Abstract
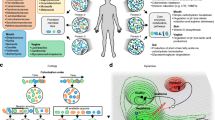
There is evidence from numerous studies that dysbiosis of the microbiome provokes various immune-mediated diseases, obesity, diabetes, and cancers by regulating metabolites, host genetics, environmental elements, and stress. Such reports are yet to define an accurate regulatory network for host-gut microbiome communication. miRNAs have recently emerged as crucial mediators of this communication, as portrayed by their interaction with the host microbiome. This mini-review summarizes the bi-direction effects between miRNA and microbiome and elucidates their role in carcinogenesis. An in-depth understanding of the association of miRNA with host-microbiome could be valuable to improve cancer remission, diagnosis, and treatment, and may help to potential tumor markers.
Similar content being viewed by others
References
Allegra A, Innao V, Allegra AG, Ettari R, Pugliese M, Pulvirenti N, Musolino C (2019) Role of the microbiota in hematologic malignancies. Neth J Med 77(2):67–80 (PMID: 30895929)
Aykut B, Pushalkar S, Chen R, Li Q, Abengozar R, Kim JI, Shadaloey SA, Wu D, Preiss P, Verma N, Guo Y, Saxena A, Vardhan M, Diskin B, Wang W, Leinwand J, Kurz E, Kochen Rossi JA, Hundeyin M, Zambrinis C, Li X, Saxena D, Miller G (2019) The fungal mycobiome promotes pancreatic oncogenesis via activation of MBL. Nature 574(7777):264–267. https://doi.org/10.1038/s41586-019-1608-2 (Epub 2019 Oct 2. PMID: 31578522; PMCID: PMC6858566)
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116(2):281–297. https://doi.org/10.1016/s0092-8674(04)00045-5 (PMID: 14744438)
Bhatt AP, Redinbo MR, Bultman SJ (2017) The role of the microbiome in cancer development and therapy. CA Cancer J Clin 67(4):326–344. https://doi.org/10.3322/caac.21398 (PMID: 28481406; PMCID: PMC5530583)
Burns MB, Montassier E, Abrahante J, Priya S, Niccum DE, Khoruts A, Starr TK, Knights D, Blekhman R (2018) Colorectal cancer mutational profiles correlate with defined microbial communities in the tumor microenvironment. PLoS Genet 14(6):e1007376. https://doi.org/10.1371/journal.pgen.1007376 (PMID:29924794;PMCID:PMC6028121)
Coker OO, Dai Z, Nie Y, Zhao G, Cao L, Nakatsu G, Wu WK, Wong SH, Chen Z, Sung JJY, Yu J (2018) Mucosal microbiome dysbiosis in gastric carcinogenesis. Gut 67(6):1024–1032. https://doi.org/10.1136/gutjnl-2017-314281 (PMID: 28765474; PMCID: PMC5969346)
Cooks T, Pateras IS, Jenkins LM, Patel KM, Robles AI, Morris J, Forshew T, Appella E, Gorgoulis VG, Harris CC (2018) Mutant p53 cancers reprogram macrophages to tumor supporting macrophages via exosomal miR-1246. Nat Commun 9(1):771. https://doi.org/10.1038/s41467-018-03224-w (PMID:29472616;PMCID:PMC5823939)
Creemers EE, Tijsen AJ, Pinto YM (2012) Circulating microRNAs: novel biomarkers and extracellular communicators in cardiovascular disease? Circ Res 110(3):483–495. https://doi.org/10.1161/CIRCRESAHA.111.247452 (PMID: 22302755)
Dickson RP, Huffnagle GB (2015) The lung microbiome: new principles for respiratory bacteriology in health and disease. PLoS Pathog 11(7):e1004923. https://doi.org/10.1371/journal.ppat.1004923 (PMID:26158874;PMCID:PMC4497592)
Dickson RP, Martinez FJ, Huffnagle GB (2014) The role of the microbiome in exacerbations of chronic lung diseases. Lancet 384(9944):691–702. https://doi.org/10.1016/S0140-6736(14)61136-3 (PMID:25152271;PMCID:PMC4166502)
Dzutsev A, Badger JH, Perez-Chanona E, Roy S, Salcedo R, Smith CK, Trinchieri G (2017) Microbes and Cancer. Annu Rev Immunol 35:199–228. https://doi.org/10.1146/annurev-immunol-051116-052133 (Epub 2017 Jan 30 PMID: 28142322)
Gebert LFR, MacRae IJ (2019) Regulation of microRNA function in animals. Nat Rev Mol Cell Biol 20(1):21–37. https://doi.org/10.1038/s41580-018-0045-7 (PMID:30108335;PMCID:PMC6546304)
Gensollen T, Iyer SS, Kasper DL, Blumberg RS (2016) How colonization by microbiota in early life shapes the immune system. Science 352(6285):539–544. https://doi.org/10.1126/science.aad9378 (PMID:27126036;PMCID:PMC5050524)
Gomez de Agüero M, Ganal-Vonarburg SC, Fuhrer T, Rupp S, Uchimura Y, Li H, Steinert A, Heikenwalder M, Hapfelmeier S, Sauer U, Mccoy KD, Macpherson AJ (2016) The maternal microbiota drives early postnatal innate immune development. Science 351(6279):1296–1302. https://doi.org/10.1126/science.aad2571 (PMID: 26989247)
Guo S, Chen J, Chen F, Zeng Q, Liu WL, Zhang G (2020) Exosomes derived from Fusobacteriumnucleatum-infected colorectal cancer cells facilitate tumour metastasis by selectively carrying miR-1246/92b-3p/27a-3p and CXCL16. Gut 7:gutjnl-2020-321187. https://doi.org/10.1136/gutjnl-2020-321187 (Epub ahead of print. PMID: 33172926)
Izaurralde E (2015) Gene regulation. Breakers and blockers—miRNAs at work. Science 349(6246):380–382. https://doi.org/10.1126/science.1260969 (Epub 2015 Jul 23. PMID: 26206919)
Liu S, da Cunha AP, Rezende RM, Cialic R, Wei Z, Bry L, Comstock LE, Gandhi R, Weiner HL (2016) The host shapes the gut microbiota via fecal MicroRNA. Cell Host Microbe 19(1):32–43. https://doi.org/10.1016/j.chom.2015.12.005 (PMID:26764595;PMCID:PMC4847146)
Liu S, Rezende RM, Moreira TG, Tankou SK, Cox LM, Wu M, Song A, Dhang FH, Wei Z, Costamagna G, Weiner HL (2019) Oral administration of miR-30d from Feces of MS patients suppresses MS-like symptoms in mice by expanding akkermansia muciniphila. Cell Host Microbe 26(6):779-794.e8. https://doi.org/10.1016/j.chom.2019.10.008 (Epub 2019 Nov 26. PMID: 31784260; PMCID: PMC6948921)
Luo X, Burwinkel B, Tao S, Brenner H (2011) MicroRNA signatures: novel biomarker for colorectal cancer? Cancer Epidemiol Biomarkers Prev 20(7):1272–1286. https://doi.org/10.1158/1055-9965.EPI-11-0035 (Epub 2011 May 6 PMID: 21551242)
Lynch JB, Hsiao EY (2019) Microbiomes as sources of emergent host phenotypes. Science 365(6460):1405–1409. https://doi.org/10.1126/science.aay0240 (PMID: 31604267)
Mager LF, Burkhard R, Pett N, Cooke NCA, Brown K, Ramay H, Paik S, Stagg J, Groves RA, Gallo M, Lewis IA, Geuking MB, McCoy KD (2020) Microbiome-derived inosine modulates response to checkpoint inhibitor immunotherapy. Science 369(6510):1481–1489. https://doi.org/10.1126/science.abc3421 (Epub 2020 Aug 13 PMID: 32792462)
Mao Q, Jiang F, Yin R, Wang J, Xia W, Dong G, Ma W, Yang Y, Xu L, Hu J (2018) Interplay between the lung microbiome and lung cancer. Cancer Lett 415:40–48. https://doi.org/10.1016/j.canlet.2017.11.036 (Epub 2017 Dec 2 PMID: 29197615)
Mao Q, Ma W, Wang Z, Liang Y, Zhang T, Yang Y, Xia W, Jiang F, Hu J, Xu L (2020) Differential flora in the microenvironment of lung tumor and paired adjacent normal tissues. Carcinogenesis 41(8):1094–1103. https://doi.org/10.1093/carcin/bgaa044 (PMID: 32658980)
Mima K, Nakagawa S, Sawayama H, Ishimoto T, Imai K, Iwatsuki M, Hashimoto D, Baba Y, Yamashita YI, Yoshida N, Chikamoto A, Baba H (2017) The microbiome and hepatobiliary-pancreatic cancers. Cancer Lett 402:9–15. https://doi.org/10.1016/j.canlet.2017.05.001 (Epub 2017 May 17 PMID: 28527946)
Ogata-Kawata H, Izumiya M, Kurioka D, Honma Y, Yamada Y, Furuta K, Gunji T, Ohta H, Okamoto H, Sonoda H, Watanabe M, Nakagama H, Yokota J, Kohno T, Tsuchiya N (2014) Circulating exosomal microRNAs as biomarkers of colon cancer. PLoS ONE 9(4):e92921. https://doi.org/10.1371/journal.pone.0092921 (PMID:24705249;PMCID:PMC3976275)
Peck BC, Mah AT, Pitman WA, Ding S, Lund PK, Sethupathy P (2017) Functional transcriptomics in diverse intestinal epithelial cell types reveals robust MicroRNA sensitivity in intestinal stem cells to microbial status. J Biol Chem 292(7):2586–2600. https://doi.org/10.1074/jbc.M116.770099
Peng Z, Cheng S, Kou Y, Wang Z, Jin R, Hu H, Zhang X, Gong JF, Li J, Lu M, Wang X, Zhou J, Lu Z, Zhang Q, Tzeng DTW, Bi D, Tan Y, Shen L (2020) The gut microbiome is associated with clinical response to anti-PD-1/PD-L1 immunotherapy in gastrointestinal cancer. Cancer Immunol Res 8(10):1251–1261. https://doi.org/10.1158/2326-6066.CIR-19-1014 (Epub 2020 Aug 27 PMID: 32855157)
Roy S, Trinchieri G (2017) Microbiota: a key orchestrator of cancer therapy. Nat Rev Cancer 17(5):271–285. https://doi.org/10.1038/nrc.2017.13 (Epub 2017 Mar 17 PMID: 28303904)
Rubinstein MR, Wang X, Liu W, Hao Y, Cai G, Han YW (2013) Fusobacterium nucleatum promotes colorectal carcinogenesis by modulating E-cadherin/β-catenin signaling via its FadA adhesin. Cell Host Microbe 14(2):195–206. https://doi.org/10.1016/j.chom.2013.07.012 (PMID:23954158;PMCID:PMC3770529)
Singh N, Shirdel EA, Waldron L, Zhang RH, Jurisica I, Comelli EM (2012) The murine caecal microRNA signature depends on the presence of the endogenous microbiota. Int J Biol Sci 8(2):171–186. https://doi.org/10.7150/ijbs.8.171 (Epub 2011 Dec 19. PMID: 22211115; PMCID: PMC3248702)
Tailford LE, Crost EH, Kavanaugh D, Juge N (2015) Mucin glycan foraging in the human gut microbiome. Front Genet 6:81. https://doi.org/10.3389/fgene.2015.00081 (PMID:25852737;PMCID:PMC4365749)
Tarallo S, Ferrero G, Gallo G, Francavilla A, Clerico G, Realis Luc A, Manghi P, Thomas AM, Vineis P, Segata N, Pardini B, Naccarati A, Cordero F. Correction for Tarallo et al (2020) Altered fecal small RNA profiles in colorectal cancer reflect gut microbiome composition in stool samples. maSystems 5(1):e00072-e120. https://doi.org/10.1128/mSystems.00072-20 (Erratum for: mSystems. 2019;4(5): PMID: 32071156; PMCID: PMC7029216)
Teng Y, Ren Y, Sayed M, Hu X, Lei C, Kumar A, Hutchins E, Mu J, Deng Z, Luo C, Sundaram K, Sriwastva MK, Zhang L, Hsieh M, Reiman R, Haribabu B, Yan J, Jala VR, Miller DM, Van Keuren-Jensen K, Merchant ML, McClain CJ, Park JW, Egilmez NK, Zhang HG (2018) Plant-derived exosomal micrornas shape the gut microbiota. Cell Host Microbe 24(5):637-652.e8. https://doi.org/10.1016/j.chom.2018.10.001
Tomkovich S, Gharaibeh RZ, Dejea CM, Pope JL, Jiang J, Winglee K, Gauthier J, Newsome RC, Yang Y, Fodor AA, Schmittgen TD, Sears CL, Jobin C (2020) Human colon mucosal biofilms and murine host communicate via altered mRNA and microRNA expression during cancer. Msystems 5(1):e00451-e519. https://doi.org/10.1128/mSystems.00451-19 (PMID: 31937674; PMCID: PMC6967385)
Tsay JJ, Wu BG, Badri MH, Clemente JC, Shen N, Meyn P, Li Y, Yie TA, Lhakhang T, Olsen E, Murthy V, Michaud G, Sulaiman I, Tsirigos A, Heguy A, Pass H, Weiden MD, Rom WN, Sterman DH, Bonneau R, Blaser MJ, Segal LN (2018) Airway microbiota is associated with upregulation of the PI3K pathway in lung cancer. Am J Respir Crit Care Med 198(9):1188–1198. https://doi.org/10.1164/rccm.201710-2118OC (PMID:29864375;PMCID:PMC6221574)
Turnbaugh PJ, Ley RE, Hamady M, Fraser-Liggett CM, Knight R, Gordon JI (2007) The human microbiome project. Nature 449:804–810
Turroni S, Brigidi P, Cavalli A, Candela M (2018) Microbiota-host transgenomic metabolism, bioactive molecules from the inside. J Med Chem 61(1):47–61. https://doi.org/10.1021/acs.jmedchem.7b00244 (Epub 2017 Aug 3 PMID: 28745893)
Uronis JM, Mühlbauer M, Herfarth HH, Rubinas TC, Jones GS, Jobin C (2009) Modulation of the intestinal microbiota alters colitis-associated colorectal cancer susceptibility. PLoS ONE 4(6):e6026. https://doi.org/10.1371/journal.pone.0006026 (PMID:19551144;PMCID:PMC2696084)
Vogtmann E, Goedert JJ (2016) Epidemiologic studies of the human microbiome and cancer. Br J Cancer 114(3):237–242. https://doi.org/10.1038/bjc.2015.465 (Epub ahead of print. PMID: 33296049)
Wang T, Cai G, Qiu Y, Fei N, Zhang M, Pang X, Jia W, Cai S, Zhao L (2012) Structural segregation of gut microbiota between colorectal cancer patients and healthy volunteers. ISME J 6(2):320–329. https://doi.org/10.1038/ismej.2011.109 (Epub 2011 Aug 18. PMID: 21850056; PMCID: PMC3260502)
Weir TL, Manter DK, Sheflin AM, Barnett BA, Heuberger AL, Ryan EP (2013) Stool microbiome and metabolome differences between colorectal cancer patients and healthy adults. PLoS ONE 8(8):e70803. https://doi.org/10.1371/journal.pone.0070803 (PMID:23940645;PMCID:PMC3735522)
Wong-Rolle A, Wei HK, Zhao C, Jin C (2020) Unexpected guests in the tumor microenvironment: microbiome in cancer. Protein Cell. https://doi.org/10.1007/s13238-020-00813-8 (Epub ahead of print. PMID: 33296049)
Wu CW, Ng SS, Dong YJ, Ng SC, Leung WW, Lee CW, Wong YN, Chan FK, Yu J, Sung JJ (2012) Detection of miR-92a and miR-21 in stool samples as potential screening biomarkers for colorectal cancer and polyps. Gut 61(5):739–745. https://doi.org/10.1136/gut.2011.239236 (Epub 2011 Sep 19 PMID: 21930727)
Xie G, Wang X, Zhao A, Yan J, Chen W, Jiang R, Ji J, Huang F, Zhang Y, Lei S, Ge K, Zheng X, Rajani C, Alegado RA, Liu J, Liu P, Nicholson J, Jia W (2017) Sex-dependent effects on gut microbiota regulate hepatic carcinogenic outcomes. Sci Rep 7:45232. https://doi.org/10.1038/srep45232 (PMID:28345673;PMCID:PMC5366919)
Yang Y, Weng W, Peng J, Hong L, Yang L, Toiyama Y, Gao R, Liu M, Yin M, Pan C, Li H, Guo B, Zhu Q, Wei Q, Moyer MP, Wang P, Cai S, Goel A, Qin H, Ma Y (2017) Fusobacterium nucleatum increases proliferation of colorectal cancer cells and tumor development in mice by activating toll-like receptor 4 signaling to nuclear factor-κB, and up-regulating expression of microRNA-21. Gastroenterology 152(4):851-866.e24. https://doi.org/10.1053/j.gastro.2016.11.018
Yi X, Zhou H, Chao Y, Xiong S, Zhong J, Chai Z, Yang K, Liu Z (2020) Bacteria-triggered tumor-specific thrombosis to enable potent photothermal immunotherapy of cancer. Sci Adv 6(33):3546. https://doi.org/10.1126/sciadv.aba3546 (PMID: 32851163; PMCID: PMC7428325)
Yu T, Guo F, Yu Y, Sun T, Ma D, Han J, Qian Y, Kryczek I, Sun D, Nagarsheth N, Chen Y, Chen H, Hong J, Zou W, Fang JY (2017) Fusobacterium nucleatum Promotes chemoresistance to colorectal cancer by modulating autophagy. Cell 170(3):548-563.e16. https://doi.org/10.1016/j.cell.2017.07.008 (PMID:28753429;PMCID:PMC5767127)
Yuan C, Subramanian S (2019) microRNA-mediated tumor-microbiota metabolic interactions in colorectal cancer. DNA Cell Biol 38(4):281–285. https://doi.org/10.1089/dna.2018.4579 (Epub 2019 Jan 22. PMID: 30668143; PMCID: PMC6477581)
Yuan C, Burns MB, Subramanian S, Blekhman R (2018) Interaction between host microRNAs and the gut microbiota in colorectal cancer. mSystems 3(3):e00205-e217. https://doi.org/10.1128/mSystems.00205-17 (PMID: 29795787; PMCID: PMC5954203)
Zhu Z, Huang J, Li X, Xing J, Chen Q, Liu R, Hua F, Qiu Z, Song Y, Bai C, Mo YY, Zhang Z (2020) Gut microbiota regulate tumor metastasis via circRNA/miRNA networks. Gut Microbes 12(1):1788891. https://doi.org/10.1080/19490976.2020.1788891
Acknowledgements
We thank all our family for allowing us to conduct this research at the National Natural Science Foundation of China, The Project of Invigorating Health Care through Science, Technology and Education, Jiangsu Provincial Medical Innovation Team, The Project of Invigorating Health Care through Science, Technology and Education, Jiangsu Provincial Medical Outstanding Talent, Young Talents Program of Jiangsu Cancer Hospital.
Funding
This work was supported by the grants from the National Natural Science Foundation of China (Grant No. 82073211, 82002434, 82003106); The Project of Invigorating Health Care through Science, Technology and Education, Jiangsu Provincial Medical Innovation Team (CXTDA2017002); The Project of Invigorating Health Care through Science, Technology and Education, Jiangsu Provincial Medical Outstanding Talent (JCRCA2016001); Young Talents Program of Jiangsu Cancer Hospital (23).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declared no conflicts of interest in this work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, Q., Ding, H., Dong, G. et al. Bi-direction effects between microbiome and MiRNAs in carcinogenesis. J Cancer Res Clin Oncol 147, 1299–1305 (2021). https://doi.org/10.1007/s00432-021-03567-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-021-03567-w