Abstract
Background
Acute myeloid leukemia (AML) is a hematological cancer driven on by aberrant myeloid precursor cell proliferation and differentiation. A prognostic model was created in this study to direct therapeutic care.
Methods
Differentially expressed genes (DEGs) were investigated using the RNA-seq data from the TCGA-LAML and GTEx. Weighted Gene Coexpression Network Analysis (WGCNA) examines the genes involved in cancer. Find the intersection genes and construct the PPI network to discover hub genes and remove prognosis-related genes. A nomogram was produced for predicting the prognosis of AML patients using the risk prognosis model that was constructed using COX and Lasso regression analysis. GO, KEGG, and ssGSEA analysis were used to look into its biological function. TIDE score predicts immunotherapy response.
Results
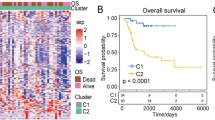
Differentially expressed gene analysis revealed 1004 genes, WGCNA analysis revealed 19,575 tumor-related genes, and 941 intersection genes in total. Twelve prognostic genes were found using the PPI network and prognostic analysis. To build a risk rating model, RPS3A and PSMA2 were examined using COX and Lasso regression analysis. The risk score was used to divide the patients into two groups, and Kaplan–Meier analysis indicated that the two groups had different overall survival rates. Univariate and multivariate COX studies demonstrated that risk score is an independent prognostic factor. According to the TIDE study, the immunotherapy response was better in the low-risk group than in the high-risk group.
Conclusions
We eventually selected out two molecules to construct prediction models that might be used as biomarkers for predicting AML immunotherapy and prognosis.





Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Arber DA, Orazi A, Hasserjian R et al (2016) The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia [J]. Blood 127(20):2391–2405
Ayyadurai VAS, Deonikar P, Mclure KG et al (2022) Molecular systems architecture of Interactome in the Acute Myeloid Leukemia Microenvironment [J]. Cancers 14(3):756
Balch CM, Riley LB, Bae YJ et al (1990) Patterns of human tumor-infiltrating lymphocytes in 120 human cancers [J]. Archiv Surg (chicago, Ill : 1960) 125(2):200–205
Barnes TA, Amir E (2018) HYPE or HOPE: the prognostic value of infiltrating immune cells in cancer [J]. Br J Cancer 118(2):e5
Binnewies M, Roberts EW, Kersten K et al (2018) Understanding the tumor immune microenvironment (TIME) for effective therapy [J]. Nat Med 24(5):541–550
Casey SC, Amedei A, Aquilano K et al (2015) Cancer prevention and therapy through the modulation of the tumor microenvironment [J]. Semi Cancer Biol 35:S199-s223
Döhner H, Estey E, Grimwade D et al (2017) Diagnosis and management of AML in adults: 2017 ELN recommendations from an international expert panel [J]. Blood 129(4):424–447
Han J, Koh YJ, Moon HR et al (2010) Adipose tissue is an extramedullary reservoir for functional hematopoietic stem and progenitor cells [J]. Blood 115(5):957–964
Han Y, Dong Y, Yang Q et al (2018) Acute Myeloid Leukemia cells express ICOS ligand to promote the expansion of regulatory T cells [J]. Front Immunol 9:2227
Han Y, Liu D, Li L (2020) PD-1/PD-L1 pathway: current researches in cancer [J]. Am J Cancer Res 10(3):727–742
Hino C, Pham B, Park D et al (2022) Targeting the Tumor Microenvironment in Acute Myeloid Leukemia: the future of immunotherapy and natural products [J]. Biomedicines 10(6):1410
Jiang Y, Li Y, Zhu B (2015) T-cell exhaustion in the tumor microenvironment [J]. Cell Death Dis 6(6):e1792
Kantarjian H (2016) Acute myeloid leukemia–major progress over four decades and glimpses into the future [J]. Am J Hematol 91(1):131–145
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis [J]. BMC Bioinformat 9:559
Liu H, Liu M, You H et al (2020) Oncogenic network and hub genes for natural killer/T-Cell Lymphoma utilizing WGCNA [J]. Front Oncol 10:223
Mantovani A, Sozzani S, Locati M et al (2002) Macrophage polarization: tumor-associated macrophages as a paradigm for polarized M2 mononuclear phagocytes [J]. Trends Immunol 23(11):549–555
Menter T, Tzankov A (2022) Tumor Microenvironment in Acute Myeloid Leukemia: adjusting Niches [J]. Front Immunol 13:811144
Nandi D, Woodward E, Ginsburg DB et al (1997) Intermediates in the formation of mouse 20S proteasomes: implications for the assembly of precursor beta subunits [J]. EMBO J 16(17):5363–5375
Naora H (1999) Involvement of ribosomal proteins in regulating cell growth and apoptosis: translational modulation or recruitment for extraribosomal activity? [J]. Immunol Cell Biol 77(3):197–205
Panagopoulos I, Gorunova L, Andersen HK et al (2018) PAN3-PSMA2 fusion resulting from a novel t(7;13)(p14;q12) chromosome translocation in a myelodysplastic syndrome that evolved into acute myeloid leukemia [J]. Exp Hematol Oncol 7:7
Qi J, Hu Z, Liu S et al (2020) Comprehensively analyzed Macrophage-Regulated genes indicate that PSMA2 Promotes colorectal cancer progression [J]. Front Oncol 10:618902
Reizis B (2019) Plasmacytoid dendritic cells: development, regulation, and function [J]. Immunity 50(1):37–50
Renosi F, Callanan M, Lefebvre C (2022) Genetics and Epigenetics in Neoplasms with Plasmacytoid Dendritic cells [J]. Cancers 14(17):4132
Rosenberg SA, Spiess P, Lafreniere R (1986) A new approach to the adoptive immunotherapy of cancer with tumor-infiltrating lymphocytes [J]. Science (new York, NY) 233(4770):1318–1321
Sakamoto K, Sato Y, Shinka T et al (2009) Proteasome subunits mRNA expressions correlate with male BMI: implications for a role in obesity [J]. Obesity (silver Spring, Md) 17(5):1044–1049
Sakamoto KM, Grant S, Saleiro D et al (2015) Targeting novel signaling pathways for resistant acute myeloid leukemia [J]. Mol Genet Metab 114(3):397–402
Shafat MS, Oellerich T, Mohr S et al (2017) Leukemic blasts program bone marrow adipocytes to generate a protumoral microenvironment [J]. Blood 129(10):1320–1332
Shallis RM, Wang R, Davidoff A et al (2019) Epidemiology of acute myeloid leukemia: recent progress and enduring challenges [J]. Blood Rev 36:70–87
Swiecki M, Colonna M (2015) The multifaceted biology of plasmacytoid dendritic cells [J]. Nat Rev Immunol 15(8):471–485
Szczepanski MJ, Szajnik M, Czystowska M et al (2009) Increased frequency and suppression by regulatory T cells in patients with acute myelogenous leukemia [J]. Clin Cancer Res off J Am Assoc Cancer Res 15(10):3325–3332
Taghiloo S, Asgarian-Omran H (2021) Immune evasion mechanisms in acute myeloid leukemia: a focus on immune checkpoint pathways [J]. Crit Rev Oncol Hematol 157:103164
Tang Y, He Y, Li C et al (2018) RPS3A positively regulates the mitochondrial function of human periaortic adipose tissue and is associated with coronary artery diseases [J]. Cell Discovery 4:52
Tettamanti S, Pievani A, Biondi A et al (2022) Catch me if you can: how AML and its niche escape immunotherapy [J]. Leukemia 36(1):13–22
Ustun C, Miller JS, Munn DH et al (2011) Regulatory T cells in acute myelogenous leukemia: is it time for immunomodulation? [J]. Blood 118(19):5084–5095
Zhou X, Liao WJ, Liao JM et al (2015) Ribosomal proteins: functions beyond the ribosome [J]. J Mol Cell Biol 7(2):92–104
Zhou C, Weng J, Liu C et al (2020) High RPS3A expression correlates with low tumor immune cell infiltration and unfavorable prognosis in hepatocellular carcinoma patients [J]. Am J Cancer Res 10(9):2768–2784
Funding
There is no funding support for this research.
Author information
Authors and Affiliations
Contributions
This study was independently completed by the author.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the publication of this paper has no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, N. Analysis of prognostic biomarker models and immune microenvironment in acute myeloid leukemia by integrative bioinformatics. J Cancer Res Clin Oncol 149, 9609–9619 (2023). https://doi.org/10.1007/s00432-023-04871-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-04871-3