Abstract
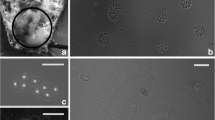
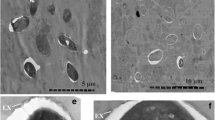
The spore morphology, chromosomal karyotype, and molecular systematic of a new microsporidian which was isolated from the domesticated silkworm Bombyx mori (Lepidoptera: Bombycidae) in Shandong, China have been studied. The spores were long oval and measured 3.4 × 1.6 μm on fresh smears. Ultrastructure of the spores was characteristic for the genus Vairimorpha: 13–15 polar filament coils, posterior vacuole, and a diplokaryon. Six chromosome bands have been separated by pulsed field gel electrophoresis. The sequenced complete rRNA gene of this isolate is 4,231 bp long. Phylogenetic analysis based on SSU rRNA gene and LSU rRNA gene both revealed that this novel microsporidian which was isolated from B. mori had close relationship to the genus Vairimorpha, not to the genus Nosema. Moreover, the organization of the rRNA units of this microsporidian is not similar to that of Nosema bombycis, but same to that of other microsporidian, such as Vairimorpha necatrix. Although this microsporidian, designed as Vairimorpha sp. BM, was isolated from B. mori, all of these informations indicate that this isolate is closely related to the Vairimorpha group.




Similar content being viewed by others
References
Canning EU, Curry A, Cheney S, Lafranchi-Tristem NJ, Haque MA (1999) Vairimorpha imperfecta n. sp., a microsporidian exhibiting an abortive octosporous sporogony in Plutella xylostella L. (Lepidoptera: Yponomeutidae). Parasitology 119(Pt 3):273–286
Choi JY, kIM JG, Choi YC, Goo TW, Chang JH, Je YH, Kim KY (2002) Nosema sp. isolated from cabbage white butterfly (Pieris rapae) collected in Korea. J Microbiol 40:199–204
Corradi N, Haag KL, Pombert JF, Ebert D, Keeling PJ (2009) Draft genome sequence of the Daphnia pathogen Octosporea bayeri: insights into the gene content of a large microsporidian genome and a model for host–parasite interactions. Genome Biol 10:R106
Down RE, Bell HA, Kirkbride-Smith AE, Edwards JP (2004) The pathogenicity of Vairimorpha necatrix (Microspora: Microsporidia) against the tomato moth, Lacanobia oleracea (Lepidoptera: Noctuidae) and its potential use for the control of lepidopteran glasshouse pests. Pest Manag Sci 60:755–764
Franzen C, Muller A (1999) Molecular techniques for detection, species differentiation, and phylogenetic analysis of microsporidia. Clin Microbiol Rev 12:243–285
Huang WF, Tsai SJ, Lo CF, Soichi Y, Wang CH (2004) The novel organization and complete sequence of the ribosomal RNA gene of Nosema bombycis. Fungal Genet Biol 41:473–481
Joseph J, Sharma S, Murthy SI, Krishna PV, Garg P, Nutheti R, Kenneth J, Balasubramanian D (2006) Microsporidial keratitis in India: 16S rRNA gene-based PCR assay for diagnosis and species identification of microsporidia in clinical samples. Invest Ophthalmol Vis Sci 47:4468–4473
Kawakami Y, Inoue T, Ito K, Kitamizu K, Hanawa C, Ando T, Iwano H, Ishihara R (1994) Identification of a chromosome harboring the small subunit ribosomal RNA gene of Nosema bombycis. J Invertebr Pathol 64:147–148
Ku CT, Wang CY, Tsai YC, Tzeng CC, Wang CH (2007) Phylogenetic analysis of two putative Nosema isolates from Cruciferous Lepidopteran pests in Taiwan. J Invertebr Pathol 95:71–76
Liu H, Pan G, Song S, Xu J, Li T, Deng Y, Zhou Z (2008) Multiple rDNA units distributed on all chromosomes of Nosema bombycis. J Invertebr Pathol 99:235–238
Malone LA, McIvor CA (1993) Pulsed-field gel electrophoresis of DNA from four microsporidian isolates. J Invertebr Pathol 61:203–205
Munderloh UG, Kurtti TJ, Ross SE (1990) Electrophoretic characterization of chromosomal DNA from two microsporidia. J Invertebr Pathol 56:243–248
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sprague V (1982) Microspora. In: Parker SP (ed) Synopsis and classification of living organism, vol. 1. McGraw-Hill, New York, pp 589–594
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tsai SJ, Lo CF, Soichi Y, Wang CH (2003) The characterization of microsporidian isolates (Nosematidae: Nosema) from five important lepidopteran pests in Taiwan. J Invertebr Pathol 83:51–59
Tsai SJ, Huang WF, Wang CH (2005) Complete sequence and gene organization of the Nosema spodopterae rRNA gene. J Eukaryot Microbiol 52:52–54
Tsai YC, Solter LF, Wang CY, Fan HS, Chang CC, Wang CH (2009) Morphological and molecular studies of a microsporidium (Nosema sp.) isolated from the thee spot grass yellow butterfly, Eurema blanda arsakia (Lepidoptera: Pieridae). J Invertebr Pathol 100:85–93
Wang LL, Chen KP, Zhang Z, Yao Q, Gao GT, Zhao Y (2006) Phylogenetic analysis of Nosema antheraeae (Microsporidia) isolated from Chinese oak silkworm, Antheraea pernyi. J Eukaryot Microbiol 53:310–313
Wittner M, Weiss LM (1999) The microsporidia and microsporidiosis. ASM Press, Washington, p 553
Acknowledgments
The authors are grateful to senior agronomist Zhaoyi Guo for providing the initial extract of the spore sample. We are grateful to everyone who generously provided us with access to the software used in this work. This work is supported by the National Natural Science Foundation of China under Grant No. 30930067, the Program of Introducing Talents of Discipline to Universities (No. B07045), and the Research Fund of the Doctoral Program of Southwest University under Grant No. swu109039.
Author information
Authors and Affiliations
Corresponding author
Additional information
Nucleotide sequence reported in this study has been submitted to the GenBank™, EMBL, and DDBJ databases under the accession number HQ891818.
Rights and permissions
About this article
Cite this article
Liu, H., Pan, G., Li, T. et al. Ultrastructure, chromosomal karyotype, and molecular phylogeny of a new isolate of microsporidian Vairimorpha sp. BM (Microsporidia, Nosematidae) from Bombyx mori in China. Parasitol Res 110, 205–210 (2012). https://doi.org/10.1007/s00436-011-2470-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-011-2470-9