plasmodium falciparum. PLoS ONE 9:e113220. https://doi.org/10.1371/journal.pone.0113220
Peng M, Chen F, Wu Z et al (2021) Endoplasmic reticulum stress, a target for drug design and drug resistance in parasitosis. Front Microbiol 12. https://doi.org/10.3389/fmicb.2021.670874
Picot S, Burnod J, Bracchi V et al (1997) Apoptosis related to chloroquine sensitivity of the human malaria parasite Plasmodium falciparum. Trans R Soc Trop Med Hyg 91:590–591. https://doi.org/10.1016/s0035-9203(97)90039-0
Pollitt LC, Colegrave N, Khan SM et al (2010) Investigating the evolution of apoptosis in malaria parasites: the importance of ecology. Parasit Vectors 3:105. https://doi.org/10.1186/1756-3305-3-105
Rodriguez-Rocha H, Garcia-Garcia A, Panayiotidis MI et al (2011) DNA damage and autophagy. Mutat Res 711:158–166. https://doi.org/10.1016/j.mrfmmm.2011.03.007
Timothy FM, Zininga T (2023) Small heat shock proteins as modulators of cell death in Plasmodium falciparum parasites and its human host. Front Cell Death 2. https://doi.org/10.3389/fceld.2023.1322780
Unal S (2022) Characterization of endoplasmic reticulum stress-induced cell death pathways in rodent malaria parasites. Master’s Thesis (Thesis Number: 732310) Istanbul University, Turkey. https://tez.yok.gov.tr/UlusalTezMerkezi/TezGoster?key=kScA8XnrRb0WogXqPGFku88Vbh5NbLzs4QRjCJ_1Cc9Hx_Vp5CztRx3aYkE0syr
Walter P, Ron D (2011) The unfolded protein response: from stress pathway to homeostatic regulation. Science 334:1081–1086. https://doi.org/10.1126/science.1209038
World Malaria Report (2023) World Health Organization 2023, Geneva. Licence: CC BY-NC-SA 3.0 IGO
Yao X, Tu Y, Xu Y et al (2020) Endoplasmic reticulum stress confers 5-fluorouracil resistance in breast cancer cell via the GRP78/OCT4/lncRNA MIAT/AKT pathway. Am J Cancer Res 10:838–855
Zhang Z, Gao W, Zhou L et al (2019) Repurposing brigatinib for the treatment of colorectal cancer based on inhibition of ER-phagy. Theranostics 9:4878–4892. https://doi.org/10.7150/thno.36254
Acknowledgements
The authors are thankful to Elif Kurt for her help in animal care and insectary maintenance. The authors are thankful to the Beykoz Institute of Life Sciences and Biotechnology, Bezmialem Vakif University, for providing all necessary support.
Funding
Open access funding provided by the Scientific and Technological Research Council of Türkiye (TÜBİTAK). This work was supported by the Scientific Research Projects Coordination Unit of Istanbul University. Project number: 34860.
Author information
Authors and Affiliations
Contributions
Conceptualization: Bedia Palabiyik and Ahmed Sayed Ibrahim Aly. Funding acquisition: Bedia Palabiyik and Ahmed Sayed Ibrahim Aly. Methodology: Sinem Unal, Umit Yasar Kina, Mohd Kamil. Formal analysis: Umit Yasar Kina, Mohd Kamil. Writing—Original Draft: Sinem Unal, Umit Yasar Kina. Visualization: Umit Yasar Kina. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This study was reviewed and approved by the Experimental Animals Ethical Committee of Bezmialem Vakif University, Istanbul, Turkey (permit number: 2020/235).
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Handling Editor: Una Ryan
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Sinem Unal and Umit Y. Kina contributed equally to this work and share first authorship.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Unal, S., Kina, U.Y., Kamil, M. et al. Drug-induced ER stress leads to induction of programmed cell death pathways of the malaria parasite. Parasitol Res 123, 263 (2024). https://doi.org/10.1007/s00436-024-08281-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00436-024-08281-3