Abstract
Background
COVID-19 has created a significant risk to worldwide public health. According to recent research, C-type lectins may be SARS-CoV-2 receptors. Layilin (LAYN), a broadly expressed integral membrane hyaluronan receptor with a C-type lectin structural domain, is a gene related to cell senescence. There are a few studies on C-type lectins in pan-cancer, and no pan-cancer analysis has been conducted for LAYN.
Methods
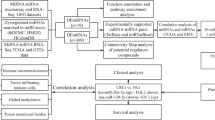
The genotype tissue expression (GTEx) portal and the cancer genome map (TCGA) database were used to collect samples from healthy and cancer patients. Bioinformatics methods are used to construct immune landscape, mutation landscape, and stemness landscape of LAYN. The single-cell sequencing data were used from the CancerSEA website to analyze the functions of LAYN. The prognosis potential of LAYN was discussed based on machine learning.
Results
LAYN is differentially expressed among cancers. Survival analysis indicated that LAYN was related to a poor overall survival (OS) rate in cancers, like HNSC, MESO, and OV. Mutational landscapes of LAYN in SKCM and STAD were constructed. LAYN was negatively related to Tumor Mutation Burden (TMB) in THCA, PRAD, and UCEC, and with the Microsatellite Instability (MSI) in STAD, LUAD, and UCEC. The immune landscape in pan-cancer suggested that LAYN may be involved in tumor immune escape. LAYN plays a crucial role in the infiltration of immune cells in malignant tumors. LAYN participates in methylation modifications and affects tumor proliferation and metastasis by regulating stemness. Analysis of single-cell sequencing data suggests that LAYN may participate in several biological processes, like stemness, apoptosis, and DNA repair. LAYN transcript was predicted as a liquid–liquid phase separation (LLPS)-related RNA. The results of KIRC were verified in the GEO and ArrayExpress databases. Furthermore, prognostic models based on machine learning of LAYN-related genes were established. Hsa-miR-153-5p and hsa-miR-505-3p may be the upstream miRNAs of LAYN and have a high value for tumor prognosis.
Conclusion
This study elucidated the functional mechanisms of LAYN from a pan-cancer perspective and provided novel insights into cancer prognosis, metastasis, and immunotherapy. LAYN has the potential to become a new target of mRNA vaccines and molecular therapies in tumors.















Similar content being viewed by others
Data availability
The datasets presented in this study can be found in online repositories. The names of the repository/repositories can be found in the article/supplementary material.
References
Adachi T, Arito M, Suematsu N et al (2015) Roles of layilin in TNF-α-induced epithelial-mesenchymal transformation of renal tubular epithelial cells. Biochem Biophys Res Commun 467(1):63–69. https://doi.org/10.1016/j.bbrc.2015.09.121
Balta E, Wabnitz GH, Samstag Y (2021) Hijacked immune cells in the tumor microenvironment: molecular mechanisms of immunosuppression and cues to improve T cell-based immunotherapy of solid tumors. Int J Mol Sci 22(11):5736. https://doi.org/10.3390/ijms22115736. (Published 2021 May 27)
Bono P, Rubin K, Higgins JM et al (2001) Layilin, a novel integral membrane protein, is a hyaluronan receptor. Mol Biol Cell 12(4):891–900. https://doi.org/10.1091/mbc.12.4.891
Chen Y, Wang X (2020) miRDB: an online database for prediction of functional microRNA targets. Nucleic Acids Res 48(D1):D127–D131. https://doi.org/10.1093/nar/gkz757
Chen Z, Zhuo W, Wang Y, Ao X, An J (2008) Down-regulation of layilin, a novel hyaluronan receptor, via RNA interference, inhibits invasion and lymphatic metastasis of human lung A549 cells. Biotechnol Appl Biochem 50(Pt 2):89–96. https://doi.org/10.1042/BA20070138
Chen L, Heikkinen L, Wang C, Yang Y, Sun H, Wong G (2019) Trends in the development of miRNA bioinformatics tools. Brief Bioinform 20(5):1836–1852. https://doi.org/10.1093/bib/bby054
Chen P, Hsu WH, Han J, Xia Y, DePinho RA (2021) Cancer stemness meets immunity: from mechanism to therapy. Cell Rep 34(1):108597. https://doi.org/10.1016/j.celrep.2020.108597
Cortes-Ciriano I, Lee S, Park WY, Kim TM, Park PJ (2017) A molecular portrait of microsatellite instability across multiple cancers. Nat Commun 8:15180. https://doi.org/10.1038/ncomms15180. (Published 2017 Jun 6)
Drake CG, Jaffee E, Pardoll DM (2006) Mechanisms of immune evasion by tumors. Adv Immunol 90:51–81. https://doi.org/10.1016/S0065-2776(06)90002-9
Gao J, Aksoy BA, Dogrusoz U et al (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6(269):pl1. https://doi.org/10.1126/scisignal.2004088. (Published 2013 Apr 2)
GTEx Consortium (2013) The genotype-tissue expression (GTEx) project. Nat Genet 45(6):580–585. https://doi.org/10.1038/ng.2653
Haruehanroengra P, Zheng YY, Zhou Y, Huang Y, Sheng J (2020) RNA modifications and cancer. RNA Biol 17(11):1560–1575. https://doi.org/10.1080/15476286.2020.1722449
Jhunjhunwala S, Hammer C, Delamarre L (2021) Antigen presentation in cancer: insights into tumour immunogenicity and immune evasion. Nat Rev Cancer 21(5):298–312. https://doi.org/10.1038/s41568-021-00339-z
Kaji T, Arito M, Tsutiya A et al (2019) Layilin enhances the invasive ability of malignant glioma cells via SNAI1 signaling. Brain Res 1719:140–147. https://doi.org/10.1016/j.brainres.2019.05.034
Kobayashi H, Looker HC, Satake E et al (2022) Results of untargeted analysis using the SOMAscan proteomics platform indicates novel associations of circulating proteins with risk of progression to kidney failure in diabetes. Kidney Int 102(2):370–381. https://doi.org/10.1016/j.kint.2022.04.022
Li T, Fu J, Zeng Z et al (2020) TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res 48(W1):W509–W514. https://doi.org/10.1093/nar/gkaa407
Li Z, Zhao S, Zhu S, Fan Y (2021) MicroRNA-153-5p promotes the proliferation and metastasis of renal cell carcinoma via direct targeting of AGO1. Cell Death Dis 12(1):33. https://doi.org/10.1038/s41419-020-03306-y. (Published 2021 Jan 4)
Liu M, Li H, Luo X et al (2022) RPS: a comprehensive database of RNAs involved in liquid-liquid phase separation. Nucleic Acids Res 50(D1):D347–D355. https://doi.org/10.1093/nar/gkab986
Ma S, Chen C, Ji X et al (2019) The interplay between m6A RNA methylation and noncoding RNA in cancer. J Hematol Oncol 12(1):121. https://doi.org/10.1186/s13045-019-0805-7. (Published 2019 Nov 22)
Ma Y, Gu Y, Zhang X et al (2022) High expression of MUC5AC, MUC5B, and layilin plays an essential role in prediction in the development of plastic bronchitis caused by MPP. Front Microbiol 13:911228. https://doi.org/10.3389/fmicb.2022.911228. (Published 2022 Jun 13)
Mahuron KM, Moreau JM, Glasgow JE et al (2020) Layilin augments integrin activation to promote antitumor immunity. J Exp Med 217(9):e20192080. https://doi.org/10.1084/jem.20192080
Malta TM, Sokolov A, Gentles AJ et al (2018) Machine learning identifies stemness features associated with oncogenic dedifferentiation. Cell 173(2):338-354.e15. https://doi.org/10.1016/j.cell.2018.03.034
Murciano-Goroff YR, Warner AB, Wolchok JD (2020) The future of cancer immunotherapy: microenvironment-targeting combinations. Cell Res 30(6):507–519. https://doi.org/10.1038/s41422-020-0337-2
Navarro Gonzalez J, Zweig AS, Speir ML et al (2021) The UCSC genome browser database: 2021 update. Nucleic Acids Res 49(D1):D1046–D1057. https://doi.org/10.1093/nar/gkaa1070
Pan JH, Zhou H, Cooper L et al (2019) LAYN Is a prognostic biomarker and correlated with immune infiltrates in gastric and colon cancers. Front Immunol 10:6. https://doi.org/10.3389/fimmu.2019.00006. (Published 2019 Jan 29)
Papanicolau-Sengos A, Aldape K (2022) DNA methylation profiling: an emerging paradigm for cancer diagnosis. Annu Rev Pathol 17:295–321. https://doi.org/10.1146/annurev-pathol-042220-022304
Peng PH, Hsu KW, Wu KJ (2021) Liquid-liquid phase separation (LLPS) in cellular physiology and tumor biology. Am J Cancer Res 11(8):3766–3776 (Published 2021 Aug 15)
Romano S, Tufano M, D’Arrigo P et al (2020) Cell stemness, epithelial-to-mesenchymal transition, and immunoevasion: intertwined aspects in cancer metastasis. Semin Cancer Biol 60:181–190. https://doi.org/10.1016/j.semcancer.2019.08.015
Roma-Rodrigues C, Mendes R, Baptista PV et al (2019) Targeting tumor microenvironment for cancer therapy. Int J Mol Sci 20(4):840. https://doi.org/10.3390/ijms20040840. (Published 2019 Feb 15)
Rovillain E, Mansfield L, Lord CJ et al (2011) An RNA interference screen for identifying downstream effectors of the p53 and pRB tumour suppressor pathways involved in senescence. BMC Genomics 12:355. https://doi.org/10.1186/1471-2164-12-355. (Published 2011 Jul 8)
Samstein RM, Lee CH, Shoushtari AN et al (2019) Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat Genet 51(2):202–206. https://doi.org/10.1038/s41588-018-0312-8
Siegel RL, Miller KD, Fuchs HE, Jemal A (2022) Cancer statistics, 2022. CA Cancer J Clin 72(1):7–33. https://doi.org/10.3322/caac.21708
Song P, Tayier S, Cai Z, Jia G (2021) RNA methylation in mammalian development and cancer. Cell Biol Toxicol 37(6):811–831. https://doi.org/10.1007/s10565-021-09627-8
Szklarczyk D, Gable AL, Nastou KC et al (2021) The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets [published correction appears in Nucleic Acids Res. 2021 Oct 11;49(18):10800]. Nucleic Acids Res 49(D1):D605–D612. https://doi.org/10.1093/nar/gkaa1074
Tang Z, Kang B, Li C, Chen T, Zhang Z (2019a) GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res 47(W1):W556–W560. https://doi.org/10.1093/nar/gkz430
Tang Y, Wu B, Huang S et al (2019b) Downregulation of miR-505-3p predicts poor bone metastasis-free survival in prostate cancer. Oncol Rep 41(1):57–66. https://doi.org/10.3892/or.2018.6826
(2018) The TCGA Legacy. Cell. 173(2):281-282. doi: https://doi.org/10.1016/j.cell.2018.03.049
Uhlen M, Zhang C, Lee S et al (2017) A pathology atlas of the human cancer transcriptome. Science 357(6352):eaan2507. https://doi.org/10.1126/science.aan2507
Wang Y, Liu S, Zhang Y, Yang J (2019) Myosin heavy chain 9: oncogene or tumor suppressor gene? Med Sci Monit 25:888–892. https://doi.org/10.12659/MSM.912320. (Published 2019 Jan 31)
Wang Y, Wu N, Zhang J, Wang H, Men X (2020) MiR-153-5p enhances the sensitivity of triple-negative breast cancer cells to paclitaxel by inducing G2M phase arrest. Onco Targets Ther 13:4089–4097. https://doi.org/10.2147/OTT.S241640. (Published 2020 May 12)
Wegener KL, Basran J, Bagshaw CR et al (2008) Structural basis for the interaction between the cytoplasmic domain of the hyaluronate receptor layilin and the talin F3 subdomain. J Mol Biol 382(1):112–126. https://doi.org/10.1016/j.jmb.2008.06.087
Weng L, Smits P, Wauters J, Merregaert J (2002) Molecular cloning and characterization of human chondrolectin, a novel type I transmembrane protein homologous to C-type lectins. Genomics 80(1):62–70. https://doi.org/10.1006/geno.2002.6806
Xia C, Dong X, Li H et al (2022) Cancer statistics in China and United States, 2022: profiles, trends, and determinants. Chin Med J (Engl) 135(5):584–590. https://doi.org/10.1097/CM9.0000000000002108. (Published 2022 Feb 9)
Yang W, Soares J, Greninger P et al (2013) Genomics of drug sensitivity in cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res 41(Database issue):D955–D961. https://doi.org/10.1093/nar/gks1111
Yang G, Zheng RY, Jin ZS (2019) Correlations between microsatellite instability and the biological behaviour of tumours. J Cancer Res Clin Oncol 145(12):2891–2899. https://doi.org/10.1007/s00432-019-03053-4
Yang B, Wang JQ, Tan Y, Yuan R, Chen ZS, Zou C (2021) RNA methylation and cancer treatment. Pharmacol Res 174:105937. https://doi.org/10.1016/j.phrs.2021.105937
Yuan H, Yan M, Zhang G et al (2019) CancerSEA: a cancer single-cell state atlas. Nucleic Acids Res 47(D1):D900–D908. https://doi.org/10.1093/nar/gky939
Zhao P, Guan H, Dai Z, Ma Y, Zhao Y, Liu D (2019) Long noncoding RNA DLX6-AS1 promotes breast cancer progression via miR-505-3p/RUNX2 axis. Eur J Pharmacol 865:172778. https://doi.org/10.1016/j.ejphar.2019.172778
Acknowledgements
Not applicable.
Funding
This study was supported by the National Key Research and Development Program of China (grant number: 2018YFC2002202).
Author information
Authors and Affiliations
Contributions
WJ designed the work. WJ and WJ analyzed the data and prepared the manuscript. WJ, LJ, and ZY edited and revised the manuscript. All the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jiawen, W., Jinfu, W., Jianyong, L. et al. Comprehensive landscape of the miRNA-regulated prognostic marker LAYN with immune infiltration and stemness in pan-cancer. J Cancer Res Clin Oncol 149, 10989–11011 (2023). https://doi.org/10.1007/s00432-023-04986-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-04986-7