Abstract
Purpose
FOXA1, as a pioneering transcription factor, has been shown to drive prostate cancer progression. Previous studies showed that FOXA1 expression in prostate cancer was positively associated with cancer angiolymphatic invasion and metastasis. However, the mechanism underlying the correlation between FOXA1 and prostate cancer angiolymphatic invasion and metastasis remains largely unclear.
Methods
Herein, we set out to investigate the role of FOXA1 in the interactions between prostate cancer cells and endothelial cells. Endothelial cells' phenotypes were assessed through CCK‐8 assay, Transwell migration assay, and tube formation assay. The angiogenic factors acting on endothelial cells mediated by FOXA1were characterized by RNA-seq, qPCR array, angiogenesis cytokines array, and ELISA assay. The impact of FOXA1 on tumor angiogenesis was examined in a xenograft model in nude mice. The effect of FOXA1 on prostate cancer angiogenesis was validated on a primary prostate cancer tissue microarray.
Results
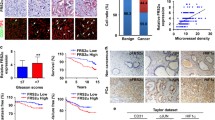
FOXA1 expression in prostate cancer cells promoted endothelial cell proliferation, migration, and tube formation in vitro. Mechanistically, FOXA1 increased pro-angiogenic factors production, including EGF, Endothelin-1, and Endoglin. Moreover, in vivo study showed that FOXA1 facilitated tumor angiogenesis. Furthermore, clinical samples investigation indicated that FOXA1 enhanced prostate cancer angiogenesis.
Conclusion
Overall, these findings illustrated a tumor angiogenesis-promoting role of FOXA1 in prostate cancer.









Similar content being viewed by others
References
Adams EJ et al (2019) FOXA1 mutations alter pioneering activity, differentiation and prostate cancer phenotypes. Nature 571:408–412. https://doi.org/10.1038/s41586-019-1318-9
Attard G et al (2016) Prostate cancer. Lancet 387:70–82. https://doi.org/10.1016/S0140-6736(14)61947-4
Ausprunk DH, Folkman J (1977) Migration and proliferation of endothelial cells in preformed and newly formed blood vessels during tumor angiogenesis. Microvasc Res 14:53–65. https://doi.org/10.1016/0026-2862(77)90141-8
Bagnato A, Spinella F (2003) Emerging role of endothelin-1 in tumor angiogenesis. Trends Endocrinol Metab 14:44–50. https://doi.org/10.1016/s1043-2760(02)00010-3
Boorjian SA et al (2007) Long-term outcome after radical prostatectomy for patients with lymph node positive prostate cancer in the prostate specific antigen era. J Urol 178:864–870. https://doi.org/10.1016/j.juro.2007.05.048 (Discussion 870-861)
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68:394–424. https://doi.org/10.3322/caac.21492
Castonguay R et al (2011) Soluble endoglin specifically binds bone morphogenetic proteins 9 and 10 via its orphan domain, inhibits blood vessel formation, and suppresses tumor growth. J Biol Chem 286:30034–30046. https://doi.org/10.1074/jbc.M111.260133
Chen F et al (2015) New horizons in tumor microenvironment biology: challenges and opportunities. BMC Med 13:45. https://doi.org/10.1186/s12916-015-0278-7
del Castillo G et al (2015) Soluble endoglin antagonizes Met signaling in spindle carcinoma cells. Carcinogenesis 36:212–222. https://doi.org/10.1093/carcin/bgu240
Folkman J, Klagsbrun M (1987) Angiogenic factors. Science 235:442–447. https://doi.org/10.1126/science.2432664
Fonsatti E et al (2000) Endoglin is a suitable target for efficient imaging of solid tumors: in vivo evidence in a canine mammary carcinoma model. Clin Cancer Res off J Am Assoc Can Res 6:2037–2043
Fonsatti E et al (2001) Endoglin: an accessory component of the TGF-beta-binding receptor-complex with diagnostic, prognostic, and bioimmunotherapeutic potential in human malignancies. J Cell Physiol 188:1–7
Gallardo-Vara E et al (2019) Endoglin protein interactome profiling identifies TRIM21 and Galectin-3 as new binding partners. Cells. https://doi.org/10.3390/cells8091082
Gao S et al (2019) Forkhead domain mutations in FOXA1 drive prostate cancer progression. Cell Res 29:770–772. https://doi.org/10.1038/s41422-019-0203-2
Gao S et al (2020) Chromatin binding of FOXA1 is promoted by LSD1-mediated demethylation in prostate cancer. Nat Genet 52:1011–1017. https://doi.org/10.1038/s41588-020-0681-7
Gerhardt J et al (2012) FOXA1 promotes tumor progression in prostate cancer and represents a novel hallmark of castration-resistant prostate cancer. Am J Pathol 180:848–861. https://doi.org/10.1016/j.ajpath.2011.10.021
Gimbrone MA Jr, Leapman SB, Cotran RS, Folkman J (1972) Tumor dormancy in vivo by prevention of neovascularization. J Exp Med 136:261–276. https://doi.org/10.1084/jem.136.2.261
Hanahan D, Folkman J (1996) Patterns and emerging mechanisms of the angiogenic switch during tumorigenesis. Cell 86:353–364. https://doi.org/10.1016/s0092-8674(00)80108-7
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144:646–674. https://doi.org/10.1016/j.cell.2011.02.013
Hawinkels LJ et al (2010) Matrix metalloproteinase-14 (MT1-MMP)-mediated endoglin shedding inhibits tumor angiogenesis. Cancer Res 70:4141–4150. https://doi.org/10.1158/0008-5472.CAN-09-4466
Hida K, Maishi N, Annan DA, Hida Y (2018) Contribution of tumor endothelial cells in cancer progression. Int J Mol Sci. https://doi.org/10.3390/ijms19051272
Imamura Y et al (2012) FOXA1 promotes tumor progression in prostate cancer via the insulin-like growth factor binding protein 3 pathway. PLoS ONE 7:e42456. https://doi.org/10.1371/journal.pone.0042456
Jain RK, Mehta RJ, Nakshatri H, Idrees MT, Badve SS (2011) High-level expression of forkhead-box protein A1 in metastatic prostate cancer. Histopathology 58:766–772. https://doi.org/10.1111/j.1365-2559.2011.03796.x
Jin HJ, Zhao JC, Ogden I, Bergan RC, Yu J (2013) Androgen receptor-independent function of FoxA1 in prostate cancer metastasis. Cancer Res 73:3725–3736. https://doi.org/10.1158/0008-5472.CAN-12-3468
Kim J et al (2017) FOXA1 inhibits prostate cancer neuroendocrine differentiation. Oncogene 36:4072–4080. https://doi.org/10.1038/onc.2017.50
Kopetz ES, Nelson JB, Carducci MA (2002) Endothelin-1 as a target for therapeutic intervention in prostate cancer. Invest New Drugs 20:173–182
Lawera A et al (2019) Role of soluble endoglin in BMP9 signaling. Proc Natl Acad Sci USA 116:17800–17808. https://doi.org/10.1073/pnas.1816661116
Li CG, Wilson PB, Bernabeu C, Raab U, Wang JM, Kumar S (1998) Immunodetection and characterisation of soluble CD105-TGFbeta complexes. J Immunol Methods 218:85–93. https://doi.org/10.1016/s0022-1759(98)00118-5
Li C, Guo B, Wilson PB, Stewart A, Byrne G, Bundred N, Kumar S (2000) Plasma levels of soluble CD105 correlate with metastasis in patients with breast cancer. Int J Cancer 89:122–126. https://doi.org/10.1002/(sici)1097-0215(20000320)89:2%3c122::aid-ijc4%3e3.0.co;2-m
Li J et al (2020) A genomic and epigenomic atlas of prostate cancer in Asian populations. Nature 580:93–99. https://doi.org/10.1038/s41586-020-2135-x
Litwin MS, Tan H-J (2017) The diagnosis and treatment of prostate cancer: a review. JAMA 317:2532–2542. https://doi.org/10.1001/jama.2017.7248
Nassiri F et al (2011) Endoglin (CD105): a review of its role in angiogenesis and tumor diagnosis, progression and therapy. Anticancer Res 31:2283–2290
Parolia A et al (2019) Distinct structural classes of activating FOXA1 alterations in advanced prostate cancer. Nature 571:413–418. https://doi.org/10.1038/s41586-019-1347-4
Pérez-Gómez E et al (2007) A role for endoglin as a suppressor of malignancy during mouse skin carcinogenesis. Can Res 67:10268–10277
Pomerantz MM et al (2015) The androgen receptor cistrome is extensively reprogrammed in human prostate tumorigenesis. Nat Genet 47:1346–1351. https://doi.org/10.1038/ng.3419
Robinson JL et al (2014) Elevated levels of FOXA1 facilitate androgen receptor chromatin binding resulting in a CRPC-like phenotype. Oncogene 33:5666–5674. https://doi.org/10.1038/onc.2013.508
Sholley MM, Ferguson GP, Seibel HR, Montour JL, Wilson JD (1984) Mechanisms of neovascularization vascular sprouting can occur without proliferation of endothelial cells. Lab Invest 51:624–634
Song B et al (2019) Targeting FOXA1-mediated repression of TGF-beta signaling suppresses castration-resistant prostate cancer progression. J Clin Invest 129:569–582. https://doi.org/10.1172/JCI122367
Sunkel B et al (2016) Integrative analysis identifies targetable CREB1/FoxA1 transcriptional co-regulation as a predictor of prostate cancer recurrence. Nucleic Acids Res 44:4105–4122. https://doi.org/10.1093/nar/gkv1528
Swami U, McFarland TR, Nussenzveig R, Agarwal N (2020) Advanced prostate cancer: treatment advances and future directions. Trends Cancer 6:702–715. https://doi.org/10.1016/j.trecan.2020.04.010
ten Dijke P, Goumans MJ, Pardali E (2008) Endoglin in angiogenesis and vascular diseases. Angiogenesis 11:79–89. https://doi.org/10.1007/s10456-008-9101-9
Trojan L, Thomas D, Knoll T, Grobholz R, Alken P, Michel MS (2004) Expression of pro-angiogenic growth factors VEGF, EGF and bFGF and their topographical relation to neovascularisation in prostate cancer. Urol Res 32:97–103. https://doi.org/10.1007/s00240-003-0383-5
Wang QB et al (2009) Androgen receptor regulates a distinct transcription program in androgen-independent prostate. Cancer Cell 138:245–256. https://doi.org/10.1016/j.cell.2009.04.056
Xu B, Song B, Lu X, Kim J, Hu M, Zhao JC, Yu J (2019) Altered chromatin recruitment by FOXA1 mutations promotes androgen independence and prostate cancer progression. Cell Res 29:773–775. https://doi.org/10.1038/s41422-019-0204-1
Zhang C et al (2011) Definition of a FoxA1 Cistrome that is crucial for G1 to S-phase cell-cycle transit in castration-resistant prostate cancer. Cancer Res 71:6738–6748. https://doi.org/10.1158/0008-5472.CAN-11-1882
Zhao J et al (2020) RNA-binding protein Musashi2 stabilizing androgen receptor drives prostate cancer progression. Cancer Sci 111:369–382. https://doi.org/10.1111/cas.14280
Zhong H et al (2000) Modulation of hypoxia-inducible factor 1α expression by the epidermal growth factor/phosphatidylinositol 3-Kinase/PTEN/AKT/FRAP pathway in human prostate cancer cells: implications for tumor angiogenesis and therapeutics. Cancer Res 60:1541–1545
Zhou S et al (2020) Noncoding mutations target cis-regulatory elements of the FOXA1 plexus in prostate cancer. Nat Commun 11:441. https://doi.org/10.1038/s41467-020-14318-9
Acknowledgements
We thank Prof. Deshui Jia for his helpful suggestions, Dr. Chenyi Jiang, for his precious technical support and coaching. This work was supported by the Natural Science Foundation of Shanghai, China (Grant no: 19ZR1441200).
Funding
This work was sponsored by the Natural Science Foundation of Shanghai, China (Grant no: 19ZR1441200).
Author information
Authors and Affiliations
Contributions
Conception and design: XW, BH. Development of methodology: YS, XW. Acquisition of data: YS, YZ. Analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis): YS, YZ. Drafting the article or revising it critically for important intellectual content: YS, XW. Study supervision: BH, XW.
Corresponding authors
Ethics declarations
Conflict of interest
None.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
432_2021_3730_MOESM1_ESM.pdf
Supplementary Fig. S1 Identification of primary HUVEC cells by immunofluorescence.a. Bright field of primary HUVEC cells. b. Immunofluorescence microscopy of primary HUVEC cells stained with DAPI. c. Immunofluorescence microscopy of primary HUVEC cells stained with vWF. d. Immunofluorescence microscopy of primary HUVEC cells stained with CD31. Scale bars, 50 μm (PDF 1272 KB)
432_2021_3730_MOESM2_ESM.pdf
Supplementary Fig. S2 Functional enrichment analysis of differentially expressed genes between oeNC-DU145 and oeFOXA1-DU145 cells. a. The top 10 upregulated pathways between oeNC-DU145 and oeFOXA1-DU145 cells identified by the GO enrichment analysis. b. The top 20 upregulated pathways in oeFOXA1-DU145 cells were identified by the KEGG enrichment analysis (PDF 264 KB)
432_2021_3730_MOESM3_ESM.pdf
Supplementary Fig. S3 Quantitative RT-PCR analyses of the mRNA extracted from the excised xenograft tumors containing oeNC-DU145 (n=5) or oeFOXA1-DU145 cells (n=6). a. Quantitative PCR determination of human FOXA1, mouse CD31, mouse VE-Cadherin. b. Quantitative PCR determination of human EGF, Endothelin-1, Endoglin. mCD31, mouse CD31; mVE-Cad, mouse VE-Cadherin. **P<0.01; ***P<0.001 (PDF 17593 KB)
432_2021_3730_MOESM4_ESM.tif
Supplementary Fig. S4 Functional assays of HUVEC cells cultured with different LNCaP cells or LNCaP conditioned media. a. Quantitative RT-PCR assays of FOXA1 expression in LNCaP cells with FOXA1 knockdown using two different shRNAs. b. Western blotting assays of FOXA1 expression in LNCaP cells with FOXA1 knockdown using two different shRNAs. c. Growth curves of HUVECs co-cultured with different LNCaP cells. d. Growth curves of HUVECs cultured with conditioned media from various LNCaP cells. e. Transwell migration assays of HUVECs co-cultured with different LNCaP cells. Scale bars, 50 μm. f. Transwell migration assays of HUVECs cultured with conditioned media from various LNCaP cells. Scale bars, 50 μm. g. Tube formation assays of HUVECs co-cultured with different LNCaP cells. Scale bars, 250 μm. h. Tube formation assays of HUVECs cultured with conditioned media from various LNCaP cells. Scale bars, 250 μm. *P<0.05. **P<0.01. ***P<0.001 (TIF 11205 KB)
432_2021_3730_MOESM5_ESM.tif
Supplementary Fig. S5 FOXA1 knockdown in prostate cancer cells suppressed the secretion of pro-angiogenic cytokines. Quantitative PCR determination of HIF-1α, VEGFA, FGF2, HGF, TGF-β, Endoglin, Endothelin-1, EGF, THBS1, and PLG gene expression in shNC-LNCaP and shFOXA1-LNCaP cells. ns: not significant. **P<0.01. ***P<0.001 (TIF 901 KB)
432_2021_3730_MOESM6_ESM.tif
Supplementary Fig. S6 The pro-migration of endothelial cells by FOXA1 overexpression in prostate cancer cells was reversed by antibodies to EGF, Endoglin, and Endothelin-1. (oeFOXA1-DU145-CM as the control group, the other groups as the case groups. **P<0.01; ***P<0.001; ****P<0.0001) (TIF 8824 KB)
Rights and permissions
About this article
Cite this article
Su, Y., Zhang, Y., Zhao, J. et al. FOXA1 promotes prostate cancer angiogenesis by inducing multiple pro-angiogenic factors expression. J Cancer Res Clin Oncol 147, 3225–3243 (2021). https://doi.org/10.1007/s00432-021-03730-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-021-03730-3