Abstract
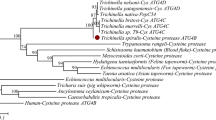
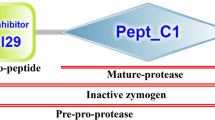
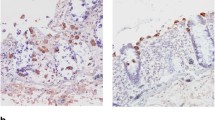
Although the functions of cysteine proteases involved in the pathogenicity and differentiation of Entamoeba histolytica have been demonstrated, little is known about the functions of serine proteases. We examined the involvement of serine proteases in amoebic excystation and metacystic development using inhibitors specific for serine proteases. Entamoeba invadens IP-1 strain was used as the model of excystation and metacystic development of E. histolytica. Four serine protease inhibitors, phenylmethanesulfonyl fluoride (PMSF), 4-(2-aminoethyl) bezensulfonylfluoride hydrochloride, 3, 4-dichloroisocoumarin, and N-tosyl-phe-chloromethylketone, decreased the number of metacystic amoebae in a dose-dependent manner, without showing cytotoxicity to cysts. PMSF inhibited not only the increase but also the development of metacystic amoebae as determined by the change of nucleus number from four- to one-nucleate amoebae. The protease activity in cyst lysates was also inhibited by PMSF and the band of protease on gelatin sodium dodecyl sulfate polyacrylamide gel electrophoresis was weaker than controls when treated with PMSF. Three serine protease families, S28 (three types), S9 (two), and S26 (one) were retrieved from the database of E. invadens. Phylogenetic analysis revealed that amebic enzymes from the serine protease families formed different clades from those from other organisms. The expression levels of these serine proteases in cysts 5 h after the induction of excystation as assessed by real-time reverse transcriptase polymerase chain reaction (RT-PCR) were higher than those observed prior to induction assayed by real-time RT-PCR; the increase in one type of S9 (named S9-3) expression was the highest. The expression of S9 enzymes also increased from cysts to trophozoites higher than the other family serine proteases. Thus, the results show that Entamoeba uses their serine proteases in the excystation and metacystic development, which leads to successful infection.









Similar content being viewed by others
References
Arroyo-Begovich A, Carbez-Trejo A (1982) Location of chitin in the cyst wall of Entamoeba invadens with the colloidal gold tracers. J Parasitol 68:253–258
Barrios-Ceballos MP, Martínez-Gallardo NA, Anaya-Velázquez F, Mirelman D, Padilla-Vaca F (2005) A novel protease from Entamoeba histolytica homologous to members of the family S28 of serine proteases. Exp Parasitol 110:270–275
Bruchhaus I, Loftus BJ, Hall N, Tannich E (2003) The intestinal protozoan parasite Entamoeba histolytica contains 20 cysteine protease genes, of which only a small subset is expressed during in vitro cultivation. Eukaryot Cell 2:501–509
Chávez-Munguía B, Cristóbal-Ramos AR, González-Robles A, Tsutsumi V, Martinez-Palomo A (2003) Ultrastructural study of Entamoeba invadens encystation and excystation. J Submicrosc Cytol Pathol 35:235–243
Chávez-Munguía B, Omaña-Molina M, González-Lázaro M, González-Robles A, Cedillo-Rivera R, Bonilla P, Martinez-Palomo A (2007) Ultrastructure of cyst differentiation in parasitic protozoa. Parasitol Res 100:1169–1175
Cleveland LR, Sanders EP (1930) Encystation, multiple fission without encystment, excystation, metacystic development, and variation in a pure line and nine strains of Entamoeba histolytica. Arch Protist 70:223–266
Diamond LS, Harlow DR, Cunnick CC (1978) A new medium for the axenic cultivation of Entamoeba histolytica and other Entamoeba. Trans R So Trop Med Hyg 72:431–432
Dobell C (1928) Researches on the intestinal protozoa of monkeys and man. Parasitology 20:357–412
Eichinger D (1997) Encystation of Entamoeba parasites. Bioessays 19:633–639
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Frisardi M, Ghosh SK, Field J, Van Dellen K, Rogers R, Robbins P, Samuelson J (2000) The most abundant glycoprotein of amebic cyst walls (Jacob) is a lectin with five cys-rich, chitin-binding domains. Infect Immun 68:4217–4224
Garcia-Zapien AG, Hernandez-Gutierrez R, Mora-Galindo J (1995) Simultaneous growth and mass encystation of Entamoeba invadens under axenic conditions. Arch Med Res 26:257–262
Geiman QM, Ratcliffe HL (1936) Morphology and life-cycle of an amoeba producing amoebiasis in reptiles. Parasitology 28:208–230
Kanehisa M, Goto S (2000) KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30
Keene WE, Petitt MG, Allen S, McKerrow JH (1986) The major neutral proteinase of Entamoeba histolytica. J Exp Med 163:536–549
Kumagai M, Kobayashi S, Okita T, Ohtomo H (2001) Modifications of Kohn's chlorazol black E staining and Wheatley's trichrome staining for temporary wet mount and permanent preparation of Entamoeba histolytica. J Parasitol 87:701–704
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:681–685
León G, Fiori C, Das P, Moreno M, Tovar R, Sánchez-Sala JL, Muñoz ML (1997) Electron probe analysis and biochemical characterization of electron-dense granules secreted by Entamoeba histolytica. Mol Biochem Res 85:233–242
López-Romero E, Villagómez-Castro JC (1993) Encystation in Entamoeba invadens. Parasitol Today 9:225–227
Makioka A, Kumagai M, Ohtomo H, Kobayashi S, Takeuchi T (2002) Effect of proteasome inhibitors on the growth, encystation, and excystation of Entamoeba histolytica and Entamoeba invadens. Parasitol Res 88:454–459
Makioka A, Kumagai M, Kobayashi S, Takeuchi T (2005) Entamoeba invadens: cysteine protease inhibitors block excystation and metacystic development. Exp Parasitol 109:27–32
McConnachie EW (1955) Studies on Entamoeba invadens Rodhain, 1934, in vitro, and its relationship to some other species of Entamoeba. Parasitol 45:452–481
McKerrow JH (1989) Parasite proteases. Exp Parasitol 68:111–115
Page RD (1996) TreeView: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Que X, Reed SL (2000) Cysteine proteinases and the pathogenesis of amebiasis. Clin Microbiol Rev 13:196–206
Rengpien S, Bailey GB (1975) Differentiation of Entamoeba: a new medium and optimal conditions for axenic encystation of E. invadens. J Parasitol 61:24–30
Riahi Y, Ankri S (2000) Involvement of serine proteases during encystation of Entamoeba invadens. Arch Med Res 31:S187–S189
Rosenthal PJ (1999) Proteases of protozoan parasites. Adv Parasitol 43:106–139
Saitou N, Nei M (1987) The neighbor-joining methods: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sajid M, McKerrow JH (2002) Cysteine proteases of parasitic organisms. Mol Biochem Parasitol 120:1–21
Sanchez L, Enea V, Eichinger D (1994) Identification of a developmentally regulated transcript expressed during encystation of Entamoeba invadens. Mol Biochem Parasitol 67:125–135
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tillack M, Biller L, Irmer H, Freitas M, Gomes MA, Tannich E, Bruchhaus I (2007) The Entamoeba histolytica genome: primary structure and expression of proteolytic enzymes. BMC Genomics 8:170–185
Yousef GM, Kopolovic AD, Elliott MB, Diamandis EP (2003) Genomic overview of serine proteases. Biochem Biophys Res Commun 305:28–36
Acknowledgments
We thank N. Watanabe and J. Watanabe for their valuable discussions with us, L. S. Diamond for supplying the E. invadens, K. Hiranuka and T. Katayama for their search of databases, and T. Yamashita and T. Tadano for their technical assistance. This work was supported in part by a Grant-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports and Technology of Japan.
Author information
Authors and Affiliations
Corresponding author
Additional information
An erratum to this article can be found at http://dx.doi.org/10.1007/s00436-009-1564-0
Rights and permissions
About this article
Cite this article
Makioka, A., Kumagai, M., Kobayashi, S. et al. Involvement of serine proteases in the excystation and metacystic development of Entamoeba invadens . Parasitol Res 105, 977–987 (2009). https://doi.org/10.1007/s00436-009-1478-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-009-1478-x