Abstract
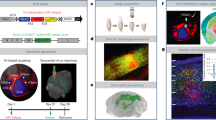
The impressive functions of the brain rely on an extensive connectivity matrix between specific neurons, the architecture of which is frequently characterized by one brain nucleus/region connecting to multiple targets, either via collaterals of the same projection neuron or several, differentially specified neurons. Delineating the fine architecture of projection neuron subsets in a specific brain region could greatly facilitate its circuit, computational, and functional resolution. Here, we developed multiple fluorescent rabies viruses (RV) to delineate the fine organization of corticothalamic projection neuron subsets in the primary visual cortex (V1). By simultaneously retrograde labeling multiple distinct subsets of corticothalamic projection neurons in V1 from their target nuclei in thalamus (dLGN, LP, LD), we observed that V1-dLGN corticothalamic projection neurons were densely concentrated in layer VI, except for several sparsely scattered neurons in layer V, while V1-LP and V1-LD corticothalamic projection neurons were localized to both layers V and VI. Meanwhile, we observed a fraction of V1 corticothalamic projection neurons targeting two thalamic nuclei, which was further confirmed by fMOST whole-brain imaging. The multiple fluorescent RV tracing tools can be extensively applied to resolve the architecture of projection neuron subsets in certain brain regions, with a strong potential to delineate the computational and functional organization of these brain regions.




Similar content being viewed by others
Availability of data and material
The authors confirm that the data supporting the findings of this study are available within the article and the supplementary materials.
References
Barbas H (2015) General cortical and special prefrontal connections: principles from structure to function. Annu Rev Neurosci 38:269–289. https://doi.org/10.1146/annurev-neuro-071714-033936
Betley JN, Sternson SM (2011) Adeno-associated viral vectors for mapping, monitoring, and manipulating neural circuits. Hum Gene Ther 22:669–677. https://doi.org/10.1089/hum.2010.204
Broussard GJ, Liang R, Tian L (2014) Monitoring activity in neural circuits with genetically encoded indicators. Front Mol Neurosci 7:97. https://doi.org/10.3389/fnmol.2014.00097
Calhoon GG, Tye KM (2015) Resolving the neural circuits of anxiety. Nat Neurosci 18:1394–1404. https://doi.org/10.1038/nn.4101
Callaway EM (2008) Transneuronal circuit tracing with neurotropic viruses. Curr Opin Neurobiol 18:617–623. https://doi.org/10.1016/j.conb.2009.03.007
Callaway EM, Luo L (2015) Monosynaptic Circuit Tracing with Glycoprotein-Deleted Rabies Viruses. J Neurosci 35:8979–8985. https://doi.org/10.1523/JNEUROSCI.0409-15.2015
Cetin A, Komai S, Eliava M, Seeburg PH, Osten P (2006) Stereotaxic gene delivery in the rodent brain. Nat Protoc 1:3166–3173. https://doi.org/10.1038/nprot.2006.450
Chatterjee S et al (2018) Nontoxic, double-deletion-mutant rabies viral vectors for retrograde targeting of projection neurons. Nat Neurosci 21:638–646. https://doi.org/10.1038/s41593-018-0091-7
Ciabatti E, Gonzalez-Rueda A, Mariotti L, Morgese F, Tripodi M (2017) Life-long genetic and functional access to neural circuits using self-inactivating rabies virus. Cell 170:382–392. https://doi.org/10.1016/j.cell.2017.06.014
Diao YP et al (2018) Reciprocal connections between cortex and thalamus contribute to retinal axon targeting to dorsal lateral geniculate nucleus. Cereb Cortex 28:1168–1182. https://doi.org/10.1093/cercor/bhx028
Dulka BN, Bress KS, Grizzell JA, Cooper MA (2018) Social dominance modulates stress-induced neural activity in medial prefrontal cortex projections to the basolateral amygdala. Neuroscience 388:274–283. https://doi.org/10.1016/j.neuroscience.2018.07.042
Ginger M, Haberl M, Conzelmann K-K, Schwarz MK, Frick A (2013) Revealing the secrets of neuronal circuits with recombinant rabies virus technology. Front Neural Circuits 7:2–2. https://doi.org/10.3389/fncir.2013.00002
Gong H et al (2016) High-throughput dual-colour precision imaging for brain-wide connectome with cytoarchitectonic landmarks at the cellular level. Nat Commun 7:12142. https://doi.org/10.1038/ncomms12142
Gong H et al (2013) Continuously tracing brain-wide long-distance axonal projections in mice at a one-micron voxel resolution. Neuroimage 74:87–98. https://doi.org/10.1016/j.neuroimage.2013.02.005
Harris KD, Shepherd GM (2015) The neocortical circuit: themes and variations. Nat Neurosci 18:170–181. https://doi.org/10.1038/nn.3917
Hasse JM, Briggs F (2017) Corticogeniculate feedback sharpens the temporal precision and spatial resolution of visual signals in the ferret. Proc Natl Acad Sci U S A 114:E6222–E6230. https://doi.org/10.1073/pnas.1704524114
Janak PH, Tye KM (2015) From circuits to behaviour in the amygdala. Nature 517:284–292. https://doi.org/10.1038/nature14188
Jorntell H (2013) Neural circuits in movement control. J Physiol 591:5421–5423. https://doi.org/10.1113/jphysiol.2013.265603
Kim EJ, Jacobs MW, Ito-Cole T, Callaway EM (2016) Improved monosynaptic neural circuit tracing using engineered rabies virus glycoproteins. Cell reports 15:692–699. https://doi.org/10.1016/j.celrep.2016.03.067
Kim J, Matney CJ, Blankenship A, Hestrin S, Brown SP (2014) Layer 6 corticothalamic neurons activate a cortical output layer, layer 5a. J Neurosci 34:9656–9664. https://doi.org/10.1523/JNEUROSCI.1325-14.2014
Kitamura T et al (2017) Engrams and circuits crucial for systems consolidation of a memory. Science 356:73–78. https://doi.org/10.1126/science.aam6808
Lanciego JL, Wouterlood FG (2020) Neuroanatomical tract-tracing techniques that did go viral. Brain Struct Funct 225:1193–1224. https://doi.org/10.1007/s00429-020-02041-6
Mu D et al (2017) A central neural circuit for itch sensation. Science 357:695–699. https://doi.org/10.1126/science.aaf4918
Naumann T, Härtig W, Frotscher M (2000) Retrograde tracing with Fluoro-Gold: different methods of tracer detection at the ultrastructural level and neurodegenerative changes of back-filled neurons in long-term studies. J Neurosci Methods 103:11–21. https://doi.org/10.1016/s0165-0270(00)00292-2
Oh SW et al (2014) A mesoscale connectome of the mouse brain. Nature 508:207–214. https://doi.org/10.1038/nature13186
Osakada F, Callaway EM (2013) Design and generation of recombinant rabies virus vectors. Nat Protoc 8:1583–1601. https://doi.org/10.1038/nprot.2013.094
Osakada F, Mori T, Cetin AH, Marshel JH, Virgen B, Callaway EM (2011) New rabies virus variants for monitoring and manipulating activity and gene expression in defined neural circuits. Neuron 71:617–631. https://doi.org/10.1016/j.neuron.2011.07.005
Park HG, Carmel JB (2016) Selective manipulation of neural circuits. Neurotherapeutics 13:311–324. https://doi.org/10.1007/s13311-016-0425-7
Peirs C, Seal RP (2016) Neural circuits for pain: Recent advances and current views. Science 354:578–584. https://doi.org/10.1126/science.aaf8933
Peltékian E, Garcia L, Danos O (2002) Neurotropism and retrograde axonal transport of a canine adenoviral vector: a tool for targeting key structures undergoing neurodegenerative processes. Mol Ther 5:25–32. https://doi.org/10.1006/mthe.2001.0517
Petersen CC (2014) Cortical control of whisker movement. Annu Rev Neurosci 37:183–203. https://doi.org/10.1146/annurev-neuro-062012-170344
Pollak Dorocic I et al (2014) A whole-brain atlas of inputs to serotonergic neurons of the dorsal and median raphe nuclei. Neuron 83:663–678. https://doi.org/10.1016/j.neuron.2014.07.002
Reardon TR et al (2016) Rabies Virus CVS-N2c(Delta G) strain enhances retrograde synaptic transfer and neuronal viability. Neuron 89:711–724. https://doi.org/10.1016/j.neuron.2016.01.004
Ren J et al (2018) Anatomically defined and functionally distinct dorsal raphe serotonin sub-systems. Cell 175:472–487. https://doi.org/10.1016/j.cell.2018.07.043
Sarno E, Robison AJ (2018) Emerging role of viral vectors for circuit-specific gene interrogation and manipulation in rodent brain. Pharmacol Biochem Behav 174:2–8. https://doi.org/10.1016/j.pbb.2018.04.008
Saunders A, Sabatini BL (2015) Cre activated and inactivated recombinant adeno-associated viral vectors for neuronal anatomical tracing or activity manipulation. Curr Protoc Neurosci 72:1241–12415. https://doi.org/10.1002/0471142301.ns0124s72
Scammell TE, Arrigoni E, Lipton JO (2017) Neural circuitry of wakefulness and sleep. Neuron 93:747–765. https://doi.org/10.1016/j.neuron.2017.01.014
Schwarz LA et al (2015) Viral-genetic tracing of the input-output organization of a central noradrenaline circuit. Nature 524:88–92. https://doi.org/10.1038/nature14600
Silva BA, Gross CT, Graff J (2016) The neural circuits of innate fear: detection, integration, action, and memorization. Learn Mem 23:544–555. https://doi.org/10.1101/lm.042812.116
Sun L et al (2019) Differences in neurotropism and neurotoxicity among retrograde viral tracers. Mol Neurodegener 14:8. https://doi.org/10.1186/s13024-019-0308-6
Suzuki T, Morimoto N, Akaike A, Osakada F (2020) Multiplex neural circuit tracing with G-deleted rabies viral vectors. Front Neural Circuits 13:77. https://doi.org/10.3389/fncir.2019.00077
Tasic B et al (2016) Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat Neurosci 19:335–346. https://doi.org/10.1038/nn.4216
Tervo DGR et al (2016) A Designer AAV variant permits efficient retrograde access to projection neurons. Neuron 92:372–382. https://doi.org/10.1016/j.neuron.2016.09.021
Ugolini G (2010) Advances in viral transneuronal tracing. J Neurosci Methods 194:2–20. https://doi.org/10.1016/j.jneumeth.2009.12.001
Wickersham IR, Finke S, Conzelmann KK, Callaway EM (2007a) Retrograde neuronal tracing with a deletion-mutant rabies virus. Nat Methods 4:47–49. https://doi.org/10.1038/nmeth999
Wickersham IR et al (2007b) Monosynaptic restriction of transsynaptic tracing from single, genetically targeted neurons. Neuron 53:639–647. https://doi.org/10.1016/j.neuron.2007.01.033
Xiong H et al (2014) Chemical reactivation of quenched fluorescent protein molecules enables resin-embedded fluorescence microimaging. Nat Commun 5:3992. https://doi.org/10.1038/ncomms4992
Acknowledgement
We thank Dr. Fuqiang Xu (Wuhan Institute of Physics and Mathematics, Chinese Academy of Sciences) and Dr. Edward Callaway (The SALK Institute, USA) for rRV packing system, and Dr. Xiaobin He and Ting Ding (Wuhan Institute of Physics and Mathematics, Chinese Academy of Sciences) for suggestions related to the RV preparation. This work was supported by the National Natural Science Foundation of China (Grant Nos. 31700934, 91632110, 31371106) and the Huazhong Agricultural University Scientific & Technological Self-innovation Foundation (Program Nos. 2662017PY082, 52204-13002).
Funding
This work was supported by the National Natural Science Foundation of China (Grant Nos. 31700934, 91632110, 31371106) and the Huazhong Agricultural University Scientific & Technological Self-innovation Foundation (Program Nos. 2662017PY082, 52204–13002).
Author information
Authors and Affiliations
Contributions
GC, JD, LL, and YT conceived and designed the project. LL, YT, and XS prepared the viruses and performed most of the neural tracing, imaging, and data analysis work. LS, JY, and SZ contributed to the development of figures. HG, XZ and XL performed fMOST imaging. ZH performed confocal microscopy imaging. ZF, HW and KR helped edit the manuscript. GC, JD, LL and YT wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics approval
All animal care procedures and experiments were approved by the Research Ethics Committee, Huazhong Agricultural University, Hubei, China (HZAUMO-2016–021) and carried out in accordance with the Guide for the Care and Use of Laboratory Animals from Research Ethics Committee, Huazhong Agricultural University.
Consent to participate
Not applicable.
Consent for publication
The authors confirm that the work described here has not been published or under consideration in any other journal. The authors agree to publication in the journal of brain structure and function by Springer.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file1 (WMV 27037 KB)
Rights and permissions
About this article
Cite this article
Li, L., Tang, Y., Sun, L. et al. Delineating the organization of projection neuron subsets in primary visual cortex with multiple fluorescent rabies virus tracing. Brain Struct Funct 226, 951–961 (2021). https://doi.org/10.1007/s00429-021-02250-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00429-021-02250-7