Abstract
Purpose
The potential role of epithelium-specific genes through the adenoma-carcinoma sequence in the development of colorectal cancer (CRC) remains unknown. Therefore, we integrated single-cell RNA sequencing and bulk RNA sequencing data to select diagnosis and prognosis biomarkers for CRC.
Methods
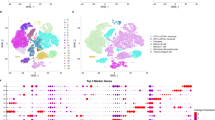
The CRC scRNA-seq dataset was used to describe the cellular landscape of normal intestinal mucosa, adenoma and CRC and to further select epithelium-specific clusters. Differentially expressed genes (DEGs) of epithelium-specific clusters were identified between intestinal lesion and normal mucosa in the scRNA-seq data throughout the adenoma-carcinoma sequence. Diagnostic biomarkers and prognostic biomarker (the risk score) for CRC were selected in the bulk RNA-seq dataset based on DEGs shared by the adenoma epithelium-specific cluster and the CRC epithelium-specific cluster (shared-DEGs).
Results
Among the 1063 shared-DEGs, we selected 38 gene expression biomarkers and 3 methylation biomarkers that had promising diagnostic power in plasma. Multivariate Cox regression identified 174 shared-DEGs as prognostic genes for CRC. We combined 1000 times LASSO-Cox regression and two-way stepwise regression to select 10 prognostic shared-DEGs to construct the risk score in the CRC meta-dataset. In the external validation dataset, the 1- and 5-year AUCs of the risk score were higher than those of stage, the pyroptosis-related genes (PRG) score and the cuproptosis-related genes (CRG) score. In addition, the risk score was closely associated with the immune infiltration of CRC.
Conclusion
The combined analysis of the scRNA-seq dataset and the bulk RNA-seq dataset in this study provides reliable biomarkers for the diagnosis and prognosis of CRC.






Similar content being viewed by others
Data availability
Data for this study can be found in The Cancer Genome Atlas (TCGA, https://portal.gdc.cancer.gov/) and the Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo/). The names of the repository/repositories and accession number(s) can be found in the article/supplementary material.
References
Ammendola M, Sacco R, Sammarco G, Donato G, Montemurro S, Ruggieri E, Patruno R, Marech I, Cariello M, Vacca A, Gadaleta CD, Ranieri G (2014) Correlation between serum tryptase, mast cells positive to tryptase and microvascular density in colo-rectal cancer patients: possible biological-clinical significance. PLoS One 9(6):e99512
Andersen GB, Tost J (2018) A Summary of the biological processes, disease-associated changes, and clinical applications of DNA methylation. Methods Mol Biol 1708:3–30
Anghel SA, Ionita-Mindrican CB, Luca I, Pop AL (2021) Promising epigenetic biomarkers for the early detection of colorectal cancer: a systematic review. Cancers (basel) 13(19):4965
Aristin Revilla S, Kranenburg O, Coffer PJ (2022) Colorectal cancer-infiltrating regulatory T cells: functional heterogeneity, metabolic adaptation, and therapeutic targeting. Front Immunol 13:903564
Ashton TM, McKenna WG, Kunz-Schughart LA, Higgins GS (2018) Oxidative phosphorylation as an emerging target in cancer therapy. Clin Cancer Res 24(11):2482–2490
Becker WR, Nevins SA, Chen DC, Chiu R, Horning AM, Guha TK, Laquindanum R, Mills M, Chaib H, Ladabaum U, Longacre T, Shen J, Esplin ED, Kundaje A, Ford JM, Curtis C, Snyder MP, Greenleaf WJ (2022) Single-cell analyses define a continuum of cell state and composition changes in the malignant transformation of polyps to colorectal cancer. Nat Genet 54(7):985–995
Bindea G, Mlecnik B, Tosolini M, Kirilovsky A, Waldner M, Obenauf AC, Angell H, Fredriksen T, Lafontaine L, Berger A, Bruneval P, Fridman WH, Becker C, Pages F, Speicher MR, Trajanoski Z, Galon J (2013) Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity 39(4):782–795
Canellas-Socias A, Cortina C, Hernando-Momblona X, Palomo-Ponce S, Mulholland EJ, Turon G, Mateo L, Conti S, Roman O, Sevillano M, Slebe F, Stork D, Caballe-Mestres A, Berenguer-Llergo A, Alvarez-Varela A, Fenderico N, Novellasdemunt L, Jimenez-Gracia L, Sipka T, Bardia L, Lorden P, Colombelli J, Heyn H, Trepat X, Tejpar S, Sancho E, Tauriello DVF, Leedham S, Attolini CS, Batlle E (2022) Metastatic recurrence in colorectal cancer arises from residual Emp1(+) cells. Nature 611(7936):603–613
Chen B, Scurrah CR, McKinley ET, Simmons AJ, Ramirez-Solano MA, Zhu X, Markham NO, Heiser CN, Vega PN, Rolong A, Kim H, Sheng Q, Drewes JL, Zhou Y, Southard-Smith AN, Xu Y, Ro J, Jones AL, Revetta F, Berry LD, Niitsu H, Islam M, Pelka K, Hofree M, Chen JH, Sarkizova S, Ng K, Giannakis M, Boland GM, Aguirre AJ, Anderson AC, Rozenblatt-Rosen O, Regev A, Hacohen N, Kawasaki K, Sato T, Goettel JA, Grady WM, Zheng W, Washington MK, Cai Q, Sears CL, Goldenring JR, Franklin JL, Su T, Huh WJ, Vandekar S, Roland JT, Liu Q, Coffey RJ, Shrubsole MJ, Lau KS (2021) Differential pre-malignant programs and microenvironment chart distinct paths to malignancy in human colorectal polyps. Cell 184(26):6262–6280 (e26)
Chen Z, Mei K, Xiao Y, Xiong Y, Long W, Wang Q, Zhong J, Di D, Ge Y, Luo Y, Li Z, Huang Y, Gu R, Wang B (2022) Prognostic assessment of oxidative stress-related genes in colorectal cancer and new insights into tumor immunity. Oxid Med Cell Longev 2022:2518340
Cheng KJ, Alshawsh MA, Mohamed EHM, Thavagnanam S, Sinniah A, Ibrahim ZA (2020) Hmgb1: an overview of its versatile roles in the pathogenesis of colorectal cancer. Cell Oncol (dordr) 43(2):177–193
Cui J, Guo F, Yu Y, Ma Z, Hong Y, Su J, Ge Y (2022) Development and validation of a prognostic 9-gene signature for colorectal cancer. Front Oncol 12:1009698
Di Z, Zhou S, Xu G, Ren L, Li C, Ding Z, Huang K, Liang L, Yuan Y (2022) Single-cell and Wgcna uncover a prognostic model and potential oncogenes in colorectal cancer. Biol Proced Online 24(1):13
Di Nicolantonio F, Vitiello PP, Marsoni S, Siena S, Tabernero J, Trusolino L, Bernards R, Bardelli A (2021) Precision oncology in metastatic colorectal cancer—from biology to medicine. Nat Rev Clin Oncol 18(8):506–525
Dienstmann R, Vermeulen L, Guinney J, Kopetz S, Tejpar S, Tabernero J (2017) Consensus molecular subtypes and the evolution of precision medicine in colorectal cancer. Nat Rev Cancer 17(4):268
Fang Y, Zhan X (2023) Identification of biomarkers associated with the prognoses of colorectal cancer patients. Digestion 104(2):148–162
Fridman WH, Meylan M, Petitprez F, Sun CM, Italiano A, Sautes-Fridman C (2022) B cells and tertiary lymphoid structures as determinants of tumour immune contexture and clinical outcome. Nat Rev Clin Oncol 19(7):441–457
Governa V, Trella E, Mele V, Tornillo L, Amicarella F, Cremonesi E, Muraro MG, Xu H, Droeser R, Daster SR, Bolli M, Rosso R, Oertli D, Eppenberger-Castori S, Terracciano LM, Iezzi G, Spagnoli GC (2017) The interplay between neutrophils and Cd8(+) T cells improves survival in human colorectal cancer. Clin Cancer Res 23(14):3847–3858
Han L, Shi J, Zhao L, Deng J, Li Y, Zhao H, Wang H, Yan Y, Zou F (2022) Bcap31 is involved in modulating colorectal cancer cell proliferation via the Emerin/beta-catenin axis. Exp Cell Res 418(1):113265
Hinshaw DC, Shevde LA (2019) The tumor microenvironment innately modulates cancer progression. Cancer Res 79(18):4557–4566
Huang Q, Chen Z, Cheng P, Jiang Z, Wang Z, Huang Y, Yang C, Pan J, Qiu F, Huang J (2019) Lyrm2 directly regulates complex I activity to support tumor growth in colorectal cancer by oxidative phosphorylation. Cancer Lett 455:36–47
Huang X, Ke K, Jin W, Zhu Q, Zhu Q, Mei R, Zhang R, Yu S, Shou L, Sun X, Feng J, Duan T, Mou Y, Xie T, Wu Q, Sui X (2022a) Identification of genes related to 5-fluorouracil based chemotherapy for colorectal cancer. Front Immunol 13:887048
Huang H, Long Z, Xie Y, Qin P, Kuang L, Li X, Zhao Y, Zhang X, Yang L, Ma W, Xiao X, Liu Y, Sun X, Zhang M, Wang F, Hu D, Hu F (2022b) Molecular subtypes based on cuproptosis-related genes and tumor microenvironment infiltration characterization in colorectal cancer. J Oncol 2022:5034092
Jakubowska K, Koda M, Grudzinska M, Kisielewski W, Lomperta K, Famulski W (2022) Neutrophil infiltration combined with necrosis in the primary tumor is a useful prognostic indicator for three-year disease-free survival time in patients with colorectal cancer. Oncol Lett 23(6):199
Kaldma A, Klepinin A, Chekulayev V, Mado K, Shevchuk I, Timohhina N, Tepp K, Kandashvili M, Varikmaa M, Koit A, Planken M, Heck K, Truu L, Planken A, Valvere V, Rebane E, Kaambre T (2014) An in situ study of bioenergetic properties of human colorectal cancer: the regulation of mitochondrial respiration and distribution of flux control among the components of Atp synthasome. Int J Biochem Cell Biol 55:171–186
Koppad S, Basava A, Nash K, Gkoutos GV, Acharjee A (2022) Machine learning-based identification of colon cancer candidate diagnostics genes. Biology (basel) 11(3):365
Li Z, Wang H, Song B, Sun Y, Han J, Xu Z (2015) Expression of high mobility group box-1 in colorectal cancer and its clinical significance. Zhonghua Wei Chang Wai Ke Za Zhi 18(6):616–619
Lin WR, Chiang JM, Lim SN, Su MY, Chen TH, Huang SW, Chen CW, Wu RC, Tsai CL, Lin YH, Alison MR, Hsieh SY, Yu JS, Chiu CT, Yeh CT (2019) Dynamic bioenergetic alterations in colorectal adenomatous polyps and adenocarcinomas. EBioMedicine 44:334–345
Liu W, Liu Y, Zhu J, Wright E, Ding I, Rodgers GP (2008) Reduced Hgc-1 protein expression is associated with malignant progression of colon carcinoma. Clin Cancer Res 14(4):1041–1049
Liu XS, Yang JW, Zeng J, Chen XQ, Gao Y, Kui XY, Liu XY, Zhang Y, Zhang YH, Pei ZJ (2022) Slc2a1 is a diagnostic biomarker involved in immune infiltration of colorectal cancer and associated with M6a modification and cerna. Front Cell Dev Biol 10:853596
Lu H, Zhu M, Qu L, Shao H, Zhang R, Li Y (2022) Oncogenic role of Hmgb1 as an alarming in robust prediction of immunotherapy response in colorectal cancer. Cancers (basel) 14(19):4875
Luo W, Xiang W, Gan L, Che J, Li J, Wang Y, Han L, Gu R, Ye L, Wang R, Zhang X, Xu Y, Dai W, Mo S, Li Q, Cai G (2022) Bulk and single-cell transcriptome profiling reveal necroptosis-based molecular classification, tumor microenvironment infiltration characterization, and prognosis prediction in colorectal cancer. J Transl Med 20(1):235
Mori M, Barnard GF, Staniunas RJ, Jessup JM, Steele GD Jr, Chen LB (1993) Prothymosin-alpha Mrna expression correlates with that of C-Myc in human colon cancer. Oncogene 8(10):2821–2826
Nguyen LH, Goel A, Chung DC (2020) Pathways of colorectal carcinogenesis. Gastroenterology 158(2):291–302
Pages F, Kirilovsky A, Mlecnik B, Asslaber M, Tosolini M, Bindea G, Lagorce C, Wind P, Marliot F, Bruneval P, Zatloukal K, Trajanoski Z, Berger A, Fridman WH, Galon J (2009) In situ cytotoxic and memory T cells predict outcome in patients with early-stage colorectal cancer. J Clin Oncol 27(35):5944–5951
Pan Y, Liu G, Zhou F, Su B, Li Y (2018) DNA methylation profiles in cancer diagnosis and therapeutics. Clin Exp Med 18(1):1–14
Quesada-Calvo F, Massot C, Bertrand V, Longuespee R, Bletard N, Somja J, Mazzucchelli G, Smargiasso N, Baiwir D, De Pauw-Gillet MC, Delvenne P, Malaise M, Marques CC, Polus M, De Pauw E, Meuwis MA, Louis E (2017) Olfm4, Kng1 and Sec24c identified by proteomics and immunohistochemistry as potential markers of early colorectal cancer stages. Clin Proteom 14:9
Rantalainen M (2018) Application of single-cell sequencing in human cancer. Brief Funct Genom 17(4):273–282
Raut JR, Guan Z, Schrotz-King P, Brenner H (2020) Fecal DNA methylation markers for detecting stages of colorectal cancer and its precursors: a systematic review. Clin Epigenetics 12(1):122
Sammarco G, Gallo G, Vescio G, Picciariello A, De Paola G, Trompetto M, Curro G, Ammendola M (2020) Mast cells, micrornas and others: the role of translational research on colorectal cancer in the forthcoming era of precision medicine. J Clin Med 9(9):2852
Song W, Ren J, Xiang R, Kong C, Fu T (2021) Identification of pyroptosis-related subtypes, the development of a prognosis model, and characterization of tumor microenvironment infiltration in colorectal cancer. Oncoimmunology 10(1):1987636
Sparvero LJ, Asafu-Adjei D, Kang R, Tang D, Amin N, Im J, Rutledge R, Lin B, Amoscato AA, Zeh HJ, Lotze MT (2009) Rage (Receptor for Advanced Glycation Endproducts), rage ligands, and their role in cancer and inflammation. J Transl Med 7:17
Su H, Cai T, Zhang S, Yan X, Zhou L, He Z, Xue P, Li J, Zheng M, Yang X, Feng B (2021) Identification of hub genes associated with neutrophils infiltration in colorectal cancer. J Cell Mol Med 25(7):3371–3380
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global cancer statistics 2020: globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71(3):209–249
Tan G, Huang C, Chen J, Zhi F (2020) Hmgb1 released from Gsdme-mediated pyroptotic epithelial cells participates in the tumorigenesis of colitis-associated colorectal cancer through the Erk1/2 Pathway. J Hematol Oncol 13(1):149
Tanaka A, Sakaguchi S (2017) Regulatory T cells in cancer immunotherapy. Cell Res 27(1):109–118
Viale A, Pettazzoni P, Lyssiotis CA, Ying H, Sanchez N, Marchesini M, Carugo A, Green T, Seth S, Giuliani V, Kost-Alimova M, Muller F, Colla S, Nezi L, Genovese G, Deem AK, Kapoor A, Yao W, Brunetto E, Kang Y, Yuan M, Asara JM, Wang YA, Heffernan TP, Kimmelman AC, Wang H, Fleming JB, Cantley LC, DePinho RA, Draetta GF (2014) Oncogene ablation-resistant pancreatic cancer cells depend on mitochondrial function. Nature 514(7524):628–632
Viale A, Corti D, Draetta GF (2015) Tumors and mitochondrial respiration: a neglected connection. Cancer Res 75(18):3685–3686
Vinuesa CG, Linterman MA, Yu D, MacLennan IC (2016) Follicular helper T Cells. Annu Rev Immunol 34:335–368
Wang XY, Chen SH, Zhang YN, Xu CF (2018) Olfactomedin-4 in digestive diseases: a mini-review. World J Gastroenterol 24(17):1881–1887
Wang Z, Xing K, Zhang B, Zhang Y, Chai T, Geng J, Qin X, Zhang X, Xu C (2022) Identification of prognostic gene signatures by developing a Scrna-Seq-based integration approach to predict recurrence and chemotherapy benefit in stage Ii-Iii colorectal cancer. Int J Mol Sci 23(20):12460
Warburg O, Wind F, Negelein E (1927) The metabolism of tumors in the body. J Gen Physiol 8(6):519–530
Wentzensen N, Wilz B, Findeisen P, Wagner R, Dippold W, von Knebel Doeberitz M, Gebert J (2004) Identification of differentially expressed genes in colorectal adenoma compared to normal tissue by suppression subtractive hybridization. Int J Oncol 24(4):987–994
Xin C, Lai Y, Ji L, Wang Y, Li S, Hao L, Zhang W, Meng R, Xu J, Hong Y, Lou Z (2022) A Novel 9-gene signature for the prediction of postoperative recurrence in stage Ii/Iii colorectal cancer. Front Genet 13:1097234
Zhang L, Yu X, Zheng L, Zhang Y, Li Y, Fang Q, Gao R, Kang B, Zhang Q, Huang JY, Konno H, Guo X, Ye Y, Gao S, Wang S, Hu X, Ren X, Shen Z, Ouyang W, Zhang Z (2018) Lineage tracking reveals dynamic relationships of T cells in colorectal cancer. Nature 564(7735):268–272
Zheng X, Song J, Yu C, Zhou Z, Liu X, Yu J, Xu G, Yang J, He X, Bai X, Luo Y, Bao Y, Li H, Yang L, Xu M, Song N, Su X, Xu J, Ma X, Shi H (2022a) Single-cell transcriptomic profiling unravels the adenoma-initiation role of protein tyrosine kinases during colorectal tumorigenesis. Signal Transduct Target Ther 7(1):60
Zheng W, Wu J, Peng Y, Sun J, Cheng P, Huang Q (2022b) Tumor-associated neutrophils in colorectal cancer development, progression and immunotherapy. Cancers (basel) 14(19):4755
Zhou RW, Xu J, Martin TC, Zachem AL, He J, Ozturk S, Demircioglu D, Bansal A, Trotta AP, Giotti B, Gryder B, Shen Y, Wu X, Carcamo S, Bosch K, Hopkins B, Tsankov A, Steinhagen R, Jones DR, Asara J, Chipuk JE, Brody R, Itzkowitz S, Chio IIC, Hasson D, Bernstein E, Parsons RE (2022) A local tumor microenvironment acquired super-enhancer induces an oncogenic driver in colorectal carcinoma. Nat Commun 13(1):6041
Zhuo C, Xu Y, Ying M, Li Q, Huang L, Li D, Cai S, Li B (2015) Foxp3+ Tregs: heterogeneous phenotypes and conflicting impacts on survival outcomes in patients with colorectal cancer. Immunol Res 61(3):338–347
Acknowledgements
We are indebted to all the patients who participated in studies of TCGA and GEO databases.
Funding
This study was supported by the Science and Technology Development Foundation of Shenzhen (Grant number: JCYJ20190808111817192, JCYJ20210324093807020) and the Scientific Research Initiation foundation for young teachers of Shenzhen University (Grant number: 2019123, QNJS0160).
Author information
Authors and Affiliations
Contributions
Data collection and filtering: XZ, YW, LY; data analysis and figures preparation: XZ; collation of results: HH, LY; article writing: XZ; review and modify article format: XZ, YD; conceived and designed this study: BH, FH. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Ethics statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, X., Yang, L., Deng, Y. et al. Single-cell RNA-Seq and bulk RNA-Seq reveal reliable diagnostic and prognostic biomarkers for CRC. J Cancer Res Clin Oncol 149, 9805–9821 (2023). https://doi.org/10.1007/s00432-023-04882-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-04882-0