Abstract
Tetraspanin (TSPAN) protein family forms a family of transmembrane proteins that act as organizers/scaffold for other proteins. TSPANs are primarily present on plasma membranes although they are also found in other biological membranes. They are organized in tetraspanin-enriched microdomains (TEMs), which allow spatiotemporal tuning of protein functions through the control of their membrane localization. TSPAN6 and TSPAN7 are close paralogs expressed in different tissues, TSPAN7 being highly expressed in the brain. Their functions only started to be unveiled in the late 2000’s and are still poorly understood. Here, we introduce how TSPAN7 was first highlighted has a protein mutated in some forms of X-linked mental retardation, which was later proposed to be caused by defects in neuronal morphogenesis and synaptic transmission. We then discuss the impacts TSPAN7 has on cell morphology of dendritic cells and osteoclasts, through rearrangement of actin cytoskeleton and how TSPAN7 was shown to be a target of autoantibody in patients suffering from type 1 diabetes. Finally, we are addressing the double edge sword that is TSPAN7 in cancer. In the second part of this review, we address the known roles of TSPAN6 and how this protein was shown to participate in synaptic transmission and in amyloid precursor protein secretion, which may contribute to Alzheimer’s disease pathology. We conclude this review by discussing the anti-inflammatory effect of TSPAN6.




Similar content being viewed by others
References
Charrin S, Jouannet S, Boucheix C, Rubinstein E (2014) Tetraspanins at a glance. J Cell Sci 127:3641–3648. https://doi.org/10.1242/jcs.154906
Hemler ME (2005) Tetraspanin functions and associated microdomains. Nat Rev Mol Cell Biol 6:801–811. https://doi.org/10.1038/nrm1736
Termini CM, Gillette JM (2017) Tetraspanins function as regulators of cellular signaling. Front Cell Dev Biol 5:34. https://doi.org/10.3389/fcell.2017.00034
Bassani S, Passafaro M (2012) TSPAN7: a new player in excitatory synapse maturation and function. Bioarchitecture 2:95–97. https://doi.org/10.4161/bioa.20829
Bassani S, Cingolani LA, Valnegri P et al (2012) The X-linked intellectual disability protein TSPAN7 regulates excitatory synapse development and AMPAR trafficking. Neuron 73:1143–1158. https://doi.org/10.1016/j.neuron.2012.01.021
Hemler ME (2008) Targeting of tetraspanin proteins–potential benefits and strategies. Nat Rev Drug Discov 7:747–758. https://doi.org/10.1038/nrd2659
Garcia-España A, Chung P-J, Sarkar IN et al (2008) Appearance of new tetraspanin genes during vertebrate evolution. Genomics 91:326–334. https://doi.org/10.1016/j.ygeno.2007.12.005
Lu J, Li J, Liu S et al (2017) Exosomal tetraspanins mediate cancer metastasis by altering host microenvironment. Oncotarget 8:62803–62815. https://doi.org/10.18632/oncotarget.19119
Reyes R, Cardeñes B, Machado-Pineda Y, Cabañas C (2018) Tetraspanin CD9: a key regulator of cell adhesion in the immune system. Front Immunol 9:863. https://doi.org/10.3389/fimmu.2018.00863
Pileri P, Uematsu Y, Campagnoli S et al (1998) Binding of hepatitis C virus to CD81. Science 282:938–941. https://doi.org/10.1126/science.282.5390.938
Suárez H, Rocha-Perugini V, Álvarez S, Yáñez-Mó M (2018) Tetraspanins, another piece in the HIV-1 replication puzzle. Front Immunol 9:1811. https://doi.org/10.3389/fimmu.2018.01811
Fagerberg L, Hallström BM, Oksvold P et al (2014) Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol Cell Proteomics 13:397–406. https://doi.org/10.1074/mcp.M113.035600
Zemni R, Bienvenu T, Vinet MC et al (2000) A new gene involved in X-linked mental retardation identified by analysis of an X;2 balanced translocation. Nat Genet 24:167–170. https://doi.org/10.1038/72829
Abidi FE, Holinski-Feder E, Rittinger O et al (2002) A novel 2 bp deletion in the TM4SF2 gene is associated with MRX58. J Med Genet 39:430–433. https://doi.org/10.1136/jmg.39.6.430
da Maranduba CM, Sá Moreira E, Müller Orabona G et al (2004) Does the P172H mutation at the TM4SF2 gene cause X-linked mental retardation? Am J Med Genet A 124A:413–415. https://doi.org/10.1002/ajmg.a.20401
Boda B, Mas C, Muller D (2002) Activity-dependent regulation of genes implicated in X-linked non-specific mental retardation. Neuroscience 114:13–17. https://doi.org/10.1016/s0306-4522(02)00218-x
Guo M, Huang T, Cui Y et al (2008) PrPC interacts with tetraspanin-7 through bovine PrP154-182 containing alpha-helix 1. Biochem Biophys Res Commun 365:154–157. https://doi.org/10.1016/j.bbrc.2007.10.160
Perez JL, Khatri L, Chang C et al (2001) PICK1 targets activated protein kinase Calpha to AMPA receptor clusters in spines of hippocampal neurons and reduces surface levels of the AMPA-type glutamate receptor subunit 2. J Neurosci 21:5417–5428
Lee S-A, Suh Y, Lee S et al (2017) Functional expression of dopamine D2 receptor is regulated by tetraspanin 7-mediated postendocytic trafficking. FASEB J 31:2301–2313. https://doi.org/10.1096/fj.201600755RR
Usardi A, Iyer K, Sigoillot SM et al (2017) The immunoglobulin-like superfamily member IGSF3 is a developmentally regulated protein that controls neuronal morphogenesis. Dev Neurobiol 77:75–92. https://doi.org/10.1002/dneu.22412
Grozeva D, Carss K, Spasic-Boskovic O et al (2015) Targeted Next-Generation Sequencing Analysis of 1,000 Individuals with Intellectual Disability. Hum Mutat 36:1197–1204. https://doi.org/10.1002/humu.22901
Guilmatre A, Dubourg C, Mosca A-L et al (2009) Recurrent rearrangements in synaptic and neurodevelopmental genes and shared biologic pathways in schizophrenia, autism, and mental retardation. Arch Gen Psychiatry 66:947–956. https://doi.org/10.1001/archgenpsychiatry.2009.80
Piton A, Gauthier J, Hamdan FF et al (2011) Systematic resequencing of X-chromosome synaptic genes in autism spectrum disorder and schizophrenia. Mol Psychiatry 16:867–880. https://doi.org/10.1038/mp.2010.54
Merad M, Sathe P, Helft J et al (2013) The dendritic cell lineage: ontogeny and function of dendritic cells and their subsets in the steady state and the inflamed setting. Annu Rev Immunol 31:563–604. https://doi.org/10.1146/annurev-immunol-020711-074950
Ahmed Z, Kawamura T, Shimada S, Piguet V (2015) The role of human dendritic cells in HIV-1 infection. J Invest Dermatol 135:1225–1233. https://doi.org/10.1038/jid.2014.490
Bertram KM, Tong O, Royle C et al (2019) Manipulation of mononuclear phagocytes by HIV: implications for early transmission events. Front Immunol 10:2263. https://doi.org/10.3389/fimmu.2019.02263
Wu L, KewalRamani VN (2006) Dendritic-cell interactions with HIV: infection and viral dissemination. Nat Rev Immunol 6:859–868. https://doi.org/10.1038/nri1960
Bracq L, Xie M, Benichou S, Bouchet J (2018) Mechanisms for cell-to-cell transmission of HIV-1. Front Immunol 9:260. https://doi.org/10.3389/fimmu.2018.00260
Kijewski SD, Gummuluru S (2015) A mechanistic overview of dendritic cell-mediated HIV-1transinfection: the story so far. Future Virol 10:257–269. https://doi.org/10.2217/fvl.15.2
Ménager MM, Littman DR (2016) Actin dynamics regulates dendritic cell-mediated transfer of HIV-1 to T cells. Cell 164:695–709. https://doi.org/10.1016/j.cell.2015.12.036
Wiley RD, Gummuluru S (2006) Immature dendritic cell-derived exosomes can mediate HIV-1 trans infection. Proc Natl Acad Sci USA 103:738–743. https://doi.org/10.1073/pnas.0507995103
Perot BP, García-Paredes V, Luka M, Ménager MM (2020) Dendritic cell maturation regulates TSPAN7 function in HIV-1 transfer to CD4 + T lymphocytes. Front Cell Infect Microbiol 10:70. https://doi.org/10.3389/fcimb.2020.00070
Haller C, Müller B, Fritz JV et al (2014) HIV-1 Nef and Vpu are functionally redundant broad-spectrum modulators of cell surface receptors, including tetraspanins. J Virol 88:14241–14257. https://doi.org/10.1128/JVI.02333-14
Kammula EC, Mötter J, Gorgels A et al (2012) Brain transcriptome-wide screen for HIV-1 Nef protein interaction partners reveals various membrane-associated proteins. PLoS ONE 7:e51578. https://doi.org/10.1371/journal.pone.0051578
Sato K, Aoki J, Misawa N et al (2008) Modulation of human immunodeficiency virus type 1 infectivity through incorporation of tetraspanin proteins. J Virol 82:1021–1033. https://doi.org/10.1128/JVI.01044-07
Wang L, Liu L, Che Y et al (2010) Egress of HSV-1 capsid requires the interaction of VP26 and a cellular tetraspanin membrane protein. Virol J 7:156. https://doi.org/10.1186/1743-422X-7-156
McDonald D (2010) Dendritic cells and HIV-1 trans-infection. Viruses 2:1704–1717. https://doi.org/10.3390/v2081704
Kwon J-O, Lee YD, Kim H et al (2016) Tetraspanin 7 regulates sealing zone formation and the bone-resorbing activity of osteoclasts. Biochem Biophys Res Commun 477:1078–1084. https://doi.org/10.1016/j.bbrc.2016.07.046
Jurdic P, Saltel F, Chabadel A, Destaing O (2006) Podosome and sealing zone: specificity of the osteoclast model. Eur J Cell Biol 85:195–202. https://doi.org/10.1016/j.ejcb.2005.09.008
Burrack AL, Martinov T, Fife BT (2017) T cell-mediated beta cell destruction: autoimmunity and alloimmunity in the context of type 1 diabetes. Front Endocrinol (Lausanne) 8:343. https://doi.org/10.3389/fendo.2017.00343
Hald J, Galbo T, Rescan C et al (2012) Pancreatic islet and progenitor cell surface markers with cell sorting potential. Diabetologia 55:154–165. https://doi.org/10.1007/s00125-011-2295-1
McLaughlin KA, Richardson CC, Ravishankar A et al (2016) Identification of tetraspanin-7 as a target of autoantibodies in type 1 diabetes. Diabetes 65:1690–1698. https://doi.org/10.2337/db15-1058
Walther D, Eugster A, Jergens S et al (2016) Tetraspanin 7 autoantibodies in type 1 diabetes. Diabetologia 59:1973–1976. https://doi.org/10.1007/s00125-016-3997-1
Eugster A, Kraus G, Lidzba V et al (2019) Cytoplasmic ends of tetraspanin 7 harbour epitopes recognised by autoantibodies in type 1 diabetes. Diabetologia 62:805–810. https://doi.org/10.1007/s00125-019-4832-2
McLaughlin KA, Tombs MA, Christie MR (2020) Autoimmunity to tetraspanin-7 in type 1 diabetes. Med Microbiol Immunol. https://doi.org/10.1007/s00430-020-00674-2
Thurner L, Müller A, Cérutti M et al (2011) Wegener’s granuloma harbors B lymphocytes with specificities against a proinflammatory transmembrane protein and a tetraspanin. J Autoimmun 36:87–90. https://doi.org/10.1016/j.jaut.2010.09.002
Ji G, Liang H, Wang F et al (2019) TSPAN12 precedes tumor proliferation by cell cycle control in ovarian cancer. Mol Cells 42:557–567. https://doi.org/10.14348/molcells.2019.0015
Vences-Catalán F, Levy S (2018) Immune targeting of tetraspanins involved in cell invasion and metastasis. Front Immunol 9:1277. https://doi.org/10.3389/fimmu.2018.01277
Hemler ME (2014) Tetraspanin proteins promote multiple cancer stages. Nat Rev Cancer 14:49–60. https://doi.org/10.1038/nrc3640
Schaper F, van Spriel AB (2018) antitumor immunity is controlled by tetraspanin proteins. Front Immunol 9:1185. https://doi.org/10.3389/fimmu.2018.01185
Cowan AJ, Allen C, Barac A et al (2018) Global burden of multiple myeloma: a systematic analysis for the global burden of disease study 2016. JAMA Oncol 4:1221–1227. https://doi.org/10.1001/jamaoncol.2018.2128
Cheong CM, Chow AWS, Fitter S et al (2015) Tetraspanin 7 (TSPAN7) expression is upregulated in multiple myeloma patients and inhibits myeloma tumour development in vivo. Exp Cell Res 332:24–38. https://doi.org/10.1016/j.yexcr.2015.01.006
Chakraborty S (2014) In silico analysis identifies genes common between five primary gastrointestinal cancer sites with potential clinical applications. Ann Gastroenterol 27:231–236
Takagi S, Fujikawa K, Imai T et al (1995) Identification of a highly specific surface marker of T-cell acute lymphoblastic leukemia and neuroblastoma as a new member of the transmembrane 4 superfamily. Int J Cancer 61:706–715. https://doi.org/10.1002/ijc.2910610519
Wuttig D, Baier B, Fuessel S et al (2009) Gene signatures of pulmonary metastases of renal cell carcinoma reflect the disease-free interval and the number of metastases per patient. Int J Cancer 125:474–482. https://doi.org/10.1002/ijc.24353
Wuttig D, Zastrow S, Füssel S et al (2012) CD31, EDNRB and TSPAN7 are promising prognostic markers in clear-cell renal cell carcinoma revealed by genome-wide expression analyses of primary tumors and metastases. Int J Cancer 131:E693–E704. https://doi.org/10.1002/ijc.27419
Nichols L, Saunders R, Knollmann FD (2012) Causes of death of patients with lung cancer. Arch Pathol Lab Med 136:1552–1557. https://doi.org/10.5858/arpa.2011-0521-OA
Wang X, Lin M, Zhao J et al (2018) TSPAN7 promotes the migration and proliferation of lung cancer cells via epithelial-to-mesenchymal transition. Onco Targets Ther 11:8815–8822. https://doi.org/10.2147/OTT.S167902
Xiao D, He J (2010) Epithelial mesenchymal transition and lung cancer. J Thorac Dis 2:154–159. https://doi.org/10.3978/j.issn.2072-1439.2010.02.03.7
Pérez-Bercoff A, Makino T, McLysaght A (2010) Duplicability of self-interacting human genes. BMC Evol Biol 10:160. https://doi.org/10.1186/1471-2148-10-160
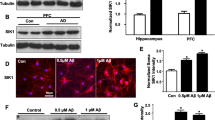
Salas IH, Callaerts-Vegh Z, Arranz AM et al (2017) Tetraspanin 6: a novel regulator of hippocampal synaptic transmission and long term plasticity. PLoS ONE 12:e0171968. https://doi.org/10.1371/journal.pone.0171968
Scheltens P, Blennow K, Breteler MMB et al (2016) Alzheimer’s disease. Lancet 388:505–517. https://doi.org/10.1016/S0140-6736(15)01124-1
Seipold L, Saftig P (2016) The emerging role of tetraspanins in the proteolytic processing of the amyloid precursor protein. Front Mol Neurosci 9:149. https://doi.org/10.3389/fnmol.2016.00149
Blalock EM, Geddes JW, Chen KC et al (2004) Incipient Alzheimer’s disease: microarray correlation analyses reveal major transcriptional and tumor suppressor responses. Proc Natl Acad Sci USA 101:2173–2178. https://doi.org/10.1073/pnas.0308512100
Bossers K, Wirz KTS, Meerhoff GF et al (2010) Concerted changes in transcripts in the prefrontal cortex precede neuropathology in Alzheimer’s disease. Brain 133:3699–3723. https://doi.org/10.1093/brain/awq258
Miller JA, Woltjer RL, Goodenbour JM et al (2013) Genes and pathways underlying regional and cell type changes in Alzheimer’s disease. Genome Med 5:48. https://doi.org/10.1186/gm452
Guix FX, Sannerud R, Berditchevski F et al (2017) Tetraspanin 6: a pivotal protein of the multiple vesicular body determining exosome release and lysosomal degradation of amyloid precursor protein fragments. Mol Neurodegener 12:25. https://doi.org/10.1186/s13024-017-0165-0
Roh JS, Sohn DH (2018) Damage-associated molecular patterns in inflammatory diseases. Immune Netw 18:e27. https://doi.org/10.4110/in.2018.18.e27
Medzhitov R (2008) Origin and physiological roles of inflammation. Nature 454:428–435. https://doi.org/10.1038/nature07201
Barton GM (2008) A calculated response: control of inflammation by the innate immune system. J Clin Investig 118:413–420. https://doi.org/10.1172/JCI34431
Leifer CA, Medvedev AE (2016) Molecular mechanisms of regulation of Toll-like receptor signaling. J Leukoc Biol 100:927–941. https://doi.org/10.1189/jlb.2MR0316-117RR
Mogensen TH (2009) Pathogen recognition and inflammatory signaling in innate immune defenses. Clin Microbiol Rev 22:240–273. https://doi.org/10.1128/CMR.00046-08
Loo Y-M, Gale M (2011) Immune signaling by RIG-I-like receptors. Immunity 34:680–692. https://doi.org/10.1016/j.immuni.2011.05.003
Saiz ML, Rocha-Perugini V, Sánchez-Madrid F (2018) Tetraspanins as organizers of antigen-presenting cell function. Front Immunol 9:1074. https://doi.org/10.3389/fimmu.2018.01074
Wang Y, Tong X, Omoregie ES et al (2012) Tetraspanin 6 (TSPAN6) negatively regulates retinoic acid-inducible gene I-like receptor-mediated immune signaling in a ubiquitination-dependent manner. J Biol Chem 287:34626–34634. https://doi.org/10.1074/jbc.M112.390401
Funding
Salaries were supported by INSERM (MMM) and ATIP-Avenir INSERM (BPP).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of commercial or financial relationships that may lead to a potential conflict of interest.
Additional information
Edited by Luise Florin.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Special Issue on Tetraspanins in Infection and Immunity.
Rights and permissions
About this article
Cite this article
Perot, B.P., Ménager, M.M. Tetraspanin 7 and its closest paralog tetraspanin 6: membrane organizers with key functions in brain development, viral infection, innate immunity, diabetes and cancer. Med Microbiol Immunol 209, 427–436 (2020). https://doi.org/10.1007/s00430-020-00681-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00430-020-00681-3