Abstract
Background
Thyroid carcinoma (THCA) is a common type of cancer worldwide, and its incidence has been increasing in recent years. Disulfidptosis, a recently defined form of metabolic-related regulated cell death (RCD), has been shown to play a sophisticated role in antitumor immunity. However, its mechanisms and functions are still poorly understood and the association between disulfidptosis and the prognosis of patients with papillary thyroid carcinoma remains to be elucidated. This study aims to investigate the connection between disulfidptosis and the prognosis of thyroid cancer, while also developing a prognostic index based on disulfidptosis genes.
Materials and methods
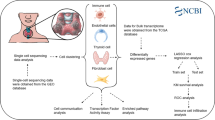
We utilized 24 genes associated with disulfidptosis to create the classification and model. To gather data, we sourced gene expression profiles, somatic mutation information, copy number variation data, and corresponding clinical data from the TCGA database for patients with thyroid cancer. Additionally, we obtained single-cell transcriptome data GSE184362 from the Gene Expression Omnibus (GEO) database for further analysis.
Results
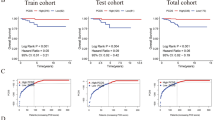
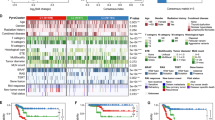
In this study, we utilized 24 genes associated with disulfidptosis to identify two distinct groups with different biological processes using non-negative matrix factorization (NMF). Our findings showed that Cluster 1 is associated with chemokines, interleukins, interferons, checkpoint genes, and other important components of the immune microenvironment. Moreover, cluster 1 patients with high IPS scores may be more sensitive to immunotherapy. We also provide drug therapeutic strategies for each cluster patients based on the IC50 of each drug. The Enet model was chosen as the optimal model with the highest C-index and showed that patients with high risk had a worse prognosis and weak cell-to-cell interactions in THCA. Finally, we established a nomogram model based on multivariable cox and logistic regression analyses to predict the overall survival of THCA patients.
Conclusion
This research provides new insight into the impact of disulfidptosis on THCA. Through a thorough examination of disulfidptosis, a new classification system has been developed that can effectively predict the clinical prognosis and drug sensitivity of THCA patients.







Similar content being viewed by others
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Aran D, Hu Z, Butte AJ (2017) xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome Biol 18:1–14
Aran D et al (2019) Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat Immunol 20(2):163–172
Bagaev A et al (2021) Conserved pan-cancer microenvironment subtypes predict response to immunotherapy. Cancer Cell 39(6):845-865. e7
Bonati L, Tang L (2021) Cytokine engineering for targeted cancer immunotherapy. Curr Opin Chem Biol 62:43–52
Charoentong P et al (2017) Pan-cancer immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep 18(1):248–262
Franses JW et al (2011) Stromal endothelial cells directly influence cancer progression. Sci Transl Med 3(66):66ra5-66ra6
Gros F, Muller S (2023) The role of lysosomes in metabolic and autoimmune diseases. Nat Rev Nephrol 19:1–18
He K et al (2016) mTOR inhibitors induce apoptosis in colon cancer cells via CHOP-dependent DR5 induction on 4E-BP1 dephosphorylation. Oncogene 35(2):148–157
Hoadley KA et al (2018) Cell-of-origin patterns dominate the molecular classification of 10,000 tumors from 33 types of cancer. Cell 173(2):291-304. e6
Hong T et al (2022) Aging-related features predict prognosis and immunotherapy efficacy in hepatocellular carcinoma. Front Immunol 13:951459
Imyanitov EN, Iyevleva AG, Levchenko EV (2021) Molecular testing and targeted therapy for non-small cell lung cancer: current status and perspectives. Crit Rev Oncol Hematol 157:103194
Jin S et al (2021) Inference and analysis of cell-cell communication using cell chat. Nat Commun 12(1):1088
Kent WJ et al (2002) The human genome browser at UCSC. Genome Res 12(6):996–1006
Korotkevich G, Sukhov V, Budin N, Shpak B, Artyomov MN, Sergushichev A (2016) Fast gene set enrichment analysis. BioRxiv 060012. https://doi.org/10.1101/060012
Liberzon A et al (2015) The molecular signatures database hallmark gene set collection. Cell Syst 1(6):417–425
Liu X et al (2023) Actin cytoskeleton vulnerability to disulfide stress mediates disulfidptosis. Nat Cell Biol 25(3):404–414
Lu X et al (2019) Immune signature-based subtypes of cervical squamous cell carcinoma tightly associated with human papillomavirus type 16 expression, molecular features, and clinical outcome. Neoplasia 21(6):591–601
Mantovani A et al (2002) Macrophage polarization: tumor-associated macrophages as a paradigm for polarized M2 mononuclear phagocytes. Trends Immunol 23(11):549–555
Mermel CH et al (2011) GISTIC2. 0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol 12:1–14
Miranda-Filho A et al (2021) Thyroid cancer incidence trends by histology in 25 countries: a population-based study. Lancet Diabetes Endocrinol 9(4):225–234
Pan Y et al (2020) Comprehensive analysis of the association between tumor-infiltrating immune cells and the prognosis of lung adenocarcinoma. J Cancer Res Ther 16(2):320–326
Pereira M et al (2020) Thyroid cancer incidence trends in the United States: association with changes in professional guideline recommendations. Thyroid 30(8):1132–1140
Pozdeyev N et al (2018) Genetic analysis of 779 advanced differentiated and anaplastic thyroid cancers. Clin Cancer Res 24(13):3059–3068
Pu W et al (2021) Single-cell transcriptomic analysis of the tumor ecosystems underlying initiation and progression of papillary thyroid carcinoma. Nat Commun 12(1):6058
Ramakrishnan R, Antonia S, Gabrilovich DI (2008) Combined modality immunotherapy and chemotherapy: a new perspective. Cancer Immunol Immunother 57:1523–1529
Rousselière A et al (2022) Distinctive phenotype for HLA-E-versus HLA-A2-restricted memory CD8 αβT cells in the course of HCMV infection discloses features shared with NKG2C+ CD57+ NK and δ2-γδT cell subsets. Front Immunol 13:1063690
Russo L, Lumeng CN (2018) Properties and functions of adipose tissue macrophages in obesity. Immunology 155(4):407–417
Stuart T et al (2019) Comprehensive integration of single-cell data. Cell 177(7):1888-1902. e21
Wei X et al (2020) Analysis of the role of the interleukins in colon cancer. Biol Res 53(1):1–9
Yang Z et al (2021) A new risk factor indicator for papillary thyroid cancer based on immune infiltration. Cell Death Dis 12(1):51
Yoshihara K et al (2013) Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun 4(1):2612
Zheng P et al (2023) Disulfidptosis: a new target for metabolic cancer therapy. J Exp Clin Cancer Res 42(1):103
Funding
None.
Author information
Authors and Affiliations
Contributions
ZF, JZ: designed this project. QZ: performed the bioinformatics analysis. ZF: wrote the manuscript and supervised the project. QZ, YD, YX, XS, QC, YZ and JM: performed the data review and modified manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethics approval and consent to participate
Not applicable.
Consent for publication
All authors agreed on the manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
432_2023_5006_MOESM1_ESM.pdf
Supplementary file1 Figure S1. Somatic mutation, and copy number variation patterns of different clusters. (A) Waterfall chart showing the somatic mutation frequency and specific mutations of top mutation rate genes in two clusters. (B) The location and CNV level of 24 disulfidptosis related genes in the TCGA-THCA cohort. (C) The amplification and deletion calculated by GISTIC in C1. (D) The amplification and deletion calculated by GISTIC in C2. (E) The GISTIC score of patients in C1. (F) The GISTIC score of patients in C2. (PDF 3088 KB)
432_2023_5006_MOESM2_ESM.pdf
Supplementary file2 Figure S2. (A) Kaplan-Meier of GYS1 and SLC7A11 in THCA patients. (B) Kaplan-Meier of INF2, PDLIM1 and SLC3A2 in THCA patients. (PDF 461 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Feng, Z., Zhao, Q., Ding, Y. et al. Identification a unique disulfidptosis classification regarding prognosis and immune landscapes in thyroid carcinoma and providing therapeutic strategies. J Cancer Res Clin Oncol 149, 11157–11170 (2023). https://doi.org/10.1007/s00432-023-05006-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-05006-4