Abstract
Background
Cirrhosis is a serious condition characterized by the replacement of healthy liver tissue with scar tissue, which can progress to liver failure if left untreated. Hepatocellular carcinoma (HCC) is a concerning complication of cirrhosis. It can be challenge to identify individuals with cirrhosis who are at high risk of developing HCC, particularly in the absence of known risk factors.
Methods
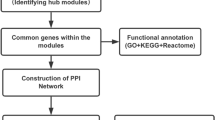
In this study, statistical and bioinformatics methods were utilized to construct a protein–protein interaction network and identify disease-related hub genes. We analyzed two hub genes, CXCL8 and CCNB1, and developed a mathematical model to predict the likelihood of developing HCC in individuals with cirrhosis. We also investigated immune cell infiltration, functional analysis under ontology terms, pathway analysis, distinct clusters of cells, and protein–drug interactions.
Results
The results indicated that CXCL8 and CCNB1 were associated with the development of cirrhosis-induced HCC. A prognostic model based on these two genes was able to predict the occurrence and survival time of HCC. In addition, the candidate drugs were also discovered based on our model.
Conclusion
The findings offer the potential for earlier detection of cirrhosis-induced HCC and provide a new instrument for clinical diagnosis, prognostication, and the development of immunological medications. This study also identified distinct clusters of cells in HCC patients using UMAP plot analysis and analyzed the expression of CXCL8 and CCNB1 within these cells, indicating potential therapeutic opportunities for targeted drug therapies to benefit HCC patients.












Similar content being viewed by others
Data availability
The datasets analysed during the current study are available from the corresponding author on reasonable request.
References
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM et al (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25(1):25–29
Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M et al (2013) NCBI GEO: archive for functional genomics data sets-update. Nucleic Acids Res 41(Database issue):D991–D995
Bidkhori G, Benfeitas R, Klevstig M, Zhang C, Nielsen J, Uhlen M et al (2018) Metabolic network-based stratification of hepatocellular carcinoma reveals three distinct tumor subtypes. Proc Natl Acad Sci USA 115(50):E11874–E11883
Blake JA, Christie KR, Dolan ME, Drabkin HJ, Hill DP, Ni L, Sitnikov D, Burgess S, Buza T, Gresham C, McCarthy F, Pillai L, Wang H, Carbon S, Dietze H, Lewis SE, Mungall CJ, Munoz-Torres MC, Feuermann M, Gaudet P, Basu S, Chisholm RL, Dodson RJ, Fey P, Mi H, Thomas PD, Muruganujan A, Poudel S, Hu JC, Aleksander SA, McIntosh BK, Renfro DP, Siegele DA, Attrill H, Brown NH, Tweedie S, Lomax J, Osumi-Sutherland D, Parkinson H, Roncaglia P, Lovering RC, Talmud PJ, Humphries SE, Denny P, Campbell NH, Foulger RE, Chibucos MC, Giglio MG, Chang HY, Finn R, Fraser M, Mitchell A, Nuka G, Pesseat S, Sangrador A, Scheremetjew M, Young SY, Stephan R, Harris MA, Oliver SG, Rutherford K, Wood V, Bahler J, Lock A, Kersey PJ, McDowall MD, Staines DM, Dwinell M, Shimoyama M, Laulederkind S, Hayman GT, Wang SJ, Petri V, Deustachio P, Matthews L, Balakrishnan R, Binkley G, Cherry JM, Costanzo MC, Demeter J, Dwight SS, Engel SR, Hitz BC, Inglis DO, Lloyd P, Miyasato SR, Paskov K, Roe G, Simison M, Nash RS, Skrzypek MS, Weng S, Wong ED, Berardini TZ, Li D, Huala E, Argasinska J, Arighi C, Auchincloss A, Axelsen K, Argoud-Puy G, Bateman A, Bely B, Blatter MC, Bonilla C, Bougueleret L, Boutet E, Breuza L, Bridge A, Britto R, Casals C, Cibrian-Uhalte E, Coudert E, Cusin I, Duek-Roggli P, Estreicher A, Famiglietti L, Gane P, Garmiri P, Gos A, Gruaz-Gumowski N, Hatton-Ellis E, Hinz U, Hulo C, Huntley R, Jungo F, Keller G, Laiho K, Lemercier P, Lieberherr D, MacDougall A, Magrane M, Martin M, Masson P, Mutowo P, O'Donovan C, Pedruzzi I, Pichler K, Poggioli D, Poux S, Rivoire C, Roechert B, Sawford T, Schneider M, Shypitsyna A, Stutz A, Sundaram S, Tognolli M, Wu C, Xenarios I, Chan J, Kishore R, Sternberg PW, Van Auken K, Muller HM, Done J, Li Y, Howe D, Westerfield M (2015) Gene Ontology Consortium: going forward. Nucleic Acids Res 43(Database issue):D1049–D1056
Chen H, Boutros PC (2011) VennDiagram: a package for the generation of highly-customizable Venn and Euler diagrams in R. BMC Bioinform 12:35
Chin CH, Chen SH, Wu HH, Ho CW, Ko MT, Lin CY (2014) cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol 8(Suppl 4):S11
Cox DR (1972) Regression models and life-tables. J R Stat Soc Ser B (methodol) 34(2):187–202
Fernando RI, Castillo MD, Litzinger M, Hamilton DH, Palena C (2011) IL-8 signaling plays a critical role in the epithelial–mesenchymal transition of human carcinoma cells IL-8 signaling in brachyury-induced tumor progression. Can Res 71(15):5296–5306
Forner A, Llovet JM, Bruix J (2012) Hepatocellular carcinoma. Lancet (london, England) 379(9822):1245–1255
Forner A, Reig M, Bruix J (2018) Hepatocellular carcinoma. Lancet (london, England) 391(10127):1301–1314
Fu T, Dai LJ, Wu SY, Xiao Y, Ma D, Jiang YZ et al (2021) Spatial architecture of the immune microenvironment orchestrates tumor immunity and therapeutic response. J Hematol Oncol 14(1):98
Gajewski TF, Schreiber H, Fu YX (2013) Innate and adaptive immune cells in the tumor microenvironment. Nat Immunol 14(10):1014–1022
Giannelli G, Koudelkova P, Dituri F, Mikulits W (2016) Role of epithelial to mesenchymal transition in hepatocellular carcinoma. J Hepatol 65(4):798–808
Hammond ME, Lapointe GR, Feucht PH, Hilt S, Gallegos CA, Gordon CA et al (1995) IL-8 induces neutrophil chemotaxis predominantly via type I IL-8 receptors. J Immunol (baltimore, Md: 1950). 155(3):1428–1433
Han Y, Wang Y, Dong X, Sun D, Liu Z, Yue J et al (2023) TISCH2: expanded datasets and new tools for single-cell transcriptome analyses of the tumor microenvironment. Nucleic Acids Res 51(D1):D1425–D1431
Iasonos A, Schrag D, Raj GV, Panageas KS (2008) How to build and interpret a nomogram for cancer prognosis. J Clin Oncol off J Am Soc Clin Oncol 26(8):1364–1370
Ito K, Murphy D (2013) Application of ggplot2 to pharmacometric graphics. CPT Pharmacomet Syst Pharmacol 2(10):e79
Kamarudin AN, Cox T, Kolamunnage-Dona R (2017) Time-dependent ROC curve analysis in medical research: current methods and applications. BMC Med Res Methodol 17(1):53
La Vecchia C, Negri E, Cavalieri d’Oro L, Franceschi S (1998) Liver cirrhosis and the risk of primary liver cancer. Eur J Cancer Prev off J Eur Cancer Prev Organ (ECP) 7(4):315–320
Lamouille S, Xu J, Derynck R (2014) Molecular mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell Biol 15(3):178–196
Lánczky A, Győrffy B (2021) Web-based survival analysis tool tailored for medical research (KMplot): development and implementation. J Med Internet Res 23(7):e27633
Lei X, Lei Y, Li JK, Du WX, Li RG, Yang J et al (2020) Immune cells within the tumor microenvironment: Biological functions and roles in cancer immunotherapy. Cancer Lett 470:126–133
Li X, Huang L, Leng X (2018) Analysis of prognostic factors of more/equal to10 years of survival for liver cancer patients after liver transplantation. J Cancer Res Clin Oncol 144(12):2465–2474
Li B, Cheng J, Wang H, Zhao S, Zhu H, Li C et al (2019) CCNB1 affects cavernous sinus invasion in pituitary adenomas through the epithelial-mesenchymal transition. J Transl Med 17(1):336
Maeser D, Gruener RF, Huang RS (2021) oncoPredict: an R package for predicting in vivo or cancer patient drug response and biomarkers from cell line screening data. Brief Bioinform. https://doi.org/10.1093/bib/bbab260
Malumbres M, Barbacid M (2005) Mammalian cyclin-dependent kinases. Trends Biochem Sci 30(11):630–641
Marrero JA (2006) Hepatocellular carcinoma. Curr Opin Gastroenterol 22(3):248–253
Maxwell PJ, Gallagher R, Seaton A, Wilson C, Scullin P, Pettigrew J et al (2007) HIF-1 and NF-kappaB-mediated upregulation of CXCR1 and CXCR2 expression promotes cell survival in hypoxic prostate cancer cells. Oncogene 26(52):7333–7345
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y et al (2015) Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 12(5):453–457
Pardoll DM (2012) The blockade of immune checkpoints in cancer immunotherapy. Nat Rev Cancer 12(4):252–264
Powell DR, Huttenlocher A (2016) Neutrophils in the tumor microenvironment. Trends Immunol 37(1):41–52
Qian BZ, Pollard JW (2010) Macrophage diversity enhances tumor progression and metastasis. Cell 141(1):39–51
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W et al (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43(7):e47
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504
Shen H, Wu H, Sun F, Qi J, Zhu Q (2021) A novel four-gene of iron metabolism-related and methylated for prognosis prediction of hepatocellular carcinoma. Bioengineered 12(1):240–251
Si T, Huang Z, Jiang Y, Walker-Jacobs A, Gill S, Hegarty R et al (2021) Expression levels of three key genes CCNB1, CDC20, and CENPF in HCC are associated with antitumor immunity. Front Oncol 11:738841
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102(43):15545–15550
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z (2017) GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res 45(W1):W98-w102
Van Calster B, Wynants L, Verbeek JFM, Verbakel JY, Christodoulou E, Vickers AJ et al (2018) Reporting and interpreting decision curve analysis: a guide for investigators. Eur Urol 74(6):796–804
Van Calster B, McLernon DJ, van Smeden M, Wynants L, Steyerberg EW (2019) Calibration: the Achilles heel of predictive analytics. BMC Med 17(1):230
Villanueva A (2019) Hepatocellular carcinoma. N Engl J Med 380(15):1450–1462
Waugh DJ, Wilson C (2008) The interleukin-8 pathway in cancer. Clin Cancer Res off J Am Assoc Cancer Res 14(21):6735–6741
Wilkerson MD, Hayes DN (2010) ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics (oxford, England) 26(12):1572–1573
Yu G, Wang LG, Han Y, He QY (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16(5):284–287
Zhang EL, Liang BY, Chen XP, Huang ZY (2015) Severity of liver cirrhosis: a key role in the selection of surgical modality for Child-Pugh A hepatocellular carcinoma. World J Surg Oncol 13:148
Zinkernagel AS, Timmer AM, Pence MA, Locke JB, Buchanan JT, Turner CE et al (2008) The IL-8 protease SpyCEP/ScpC of group A Streptococcus promotes resistance to neutrophil killing. Cell Host Microbe 4(2):170–178
Acknowledgement
We would like to express our gratitude to the patients who generously donated their tissue samples and our sincere appreciation to the GEO database for providing the valuable data used in this study. We would also like to thank Qingyuan Sun’s parents for their unwavering support and encouragement throughout the duration of this project.
Funding
This work is supported by Young Scientists Fund of the National Natural Science Foundation of China (82201383), Project funded by China Postdoctoral Science Foundation (2022M711319), and Natural Science Foundation of Shandong Province China (ZR2021MH037).
Author information
Authors and Affiliations
Contributions
Conceptualization, YW; data curation, QS; funding acquisition, CW and YW; investigation, RA and MW; methodology, QS and RA; project administration, YW; resources, JL, CL, and YW; software, QS; supervision, JL, CL, and YW; validation, RA; visualization, QS, JL, and YW; writing—original draft, QS; writing—review & editing, CW and YW.
Corresponding authors
Ethics declarations
Conflict of interest
We have no potential conflicts of interest to disclose.
Institutional review board statement
Not applicable.
Informed consent
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sun, Q., An, R., Li, J. et al. The role of CXCL8 and CCNB1 in predicting hepatocellular carcinoma in the context of cirrhosis: implications for early detection and immune-based therapies. J Cancer Res Clin Oncol 149, 11471–11489 (2023). https://doi.org/10.1007/s00432-023-05004-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-05004-6