Abstract
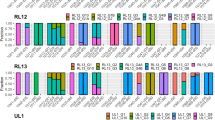
Precise identification of patients at highest risk for developing Cytomegalovirus (CMV) DNAemia may improve CMV infection management in the allogeneic hematopoietic stem cell transplantation (allo-HSCT) setting. Here, we studied the potential use of detecting free CMV micro(mi)RNAs circulating in plasma for predicting CMV DNAemia in this clinical scenario. A total of 62 adult allo-HSCT recipients were included in this prospective observational multicenter study. Plasma CMV DNA load was monitored using the CMV RealTime CMV PCR (Abbott Molecular, Des Plaines, IL, USA). Detection of mature CMV miRNAs in plasma drawn by days + 7, + 14 and + 30 after allo-HSCT was performed using the miScript PCR System (Qiagen, Hilden, Germany). Assays could be optimized for five out of the seven targeted CMV miRNAs: UL36-5p, US33-5p, UL148D, UL22A-5p and UL112-3p. Of the 62 patients included in the study, 42 developed a first episode of CMV DNAemia at a median of 35 days after allo-HSCT. All targeted CMV miRNA were detected early after transplantation, with CMV miRNA US33-5p and UL112-3p the most commonly found species at any time point; nevertheless, neither the detection rate of CMV miRNAs nor their abundance allowed discrimination between patients with subsequent CMV DNAemia and those with no CMV DNAemia. The data presented herein do not support any predictive utility of these CMV miRNAs for first episodes of CMV DNAemia in a cohort consisting primarily of allo-HSCT patients receiving haploidentical allografts.
Similar content being viewed by others
References
Romero PP, Blanco P, Giménez E, Solano C, Navarro D (2015) An update on the management and prevention of Cytomegalovirus infection following allogeneic hematopoietic stem cell transplantation. Future Virol 10:113–134
Chemaly RF, Ullmann AJ, Stoelben S, Richard MP, Bornhäuser M, Groth C, Einsele H, Silverman M, Mullane KM, Brown J, Nowak H, Kölling K, Stobernack HP, Lischka P, Zimmermann H, Rübsamen-Schaeff H, Champlin RE, Ehninger G (2014) Letermovir for Cytomegalovirus prophylaxis in hematopoietic-cell transplantation. New Engl J Med 370:1781–1789
Marty FM, Ljungman P, Chemaly RF, Maertens J, Dadwal SS, Duarte RF, Haider S, Ullmann AJ, Katayama Y, Brown J, Mullane KM, Boeckh M, Blumberg EA, Einsele H, Snydman DR, Kanda Y, DiNubile MJ, Teal VL, Wan H, Murata Y, Kartsonis NA, Leavitt RY, Badshah C (2017) Letermovir prophylaxis for Cytomegalovirus in hematopoietic-cell transplantation. New Engl J Med 377:2433–2444
Filipowicz W, Bhattacharyya Sonenberg N (2008) Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet 9:102–114
Ha M, Kim VN (2014) Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol 15:509–524
Diggins NL, Hancock MH (2018) HCMV miRNA targets reveal important cellular pathways for viral replication, latency, and reactivation. Noncoding RNA 4(4):29
Ng KR, Li JY, Gleadle JM (2015) Human Cytomegalovirus encoded microRNAs: hitting targets. Expert Rev Anti Infect Ther 13:1469–1479
Hook L, Hancock M, Landais I, Grabski R, Britt W, Nelson JA (2014) Cytomegalovirus microRNAs. Curr Opin Virol 7:40–46
Grey F (2015) Role of microRNAs in herpesvirus latency and persistence. J Gen Virol 96:739–751
Lisboa LF, Egli A, O’Shea D, Åsberg A, Hartmann A, Rollag H, Pang XL, Tyrrell DL, Kumar D, Humar A (2015) Hcmv-miR-UL22A-5p: a biomarker in transplantation with broad impact on host gene expression and potential immunological implications. Am J Transplant 15:1893–1902
Kawano Y, Kawada J, Kamiya Y, Suzuki M, Torii Y, Kimura H, Ito Y (2016) Analysis of circulating human and viral microRNAs in patients with congenital Cytomegalovirus infection. J Perinatol 36:1101–1105
Reddehase MJ, Balthesen M, Rapp M, Jonjić S, Pavić I, Koszinowski UH (1994) The conditions of primary infection define the load of latent viral genome in organs and the risk of recurrent Cytomegalovirus disease. J Exp Med 179:185–193
Sacher T, Andrassy J, Kalnins A, Dölken L, Jordan S, Podlech J, Ruzsics Z, Jauch K-W, Reddehase MJ, Koszinowski UH (2011) Shedding light on the elusive role of endothelial cells in Cytomegalovirus dissemination. PLoS Pathog 7:e1002366
Grey F, Antoniewicz A, Allen E, Saugstad J, McShea A, Carrington JC, Nelson J (2005) Identification and characterization of human Cytomegalovirus-encoded microRNAs. J Virol 79:12095–12099
Clari MA, Bravo D, Costa E, Munoz-Cobo B, Solano C, Remigia MJ, Gimenez E, Benmarzouk-Hidalgo OJ, Perez-Romero P, Navarro D (2013) Comparison of the new Abbott Real Time CMV assay and the Abbott CMV PCR Kit for the quantitation of plasma Cytomegalovirus DNAemia. Diagn Microbiol Infect Dis 75:207–209
Glucksberg H, Storb R, Fefer A, Buckner C, Neiman P, Clift R, Lerner K, Thomas E (1974) Clinical manifestations of graft-versus-host disease in human recipients of marrow from HL-A-matched sibling donors. Transplantation 18:295–304
Talaya A, Solano C, Giménez E, García Giménez JL, Vinuesa V, Alberola J, Pallardó FV, Navarro D (2017) Assessing the risk of Cytomegalovirus DNAaemia in allogeneic stem cell transplant recipients by monitoring oxidative-stress markers in plasma. J Gen Virol 98:1855–1863
Monleón D, Talaya A, Giménez E, Vinuesa V, Morales JM, Hernández-Boluda JC, Pérez A, Piñana JL, Solano C, Navarro D (2018) Validation of a plasma metabolomics model that allows anticipation of the occurrence of Cytomegalovirus DNAaemia in allogeneic stem cell transplant recipients. J Med Microbiol. https://doi.org/10.1099/jmm.0.000746
Talaya A, Giménez E, Vinuesa V, Pérez A, Amat P, Piñana JL, Albert E, Hernández-Boluda JC, Solano C, Navarro D (2019) Kinetics of inflammatory biomarkers in plasma predict the occurrence and features of Cytomegalovirus DNAemia episodes in allogeneic hematopoietic stem cell transplant recipients. Med Microbiol Immunol 1:2. https://doi.org/10.1007/s00430-019-00594-w
Stark TJ, Arnold JD, Spector DH, Yeo GW (2012) High-resolution profiling and analysis of viral and host small RNAs during human Cytomegalovirus infection. J Virol 86:226–235
Dunn W, Chou C, Li H, Hai R, Patterson D, Stolc V, Zhu H, Liu F (2003) Functional profiling of a human Cytomegalovirus genome. Proc Natl Acad Sci USA 100:14223–14228
Dunn W, Trang P, Zhong Q, Yang E, van Belle C, Liu F (2005) Human Cytomegalovirus expresses novel microRNAs during productive viral infection. Cell Microbiol 7:1684–1695
Stern-Ginossar N, Elefant N, Zimmermann A, Wolf DG, Saleh N, Biton M, Horwitz E, Prokocimer Z, Prichard M, Hahn G, Goldman-Wohl D, Greenfield C, Yagel S, Hengel H, Altuvia Y, Margalit H, Mandelboim O (2007) Host immune system gene targeting by a viral miRNA. Science 317:376–381
Stern-Ginossar N, Saleh N, Goldberg MD, Prichard M, Wolf DG, Mandelboim O (2009) Analysis of human Cytomegalovirus-encoded microRNA activity during infection. J Virol 83:10684–10693
Murphy E, Vanicek J, Robins H, Shenk T, Levine AJ (2008) Suppression of immediate-early viral gene expression by herpesvirus-coded microRNAs: implications for latency. Proc Natl Acad Sci USA 105:5453–5458
Cromer D, Tey SK, Khanna R, Davenport MP (2013) Estimating Cytomegalovirus growth rates by using only a single point. J Virol 87:3376–3381
Huang X, Yuan T, Tschannen M, Sun Z, Jacob H, Du M, Liang M, Dittmar RL, Liu Y, Liang M, Kohli M, Thibodeau SN, Boardman L, Wang L (2013) Characterization of human plasma-derived exosomal RNAs by deep sequencing. BMC Genomics 14:319
Valadi H, Ekstrom K, Bossios A, Sjostrand M, Lee JJ, Lotvall JO (2007) Exosome mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol 9:654–659
Acknowledgements
Estela Giménez holds a Juan Rodés research contract from the Carlos III Health Institute. Eliseo Albert holds a Río Ortega research contract from the Carlos III Health Institute.
Funding
This work was supported by a Fondo de Investigaciones Sanitarias (FIS) grant from Carlos III Health Institute (15/0060).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Edited by: Matthias J. Reddehase.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Talaya, A., Giménez, E., Pascual, M.J. et al. An investigation of the utility of plasma Cytomegalovirus (CMV) microRNA detection to predict CMV DNAemia in allogeneic hematopoietic stem cell transplant recipients. Med Microbiol Immunol 209, 15–21 (2020). https://doi.org/10.1007/s00430-019-00632-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00430-019-00632-7