Abstract
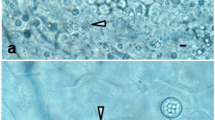
This study examined the morphological and molecular characteristics of the microsporidium Endoreticulatus sp. Zhenjiang, isolated from the silkworm (Bombyx mori). The fresh spores were oval, 2.9 ± 0.2 μm in length and 1.2 ± 0.2 μm in width. The complete rRNA cistron has a length of 4,432 bp (GenBank accession no. FJ772431), including the large subunit rRNA (2,460 bp), the internal transcribed spacer (187 bp), the small subunit rRNA (1,254 bp), the intergenic spacer (276 bp), and the 5S region (115 bp). The organization of the rRNA gene is 5′-LSU-ITS-SSU-IGS-5S-3′, which is reverse compared to the organization of most microsporidian rRNA regions. Phylogenetic analysis based on small subunit rRNA sequences showed that this isolate belongs to the genus Endoreticulatus, and is closely related to Glugoides intestinalis. Furthermore, both had a similar reverse arrangement of the rRNA gene. Our study provides another example of a microsporidian species with a novel organization of rRNA genes, demonstrating that the reverse arrangement is exhibited not only by the microsporidian genus Nosema but may also occur in a clade that contains the genera Endoreticulatus and Glugoides.



Similar content being viewed by others
References
Adl SM, Simpson AG, Farmer MA, Andersen RA, Anderson OR, Barta JR, Bowser SS, Brugerolle G, Fensome RA, Fredericq S, James TY, Karpov S, Kugrens P, Krug J, Lane CE, Lewis LA, Lodge J, Lynn DH, Mann DG, McCourt RM, Mendoza L, Moestrup O, Mozley-Standridge SE, Nerad TA, Shearer CA, Smirnov AV, Spiegel FW, Taylor MF (2005) The new higher level classification of eukaryotes with emphasis on the taxonomy of protists. J Eukaryot Microbiol 52:399–451
Baker MD, Vossbrinck CR, Maddox JV, Undeen AH (1994) Phylogenetic relationships among Vairimorpha and Nosema species (Microspora) based on ribosomal RNA sequence data. J Invertebr Pathol 64:100–106
Becnel JJ, Andreadis TG (1999) Microsporidia in insects. In: Witter M, Weiss L.M. (eds) The Microsporidia and Microsporidiosis. American Society of Microbiology Press pp 447–501
Bhat SA, Bashir I, Kamili AS (2009) Microsporidiosis of silkworm, Bombyx mori L. (Lepidoptera-bombycidae): a review. Afr J Agric Res 4:1519–1523
Corradi N, Keeling PJ (2009) Microsporidia: a journey through radical taxonomical revisions. Fungal Biol Rev 23:1–8
Dong SN, Shen ZY, Xu L, Zhu F (2010) Sequence and phylogenetic analysis of SSU rRNA gene of five microsporidia. Curr Microbiol 60:30–37
Franzen C, Müller A (1999) Molecular techniques for detection, species differentiation, and phylogenetic analysis of microsporidia. Clin Microbiol Rev 12:243–285
Goldberg AV, Molik S, Tsaousis AD, Neumann K, Kuhnke G, Delbac F (2008) Localization and functionality of microsporidian iron-sulphur cluster assembly proteins. Nature 452:624–628
Hibbett DS, Binder M, Bischoff JF, Blackwell M, Cannon PF, Eriksson OE, Huhndorf S, James T, Kirk PM, Lücking R, Thorsten Lumbsch H, Lutzoni F, Matheny PB, McLaughlin DJ, Powell MJ, Redhead S, Schoch CL, Spatafora JW, Stalpers JA, Vilgalys R, Aime MC, Aptroot A, Bauer R, Begerow D, Benny GL, Castlebury LA, Crous PW, Dai YC, Gams W, Geiser DM, Griffith GW, Gueidan C, Hawksworth DL, Hestmark G, Hosaka K, Humber RA, Hyde KD, Ironside JE, Kõljalg U, Kurtzman CP, Larsson KH, Lichtwardt R, Longcore J, Miadlikowska J, Miller A, Moncalvo JM, Mozley-Standridge S, Oberwinkler F, Parmasto E, Reeb V, Rogers JD, Roux C, Ryvarden L, Sampaio JP, Schüssler A, Sugiyama J, Thorn RG, Tibell L, Untereiner WA, Walker C, Wang Z, Weir A, Weiss M, White MM, Winka K, Yao YJ, Zhang N (2007) A higher-level phylogenetic classification of the Fungi. Mycol Res 111:509–547
Huang WF, Tsai SJ, Lo CF, Soichi Y, Wang CH (2004) The novel organization and complete sequence of the ribosomal RNA gene of Nosema bombycis. Fungal Genet Biol 41:473–481
Ironside JE (2007) Multiple losses of sex within a single genus of Microsporidia. BMC Evol Biol 29:7–48
James TY, Kauff F, Schoch CL, Matheny PB, Hofstetter V, Cox CJ, Celio G, Gueidan C, Fraker E, Miadlikowska J, Lumbsch HT, Rauhut A, Reeb V, Arnold AE, Amtoft A, Stajich JE, Hosaka K, Sung GH, Johnson D, O'Rourke B, Crockett M, Binder M, Curtis JM, Slot JC, Wang Z, Wilson AW, Schüssler A, Longcore JE, O'Donnell K, Mozley-Standridge S, Porter D, Letcher PM, Powell MJ, Taylor JW, White MM, Griffith GW, Davies DR, Humber RA, Morton JB, Sugiyama J, Rossman AY, Rogers JD, Pfister DH, Hewitt D, Hansen K, Hambleton S, Shoemaker RA, Kohlmeyer J, Volkmann-Kohlmeyer B, Spotts RA, Serdani M, Crous PW, Hughes KW, Matsuura K, Langer E, Langer G, Untereiner WA, Lücking R, Büdel B, Geiser DM, Aptroot A, Diederich P, Schmitt I, Schultz M, Yah R, Hibbett DS, Lutzoni F, McLaughlin DJ, Spatafora JW, Vilgalys R (2006) Reconstructing the early evolution of fungi using a six-gene phylogeny. Nature 443:818–822
Johny S, Kanginakudru S, Muralirangan MC, Nagaraju J (2006) Morphological and molecular characterization of a new microsporidian (Protozoa: Microsporidia) isolated from Spodoptera litura (Fabricius) (Lepidoptera: Noctuidae). Parasitology 132:1–12
Keeling PJ, Doolittle WF (1996) Alpha-tubulin from early diverging eukaryotic lineages and the evolution of the tubulin family. Mol Biol Evol 13:1297–1305
Ku CT, Wang CY, Tsai YC, Tzeng CC, Wang CH (2007) Phylogenetic analysis of two putative Nosema isolates from cruciferous lepidopteran pests in Taiwan. J Invertebr Pathol 95:71–76
Kyei-Poku G, Gauthier D, Van Frankenhuyzen K (2008) Molecular data and phylogeny of Nosema infecting Lepidopteran forest defoliators in the genera Choristoneura and Malacosoma. J Eukaryot Microbiol 55:51–58
Larsson JIR (1999) Identification of microsporidia. Acta Protozoologica 38:161–197
Nicholas KB, Nicholas HB, Deerfield DW (1997) GeneDoc: analysis and visualization of genetic variation. EMBNEW News 4:14
Refardt D, Mouton L (2007) Reverse arrangement of rRNA subunits in the microsporidium Glugoides intestinalis. J Eukaryot Microbiol 54:83–85
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Texier C, Vidau C, Vigues B, El Alaoui H, Delbac F (2010) Microsporidia: a model for minimal parasite-host interactions. Curr Opin Microbiol 13:443–449
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tsai SJ, Yamane S, Lo CF, Wang CH (2003) The characterization of microsporidian isolates (Nosematidae: Nosema) from five important lepidopteran pests in Taiwan. J Invertebr Pathol 83:51–59
Tsai SJ, Huang WF, Wang CH (2005) Complete sequence and gene organization of the Nosema spodopterae rRNA gene. J Eukaryot Microbiol 52:52–54
Vossbrinck CR, Debrunner-Vossbrinck BA (2005) Molecular phylogeny of the Microsporidia: ecological, ultrastructural and taxonomic considerations. Folia Parasitol 52:131–142
Vossbrinck CR, Baker MD, Didier ES, Debruuner-Vossbrinck BA, Shadduck JA (1993) Ribosomal DNA sequences of Encephalitozoon hellem and Encephalitozoon cuniculi: species identification and phylogenetic construction. J Eukaryot Microbiol 40:354–362
Wan YJ, Zhang L, Chen ZP (1995) Study of a pathogenic microsporidia SCM7 (Endoreticulatus sp.) isolated from the larva of silkworm, Bombyx mori. Acta Sericologica Sinica 21:168–173
Wang LL, Chen KP, Zhang Z, Yao Q, Gao GT, Zhao Y (2006) Phylogenetic analysis of Nosema antheraeae (Microsporidia) isolated from Chinese oak silkworm, Antheraea pernyi. J Eukaryot Microbiol 53:310–313
Weiss LM, Vossbrinck CR (1998) Microsporidiosis: molecular and diagnostic aspects. Adv Parasitol 40:351–395
Williams BA, Hirt RP, Lucocq JM, Embley TM (2002) A mitochondrial remnant in the microsporidian Trachipleistophora hominis. Nature 418:865–869
Zhu F, Shen ZY, Xu XF, Tao HP, Dong SN, Tang XD, Xu L (2010) Phylogenetic analysis of complete rRNA gene sequence of Nosema philosamiae isolated from the Lepidopteran Philosamia cynthia ricini. J Eukaryot Microbiol 57:294–296
Acknowledgments
This work was supported by the earmarked fund for the Modern Agro-industry Technology Research System. We are grateful to all who provided the means for us to access free software, which we have used and cited in this article. We thanked all partners and lab members for the kindly help and criticism.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
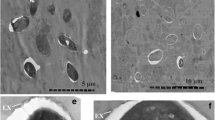
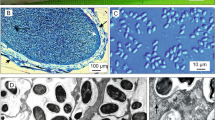
Fig. S1a
Transmission electron micrographs of Endoreticulatus sp. Zhenjiang. PV posterior vacuole. Scale bar = 0.2 μm (JPEG 7 kb)
Fig. S1b
Transmission electron micrographs of Endoreticulatus sp. Zhenjiang. PF polar filament. Scale bar = 0.2 μm (JPEG 7 kb)
Fig. S2
Phylogenetic analysis of the Endoreticulatus sp. Zhenjiang based on rRNA sequences. The numbers on the branches indicate the percentage of bootstrap replicates that supported the given topology based on the neighbor-joining method using MEGA 4.0 software. Encephalitozoon cuniculi was used as the outgroup (JPEG 54 kb)
Fig. S3
Phylogenetic analysis of the Endoreticulatus sp. Zhenjiang based on LSU rRNA subunit sequences. The numbers on the branches indicate the percentage of bootstrap replicates that supported the given topology based on the neighbor-joining method using MEGA 4.0 software. Encephalitozoon cuniculi was used as the outgroup (JPEG 46 kb)
Rights and permissions
About this article
Cite this article
Xu, X., Shen, Z., Zhu, F. et al. Phylogenetic characterization of a microsporidium (Endoreticulatus sp. Zhenjiang) isolated from the silkworm, Bombyx mori . Parasitol Res 110, 815–819 (2012). https://doi.org/10.1007/s00436-011-2560-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-011-2560-8