Abstract
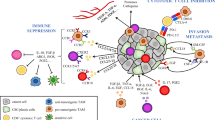
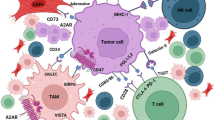
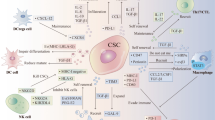
It has been for long conceived that hallmarks of cancer were intrinsic genetic features driving tumor development, proliferation, and progression, and that targeting such cell-autonomous pathways could be sufficient to achieve therapeutic cancer control. Clinical ex vivo data demonstrated that treatment efficacy often relied on the contribution of host immune responses, hence introducing the concept of tumor microenvironment (TME), namely the existence, along with tumor cells, of non-tumor components that could significantly influence tumor growth and survival. Among the complex network of TME-driving forces, immunity plays a key role and the balance between antitumor and protumor immune responses is a major driver in contrasting or promoting cancer spreading. TME is usually a very immunosuppressed milieu because of a vast array of local alterations contrasting antitumor adaptive immunity, where metabolic changes contribute to cancer dissemination by impairing T cell infiltration and favoring the accrual and activation of regulatory cells. Subcellular structures known as extracellular vesicles then help spreading immunosuppression at systemic levels by distributing genetic and protein tumor repertoire in distant tissues. A major improvement in the knowledge of TME is now pointing the attention back to tumor cells; indeed, recent findings are showing how oncogenic pathways and specific mutations in tumor cells can actually dictate the nature and the function of immune infiltrate. As our information on the reciprocal interactions regulating TME increases, finding a strategy to interfere with TME crosstalk becomes more complex and challenging. Nevertheless, TME interactions represent a promising field for the discovery of novel biomarkers and therapeutic targets for improving treatment efficacy in cancer.

Similar content being viewed by others
References
Nagarajan A, Malvi P, Wajapeyee N (2016) Oncogene-directed alterations in cancer cell metabolism. Trends Cancer 7:365–377
Huber V, Camisaschi C, Berzi A, Ferro S, Lugini L, Triulzi T, Tuccitto A, Tagliabue E, Castelli C, Rivoltini L (2017) Cancer acidity: an ultimate frontier of tumor immune escape and a novel target of immunomodulation. Semin Cancer Biol 43:74–89
Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang TH, Porta-Pardo E, Gao GF, Plaisier CL, Eddy JA, Ziv E, Culhane AC, Paull EO, IKA S, Gentles AJ, Malhotra R, Farshidfar F, Colaprico A, Parker JS, Mose LE, Vo NS, Liu J, Liu Y, Rader J, Dhankani V, Reynolds SM, Bowlby R, Califano A, Cherniack AD, Anastassiou D, Bedognetti D, Rao A, Chen K, Krasnitz A, Hu H, Malta TM, Noushmehr H, Pedamallu CS, Bullman S, Ojesina AI, Lamb A, Zhou W, Shen H, Choueiri TK, Weinstein JN, Guinney J, Saltz J, Holt RA, Rabkin CE, Cancer Genome Atlas Research Network, Lazar AJ, Serody JS, Demicco EG, Disis ML, Vincent BG, Shmulevich L (2018) The immune landscape of cancer. Immunity 48:812–830.e14
Wellenstein MD, de Visser KE (2018) Cancer-cell-intrinsic mechanisms shaping the tumor immune landscape. Immunity 48:399–416
Gainor JF, Shaw AT, Sequist LV, Fu X, Azzoli CG, Piotrowska Z, Huynh TG, Zhao L, Fulton L, Schultz KR, Howe E, Farago AF, Sullivan RJ, Stone JR, Digumarthy S, Moran T, Hata AN, Yagi Y, Yeap BY, Engelman JA, Mino-Kenudson M (2016) EGFR mutations and ALK rearrangements are associated with low response rates to PD-1 pathway blockade in non-small cell lung cancer: a retrospective analysis. Clin Cancer Res 22:4585–4593
Shin DS, Zaretsky JM, Escuin-Ordinas H, Garcia-Diaz A, Hu-Lieskovan S, Kalbasi A, Grasso CS, Hugo W, Sandoval S, Torrejon DY, Palaskas N, Rodriguez GA, Parisi G, Azhdam A, Chmielowski B, Cherry G, Seja E, Berent-Maoz B, Shintaku IP, Le DT, Pardoll DM, Diaz LA,Jr, Tumeh PC, Graeber TG, Lo RS, Comin-Anduix B, Ribas A (2017) Primary resistance to PD-1 blockade mediated by JAK1/2 mutations. Cancer Discov 7:188–201
Valenti R, Huber V, Filipazzi P, Pilla L, Sovena G, Villa A, Corbelli A, Fais S, Parmiani G, Rivoltini L (2006) Human tumor-released microvesicles promote the differentiation of myeloid cells with transforming growth factor-beta-mediated suppressive activity on T lymphocytes. Cancer Res 66:9290–9298
Condamine T, Mastio J, Gabrilovich DI (2015) Transcriptional regulation of myeloid-derived suppressor cells. J Leukoc Biol 98:913–922
Rivera LB, Bergers G (2015) Intertwined regulation of angiogenesis and immunity by myeloid cells. Trends Immunol 36:240–249
Zhang S, Ma X, Zhu C, Liu L, Wang G, Yuan X (2016) The role of myeloid-derived suppressor cells in patients with solid tumors: a meta-analysis. PLoS One 11:e0164514ù
Weber R, Fleming V, Hu X, Nagibin V, Groth C, Altevogt P, Utikal J, Umansky V (2018) Myeloid-derived suppressor cells hinder the anti-cancer activity of immune checkpoint inhibitors. Front Immunol 9:1310
Reddy SM, Reuben A, Wargo JA (2016) Influences of BRAF inhibitors on the immune microenvironment and the rationale for combined molecular and immune targeted therapy. Curr Oncol Rep 18:42-016-0531-z
Wilmott JS, Long GV, Howle JR, Haydu LE, Sharma RN, Thompson JF, Kefford RF, Hersey P, Scolyer RA (2012) Selective BRAF inhibitors induce marked T-cell infiltration into human metastatic melanoma. Clin Cancer Res 18:1386–1394
Frederick DT, Piris A, Cogdill AP, Cooper ZA, Lezcano C, Ferrone CR, Mitra D, Boni A, Newton LP, Liu C, Peng W, Sullivan RJ, Lawrence DP, Hodi FS, Overwijk WW, Lizee G, Murphy GF, Hwu P, Flaherty KT, Fisher DE, Wargo JA (2013) BRAF inhibition is associated with enhanced melanoma antigen expression and a more favorable tumor microenvironment in patients with metastatic melanoma. Clin Cancer Res 19:1225–1231
Kakavand H, Wilmott JS, Menzies AM, Vilain R, Haydu LE, Yearley JH, Thompson JF, Kefford RF, Hersey P, Long GV, Scolyer RA (2015) PD-L1 expression and tumor-infiltrating lymphocytes define different subsets of MAPK inhibitor-treated melanoma patients. Clin Cancer Res 21:3140–3148
Vergani E, Di Guardo L, Dugo M, Rigoletto S, Tragni G, Ruggeri R, Perrone F, Tamborini E, Gloghini A, Arienti F, Vergani B, Deho P, De Cecco L, Vallacchi V, Frati P, Shahaj E, Villa A, Santinami M, De Braud F, Rivoltini L, Rodolfo M (2016) Overcoming melanoma resistance to vemurafenib by targeting CCL2-induced miR-34a, miR-100 and miR-125b. Oncotarget 7:4428–4441
Zakiryanova GK, Wheeler S, Shurin MR (2018) Oncogenes in immune cells as potential therapeutic targets. Immunotargets Ther 7:21–28
Zakiryanova GK, Kustova E, Urazalieva NT, Amirbekov A, Baimuchametov ET, Nakisbekov NN, Shurin MR (2017) Alterations of oncogenes expression in NK cells in patients with cancer. Immun Inflamm Dis 5:493–502
Pello OM, De Pizzol M, Mirolo M, Soucek L, Zammataro L, Amabile A, Doni A, Nebuloni M, Swigart LB, Evan GI, Mantovani A, Locati M (2012) Role of c-MYC in alternative activation of human macrophages and tumor-associated macrophage biology. Blood 119:411–421
Li L, Ng DS, Mah WC, Almeida FF, Rahmat SA, Rao VK, Leow SC, Laudisi F, Peh MT, Goh AM, Lim JS, Wright GD, Mortellaro A, Taneja R, Ginhoux F, Lee CG, Moore PK, Lane DP (2015) A unique role for p53 in the regulation of M2 macrophage polarization. Cell Death Differ 22:1081–1093
Madapura HS, Salamon D, Wiman KG, Lain S, Klein E, Nagy N (2016) cMyc-p53 feedback mechanism regulates the dynamics of T lymphocytes in the immune response. Cell Cycle 15:1267–1275
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, Hoang CD, Diehn M, Alizadeh AA (2015) Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 12:453–457
Van Allen EM, Miao D, Schilling B, Shukla SA, Blank C, Zimmer L, Sucker A, Hillen U, MHG F, Goldinger SM, Utikal J, Hassel JC, Weide B, Kaehler KC, Loquai C, Mohr P, Gutzmer R, Dummer R, Gabriel S, Wu CJ, Schadendorf D, Garraway LA (2015) Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350:207–211
Yarchoan M, Hopkins A, Jaffee EM (2017) Tumor mutational burden and response rate to PD-1 inhibition. N Engl J Med 377:2500–2501
Turajlic S, Litchfield K, Xu H, Rosenthal R, McGranahan N, Reading JL, Wong YNS, Rowan A, Kanu N, Al Bakir M, Chambers T, Salgado R, Savas P, Loi S, Birkbak NJ, Sansregret L, Gore M, Larkin J, Quezada SA, Swanton C (2017) Insertion-and-deletion-derived tumour-specific neoantigens and the immunogenic phenotype: a pan-cancer analysis. Lancet Oncol 18:1009–1021
Le DT UJN, Wang H, Bartlett BR, Kemberling H, Eyring AD, Skora AD, Luber BS, Azad NS, Laheru D, Biedrzycki B, Donehower RC, Zaheer A, Fisher GA, Crocenzi TS, Lee JJ, Duffy SM, Goldberg RM, de la Chapelle A, Koshiji M, Bhaijee F, Huebner T, Hruban RH, Wood LD, Cuka N, Pardoll DM, Papadopoulos N, Kinzler KW, Zhou S, Cornish TC, Taube JM, Anders RA, Eshleman JR, Vogelstein B, Diaz LA Jr (2015) PD-1 blockade in tumors with mismatch-repair deficiency. N Engl J Med 372:2509–2520
Hugo W, Zaretsky JM, Sun L, Song C, Moreno BH, Hu-Lieskovan S, Berent-Maoz B, Pang J, Chmielowski B, Cherry G, Seja E, Lomeli S, Kong X, Kelley MC, Sosman JA, Johnson DB, Ribas A, Lo RS (2016) Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165:35–44
Biton J, Mansuet-Lupo A, Pecuchet N, Alifano M, Ouakrim H, Arrondeau J, Boudou-Rouquette P, Goldwasser F, Leroy K, Goc J, Wislez M, Germain C, Laurent-Puig P, Dieu-Nosjean MC, Cremer I, Herbst R, Blons HF, Damotte D (2018) TP53, STK11 and EGFR mutations predict tumor immune profile and the response to anti-PD-1 in lung adenocarcinoma. Clin Cancer Res
Davoli T, Uno H, Wooten EC, Elledge SJ (2017) Tumor aneuploidy correlates with markers of immune evasion and with reduced response to immunotherapy. Science 355:aaf8399
Roh W, Chen PL, Reuben A, Spencer CN, Prieto PA, Miller JP, Gopalakrishnan V, Wang F, Cooper ZA, Reddy SM, Gumbs C, Little L, Chang Q, Chen WS, Wani K, De Macedo MP, Chen E, Austin-Breneman JL, Jiang H, Roszik J, Tetzlaff MT, Davies MA, Gershenwald JE, Tawbi H, Lazar AJ, Hwu P, Hwu WJ, Diab A, Glitza IC, Patel SP, Woodman SE, Amaria RN, Prieto VG, Hu J, Sharma P, Allison JP, Chin L, Zhang J, Wargo JA, Futreal PA (2017) Integrated molecular analysis of tumor biopsies on sequential CTLA-4 and PD-1 blockade reveals markers of response and resistance. Sci Transl Med 9:eaah3560
Gao J, Shi LZ, Zhao H, Chen J, Xiong L, He Q, Chen T, Roszik J, Bernatchez C, Woodman SE, Chen PL, Hwu P, Allison JP, Futreal A, Wargo JA, Sharma P (2016) Loss of IFN-gamma pathway genes in tumor cells as a mechanism of resistance to anti-CTLA-4 therapy. Cell 167:397–404.e9
Skoulidis F, Goldberg ME, Greenawalt DM, Hellmann MD, Awad MM, Gainor JF, Schrock AB, Hartmaier RJ, Trabucco SE, Gay L, Ali SM, Elvin JA, Singal G, Ross JS, Fabrizio D, Szabo PM, Chang H, Sasson A, Srinivasan S, Kirov S, Szustakowski J, Vitazka P, Edwards R, Bufill JA, Sharma N, Ou SI, Peled N, Spigel DR, Rizvi H, Aguilar EJ, Carter BW, Erasmus J, Halpenny DF, Plodkowski AJ, Long NM, Nishino M, Denning WL, Galan-Cobo A, Hamdi H, Hirz T, Tong P, Wang J, Rodriguez-Canales J, Villalobos PA, Parra ER, Kalhor N, Sholl LM, Sauter JL, Jungbluth AA, Mino-Kenudson M, Azimi R, Elamin YY, Zhang J, Leonardi GC, Jiang F, Wong KK, Lee JJ, Papadimitrakopoulou VA, Wistuba II, Miller VA, Frampton GM, Wolchok JD, Shaw AT, Janne PA, Stephens PJ, Rudin CM, Geese WJ, Albacker LA, Heymach JV (2018) STK11/LKB1 mutations and PD-1 inhibitor resistance in KRAS-mutant lung adenocarcinoma. Cancer Discov 8:822–835
Mlecnik B, Bindea G, Angell HK, Sasso MS, Obenauf AC, Fredriksen T, Lafontaine L, Bilocq AM, Kirilovsky A, Tosolini M, Waldner M, Berger A, Fridman WH, Rafii A, Valge-Archer V, Pages F, Speicher MR, Galon J (2014) Functional network pipeline reveals genetic determinants associated with in situ lymphocyte proliferation and survival of cancer patients. Sci Transl Med 6:228ra37
Linsley PS, Speake C, Whalen E, Chaussabel D (2014) Copy number loss of the interferon gene cluster in melanomas is linked to reduced T cell infiltrate and poor patient prognosis. PLoS One 9:e109760
Garcia-Diaz A, Shin DS, Moreno BH, Saco J, Escuin-Ordinas H, Rodriguez GA, Zaretsky JM, Sun L, Hugo W, Wang X, Parisi G, Saus CP, Torrejon DY, Graeber TG, Comin-Anduix B, Hu-Lieskovan S, Damoiseaux R, Lo RS, Ribas A (2017) Interferon receptor signaling pathways regulating PD-L1 and PD-L2 expression. Cell Rep 19:1189–1201
Benci JL, Xu B, Qiu Y, Wu TJ, Dada H, Twyman-Saint Victor C, Cucolo L, Lee DSM, Pauken KE, Huang AC, Gangadhar TC, Amaravadi RK, Schuchter LM, Feldman MD, Ishwaran H, Vonderheide RH, Maity A, Wherry EJ, Minn AJ (2016) Tumor interferon signaling regulates a multigenic resistance program to immune checkpoint blockade. Cell 167:1540–1554.e12
Goodman AM, Kato S, Bazhenova L, Patel SP, Frampton GM, Miller V, Stephens PJ, Daniels GA, Kurzrock R (2017) Tumor mutational burden as an independent predictor of response to immunotherapy in diverse cancers. Mol Cancer Ther 16:2598–2608
Charoentong P, Finotello F, Angelova M, Mayer C, Efremova M, Rieder D, Hackl H, Trajanoski Z (2017) Pan-cancer immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep 18:248–262
Cancer Genome Atlas Network (2015) Genomic classification of cutaneous melanoma. Cell 161:1681–1696
Snyder A, Makarov V, Merghoub T, Yuan J, Zaretsky JM, Desrichard A, Walsh LA, Postow MA, Wong P, Ho TS, Hollmann TJ, Bruggeman C, Kannan K, Li Y, Elipenahli C, Liu C, Harbison CT, Wang L, Ribas A, Wolchok JD, Chan TA (2014) Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med 371:2189–2199
McGranahan N, Furness AJ, Rosenthal R, Ramskov S, Lyngaa R, Saini SK, Jamal-Hanjani M, Wilson GA, Birkbak NJ, Hiley CT, Watkins TB, Shafi S, Murugaesu N, Mitter R, Akarca AU, Linares J, Marafioti T, Henry JY, Van Allen EM, Miao D, Schilling B, Schadendorf D, Garraway LA, Makarov V, Rizvi NA, Snyder A, Hellmann MD, Merghoub T, Wolchok JD, Shukla SA, Wu CJ, Peggs KS, Chan TA, Hadrup SR, Quezada SA, Swanton C (2016) Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science 351:1463–1469
Lauss M, Donia M, Harbst K, Andersen R, Mitra S, Rosengren F, Salim M, Vallon-Christersson J, Torngren T, Kvist A, Ringner M, Svane IM, Jonsson G (2017) Mutational and putative neoantigen load predict clinical benefit of adoptive T cell therapy in melanoma. Nat Commun 8:1738-017-01460-0
Zehir A, Benayed R, Shah RH, Syed A, Middha S, Kim HR, Srinivasan P, Gao J, Chakravarty D, Devlin SM, Hellmann MD, Barron DA, Schram AM, Hameed M, Dogan S, Ross DS, Hechtman JF, DF DL, Yao J, Mandelker DL, Cheng DT, Chandramohan R, Mohanty AS, Ptashkin RN, Jayakumaran G, Prasad M, Syed MH, Rema AB, Liu ZY, Nafa K, Borsu L, Sadowska J, Casanova J, Bacares R, Kiecka IJ, Razumova A, Son JB, Stewart L, Baldi T, Mullaney KA, Al-Ahmadie H, Vakiani E, Abeshouse AA, Penson AV, Jonsson P, Camacho N, Chang MT, Won HH, Gross BE, Kundra R, Heins ZJ, Chen HW, Phillips S, Zhang H, Wang J, Ochoa A, Wills J, Eubank M, Thomas SB, Gardos SM, Reales DN, Galle J, Durany R, Cambria R, Abida W, Cercek A, Feldman DR, Gounder MM, Hakimi AA, Harding JJ, Iyer G, Janjigian YY, Jordan EJ, Kelly CM, Lowery MA, LGT M, Omuro AM, Raj N, Razavi P, Shoushtari AN, Shukla N, Soumerai TE, Varghese AM, Yaeger R, Coleman J, Bochner B, Riely GJ, Saltz LB, Scher HI, Sabbatini PJ, Robson ME, Klimstra DS, Taylor BS, Baselga J, Schultz N, Hyman DM, Arcila ME, Solit DB, Ladanyi M, Berger MF (2017) Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat Med 23:703–713
Liu G, Zhan X, Dong C, Liu L (2017) Genomics alterations of metastatic and primary tissues across 15 cancer types. Sci Rep 7:13262-017-13650-3
Grellety T, Lucchesi C, Hostein I, Auzanneau C, Khalifa E, Soubeyran I, Italiano A (2017) High-depth sequencing of paired primary and metastatic tumours: Implications for personalised medicine. Eur J Cancer 84:250–256
Garber K (2018) Blood test may predict cancer immunotherapy benefit. Science 360:1387
Eil R, Vodnala SK, Clever D, Klebanoff CA, Sukumar M, Pan JH, Palmer DC, Gros A, Yamamoto TN, Patel SJ, Guittard GC, Yu Z, Carbonaro V, Okkenhaug K, Schrump DS, Linehan WM, Roychoudhuri R, Restifo NP (2016) Ionic immune suppression within the tumour microenvironment limits T cell effector function. Nature 537:539–543
Baenke F, Chaneton B, Smith M, Van Den Broek N, Hogan K, Tang H, Viros A, Martin M, Galbraith L, Girotti MR, Dhomen N, Gottlieb E, Marais R (2016) Resistance to BRAF inhibitors induces glutamine dependency in melanoma cells. Mol Oncol 10:73–84
Gao P, Tchernyshyov I, Chang TC, Lee YS, Kita K, Ochi T, Zeller KI, De Marzo AM, Van Eyk JE, Mendell JT, Dang CV (2009) c-Myc suppression of miR-23a/b enhances mitochondrial glutaminase expression and glutamine metabolism. Nature 458:762–765
Mullen AR, Hu Z, Shi X, Jiang L, Boroughs LK, Kovacs Z, Boriack R, Rakheja D, Sullivan LB, Linehan WM, Chandel NS, DeBerardinis RJ (2014) Oxidation of alpha-ketoglutarate is required for reductive carboxylation in cancer cells with mitochondrial defects. Cell Rep 7:1679–1690
Porstmann T, Santos CR, Griffiths B, Cully M, Wu M, Leevers S, Griffiths JR, Chung YL, Schulze A (2008) SREBP activity is regulated by mTORC1 and contributes to Akt-dependent cell growth. Cell Metab 8:224–236
Tyrakis PA, Palazon A, Macias D, Lee KL, Phan AT, Velica P, You J, Chia GS, Sim J, Doedens A, Abelanet A, Evans CE, Griffiths JR, Poellinger L, Goldrath AW, Johnson RS (2016) S-2-hydroxyglutarate regulates CD8(+) T-lymphocyte fate. Nature 540:236–241
Noman MZ, Desantis G, Janji B, Hasmim M, Karray S, Dessen P, Bronte V, Chouaib S (2014) PD-L1 is a novel direct target of HIF-1alpha, and its blockade under hypoxia enhanced MDSC-mediated T cell activation. J Exp Med 211:781–790
Calcinotto A, Filipazzi P, Grioni M, Iero M, De Milito A, Ricupito A, Cova A, Canese R, Jachetti E, Rossetti M, Huber V, Parmiani G, Generoso L, Santinami M, Borghi M, Fais S, Bellone M, Rivoltini L (2012) Modulation of microenvironment acidity reverses anergy in human and murine tumor-infiltrating T lymphocytes. Cancer Res 72:2746–2756
Clambey ET, McNamee EN, Westrich JA, Glover LE, Campbell EL, Jedlicka P, de Zoeten EF, Cambier JC, Stenmark KR, Colgan SP, Eltzschig HK (2012) Hypoxia-inducible factor-1 alpha-dependent induction of FoxP3 drives regulatory T-cell abundance and function during inflammatory hypoxia of the mucosa. Proc Natl Acad Sci U S A 109:E2784–E2793
Angelin A, Gil-de-Gomez L, Dahiya S, Jiao J, Guo L, Levine MH, Wang Z, Quinn WJ 3rd, Kopinski PK, Wang L, Akimova T, Liu Y, Bhatti TR, Han R, Laskin BL, Baur JA, Blair IA, Wallace DC, Hancock WW, Beier UH (2017) Foxp3 reprograms T cell metabolism to function in low-glucose, high-lactate environments. Cell Metab 25:1282–1293.e7
Facciabene A, Peng X, Hagemann IS, Balint K, Barchetti A, Wang LP, Gimotty PA, Gilks CB, Lal P, Zhang L, Coukos G (2011) Tumour hypoxia promotes tolerance and angiogenesis via CCL28 and T(reg) cells. Nature 475:226–230
Sceneay J, Chow MT, Chen A, Halse HM, Wong CS, Andrews DM, Sloan EK, Parker BS, Bowtell DD, Smyth MJ, Moller A (2012) Primary tumor hypoxia recruits CD11b+/Ly6Cmed/Ly6G+ immune suppressor cells and compromises NK cell cytotoxicity in the premetastatic niche. Cancer Res 72:3906–3911
Li J, Wang L, Chen X, Li L, Li Y, Ping Y, Huang L, Yue D, Zhang Z, Wang F, Li F, Yang L, Huang J, Yang S, Li H, Zhao X, Dong W, Yan Y, Zhao S, Huang B, Zhang B, Zhang Y (2017) CD39/CD73 upregulation on myeloid-derived suppressor cells via TGF-beta-mTOR-HIF-1 signaling in patients with non-small cell lung cancer. Oncoimmunology 6:e1320011
Movahedi K, Laoui D, Gysemans C, Baeten M, Stange G, Van den Bossche J, Mack M, Pipeleers D, In’t Veld P, De Baetselier P, Van Ginderachter JA (2010) Different tumor microenvironments contain functionally distinct subsets of macrophages derived from Ly6C(high) monocytes. Cancer Res 70:5728–5739
Laoui D, Van Overmeire E, Di Conza G, Aldeni C, Keirsse J, Morias Y, Movahedi K, Houbracken I, Schouppe E, Elkrim Y, Karroum O, Jordan B, Carmeliet P, Gysemans C, De Baetselier P, Mazzone M, Van Ginderachter JA (2014) Tumor hypoxia does not drive differentiation of tumor-associated macrophages but rather fine-tunes the M2-like macrophage population. Cancer Res 74:24–30
Colegio OR, Chu NQ, Szabo AL, Chu T, Rhebergen AM, Jairam V, Cyrus N, Brokowski CE, Eisenbarth SC, Phillips GM, Cline GW, Phillips AJ, Medzhitov R (2014) Functional polarization of tumour-associated macrophages by tumour-derived lactic acid. Nature 513:559–563
Kuchuk O, Tuccitto A, Citterio D, Huber V, Camisaschi C, Milione M, Vergani B, Villa A, Alison MR, Carradori S, Supuran CT, Rivoltini L, Castelli C, Mazzaferro V (2018) pH regulators to target the tumor immune microenvironment in human hepatocellular carcinoma. Oncoimmunology 7:e1445452
Mendler AN, Hu B, Prinz PU, Kreutz M, Gottfried E, Noessner E (2012) Tumor lactic acidosis suppresses CTL function by inhibition of p38 and JNK/c-Jun activation. Int J Cancer 131:633–640
Chang CH, Qiu J, O'Sullivan D, Buck MD, Noguchi T, Curtis JD, Chen Q, Gindin M, Gubin MM, van der Windt GJ, Tonc E, Schreiber RD, Pearce EJ, Pearce EL (2015) Metabolic competition in the tumor microenvironment is a driver of cancer progression. Cell 162:1229–1241
Ho PC, Bihuniak JD, Macintyre AN, Staron M, Liu X, Amezquita R, Tsui YC, Cui G, Micevic G, Perales JC, Kleinstein SH, Abel ED, Insogna KL, Feske S, Locasale JW, Bosenberg MW, Rathmell JC, Kaech SM (2015) Phosphoenolpyruvate is a metabolic checkpoint of anti-tumor T cell responses. Cell 162:1217–1228
Yang L, Achreja A, Yeung TL, Mangala LS, Jiang D, Han C, Baddour J, Marini JC, Ni J, Nakahara R, Wahlig S, Chiba L, Kim SH, Morse J, Pradeep S, Nagaraja AS, Haemmerle M, Kyunghee N, Derichsweiler M, Plackemeier T, Mercado-Uribe I, Lopez-Berestein G, Moss T, Ram PT, Liu J, Lu X, Mok SC, Sood AK, Nagrath D (2016) Targeting stromal glutamine synthetase in tumors disrupts tumor microenvironment-regulated cancer cell growth. Cell Metab 24:685–700
Choi J, Stradmann-Bellinghausen B, Yakubov E, Savaskan NE, Regnier-Vigouroux A (2015) Glioblastoma cells induce differential glutamatergic gene expressions in human tumor-associated microglia/macrophages and monocytes-derived macrophages. Cancer Biol Ther 16:1205–1213
Pan M, Reid MA, Lowman XH, Kulkarni RP, Tran TQ, Liu X, Yang Y, Hernandez-Davies JE, Rosales KK, Li H, Hugo W, Song C, Xu X, Schones DE, Ann DK, Gradinaru V, Lo RS, Locasale JW, Kong M (2016) Regional glutamine deficiency in tumours promotes dedifferentiation through inhibition of histone demethylation. Nat Cell Biol 18:1090–1101
Liu PS, Wang H, Li X, Chao T, Teav T, Christen S, Di Conza G, Cheng WC, Chou CH, Vavakova M, Muret C, Debackere K, Mazzone M, Huang HD, Fendt SM, Ivanisevic J, Ho PC (2017) Alpha-ketoglutarate orchestrates macrophage activation through metabolic and epigenetic reprogramming. Nat Immunol 18:985–994
Zhao E, Maj T, Kryczek I, Li W, Wu K, Zhao L, Wei S, Crespo J, Wan S, Vatan L, Szeliga W, Shao I, Wang Y, Liu Y, Varambally S, Chinnaiyan AM, Welling TH, Marquez V, Kotarski J, Wang H, Wang Z, Zhang Y, Liu R, Wang G, Zou W (2016) Cancer mediates effector T cell dysfunction by targeting microRNAs and EZH2 via glycolysis restriction. Nat Immunol 17:95–103
Raposo G, Stoorvogel W (2013) Extracellular vesicles: exosomes, microvesicles, and friends. J Cell Biol 200:373–383
Tkach M, Thery C (2016) Communication by extracellular vesicles: where we are and where we need to go. Cell 164:1226–1232
Whiteside TL (2016) Exosomes and tumor-mediated immune suppression. J Clin Invest 126:1216–1223
Willms E, Johansson HJ, Mager I, Lee Y, Blomberg KE, Sadik M, Alaarg A, Smith CI, Lehtio J, El Andaloussi S, Wood MJ, Vader P (2016) Cells release subpopulations of exosomes with distinct molecular and biological properties. Sci Rep 6:22519
Willms E, Cabanas C, Mager I, Wood MJA, Vader P (2018) Extracellular vesicle heterogeneity: subpopulations, isolation techniques, and diverse functions in cancer progression. Front Immunol 9:738
Chen G, Huang AC, Zhang W, Zhang G, Wu M, Xu W, Yu Z, Yang J, Wang B, Sun H, Xia H, Man Q, Zhong W, Antelo LF, Wu B, Xiong X, Liu X, Guan L, Li T, Liu S, Yang R, Lu Y, Dong L, McGettigan S, Somasundaram R, Radhakrishnan R, Mills G, Lu Y, Kim J, Chen YH, Dong H, Zhao Y, Karakousis GC, Mitchell TC, Schuchter LM, Herlyn M, Wherry EJ, Xu X, Guo W (2018) Exosomal PD-L1 contributes to immunosuppression and is associated with anti-PD-1 response. Nature 560:382–386
Caivano A, Del Vecchio L, Musto P (2017) Do we need to distinguish exosomes from microvesicles in hematological malignancies? Leukemia 31:2009–2010
Huber V, Vallacchi V, Fleming V, Hu X, Cova A, Dugo M, Shahaj E, Sulsenti R, Vergani E, Filipazzi P, De Laurentiis A, Lalli L, Di Guardo L, Patuzzo R, Vergani B, Casiraghi E, Cossa M, Gualeni A, Bollati V, Arienti F, De Braud F, Mariani L, Villa A, Altevogt P, Umansky V, Rodolfo M, Rivoltini L (2018) Tumor-derived microRNAs induce myeloid suppressor cells and predict immunotherapy resistance in melanoma. J Clin Invest
Camisaschi C, Vallacchi V, Vergani E, Tazzari M, Ferro S, Tuccitto A, Kuchuk O, Shahaj E, Sulsenti R, Castelli C, Rodolfo M, Rivoltini L, Huber V (2016) Targeting immune regulatory networks to counteract immune suppression in cancer. Vaccines (Basel) 4:e38.
Zhou M, Chen J, Zhou L, Chen W, Ding G, Cao L (2014) Pancreatic cancer derived exosomes regulate the expression of TLR4 in dendritic cells via miR-203. Cell Immunol 292:65–69
Shen Y, Guo D, Weng L, Wang S, Ma Z, Yang Y, Wang P, Wang J, Cai Z (2017) Tumor-derived exosomes educate dendritic cells to promote tumor metastasis via HSP72/HSP105-TLR2/TLR4 pathway. Oncoimmunology 6:e1362527
Gobbo J, Marcion G, Cordonnier M, Dias AMM, Pernet N, Hammann A, Richaud S, Mjahed H, Isambert N, Clausse V, Rebe C, Bertaut A, Goussot V, Lirussi F, Ghiringhelli F, de Thonel A, Fumoleau P, Seigneuric R, Garrido C (2015) Restoring anticancer immune response by targeting tumor-derived exosomes with a HSP70 peptide aptamer. J Natl Cancer Inst 108:djv330
Diao J, Yang X, Song X, Chen S, He Y, Wang Q, Chen G, Luo C, Wu X, Zhang Y (2015) Exosomal Hsp70 mediates immunosuppressive activity of the myeloid-derived suppressor cells via phosphorylation of Stat3. Med Oncol 32:453-014-0453-2
Wang J, De Veirman K, Faict S, Frassanito MA, Ribatti D, Vacca A, Menu E (2016) Multiple myeloma exosomes establish a favourable bone marrow microenvironment with enhanced angiogenesis and immunosuppression. J Pathol 239:162–173
Ham S, Lima LG, EPZ C, Muller A, Lobb RJ, Krumeich S, Wen SW, Wiegmans AP, Moller A (2018) Breast cancer-derived exosomes alter macrophage polarization via gp130/STAT3 signaling. Front Immunol 9:871
Chow A, Zhou W, Liu L, Fong MY, Champer J, Van Haute D, Chin AR, Ren X, Gugiu BG, Meng Z, Huang W, Ngo V, Kortylewski M, Wang SE (2014) Macrophage immunomodulation by breast cancer-derived exosomes requires Toll-like receptor 2-mediated activation of NF-kappaB. Sci Rep 4:5750
Ying X, Wu Q, Wu X, Zhu Q, Wang X, Jiang L, Chen X, Wang X (2016) Epithelial ovarian cancer-secreted exosomal miR-222-3p induces polarization of tumor-associated macrophages. Oncotarget 7:43076–43087
Wang X, Luo G, Zhang K, Cao J, Huang C, Jiang T, Liu B, Su L, Qiu Z (2018) Hypoxic tumor-derived exosomal miR-301a mediates M2 macrophage polarization via PTEN/PI3Kgamma to promote pancreatic cancer metastasis. Cancer Res
Wang F, Li B, Wei Y, Zhao Y, Wang L, Zhang P, Yang J, He W, Chen H, Jiao Z, Li Y (2018) Tumor-derived exosomes induce PD-1(+) macrophage population in human gastric cancer that promotes disease progression. Oncogenesis 7:41-018-0049-3
Gabrusiewicz K, Li X, Wei J, Hashimoto Y, Marisetty AL, Ott M, Wang F, Hawke D, Yu J, Healy LM, Hossain A, Akers JC, Maiti SN, Yamashita S, Shimizu Y, Dunner K, Zal MA, Burks JK, Gumin J, Nwajei F, Rezavanian A, Zhou S, Rao G, Sawaya R, Fuller GN, Huse JT, Antel JP, Li S, Cooper L, Sulman EP, Chen C, Geula C, Kalluri R, Zal T, Heimberger AB (2018) Glioblastoma stem cell-derived exosomes induce M2 macrophages and PD-L1 expression on human monocytes. Oncoimmunology 7:e1412909
Challagundla KB, Wise PM, Neviani P, Chava H, Murtadha M, Xu T, Kennedy R, Ivan C, Zhang X, Vannini I, Fanini F, Amadori D, Calin GA, Hadjidaniel M, Shimada H, Jong A, Seeger RC, Asgharzadeh S, Goldkorn A, Fabbri M (2015) Exosome-mediated transfer of microRNAs within the tumor microenvironment and neuroblastoma resistance to chemotherapy. J Natl Cancer Inst 107:djv135
Mrizak D, Martin N, Barjon C, Jimenez-Pailhes AS, Mustapha R, Niki T, Guigay J, Pancre V, de Launoit Y, Busson P, Morales O, Delhem N (2014) Effect of nasopharyngeal carcinoma-derived exosomes on human regulatory T cells. J Natl Cancer Inst 107:363
Muller L, Mitsuhashi M, Simms P, Gooding WE, Whiteside TL (2016) Tumor-derived exosomes regulate expression of immune function-related genes in human T cell subsets. Sci Rep 6:20254
Yamada N, Kuranaga Y, Kumazaki M, Shinohara H, Taniguchi K, Akao Y (2016) Colorectal cancer cell-derived extracellular vesicles induce phenotypic alteration of T cells into tumor-growth supporting cells with transforming growth factor-beta1-mediated suppression. Oncotarget 7:27033–27043
Maybruck BT, Pfannenstiel LW, Diaz-Montero M, Gastman BR (2017) Tumor-derived exosomes induce CD8(+) T cell suppressors. J Immunother Cancer 5:65-017-0269-7
Wen SW, Sceneay J, Lima LG, Wong CS, Becker M, Krumeich S, Lobb RJ, Castillo V, Wong KN, Ellis S, Parker BS, Moller A (2016) The biodistribution and immune suppressive effects of breast cancer-derived exosomes. Cancer Res 76:6816–6827
Hong CS, Sharma P, Yerneni SS, Simms P, Jackson EK, Whiteside TL, Boyiadzis M (2017) Circulating exosomes carrying an immunosuppressive cargo interfere with cellular immunotherapy in acute myeloid leukemia. Sci Rep 7:14684-017-14661-w
Berchem G, Noman MZ, Bosseler M, Paggetti J, Baconnais S, Le Cam E, Nanbakhsh A, Moussay E, Mami-Chouaib F, Janji B, Chouaib S (2015) Hypoxic tumor-derived microvesicles negatively regulate NK cell function by a mechanism involving TGF-beta and miR23a transfer. Oncoimmunology 5:e1062968
Iero M, Valenti R, Huber V, Filipazzi P, Parmiani G, Fais S, Rivoltini L (2008) Tumour-released exosomes and their implications in cancer immunity. Cell Death Differ 15:80–88
Kumar B, Garcia M, Weng L, Jung X, Murakami JL, Hu X, McDonald T, Lin A, Kumar AR, DiGiusto DL, Stein AS, Pullarkat VA, Hui SK, Carlesso N, Kuo YH, Bhatia R, Marcucci G, Chen CC (2018) Acute myeloid leukemia transforms the bone marrow niche into a leukemia-permissive microenvironment through exosome secretion. Leukemia 32:575–587
Muller L, Simms P, Hong CS, Nishimura MI, Jackson EK, Watkins SC, Whiteside TL (2017) Human tumor-derived exosomes (TEX) regulate Treg functions via cell surface signaling rather than uptake mechanisms. Oncoimmunology 6:e1261243
Melo SA, Luecke LB, Kahlert C, Fernandez AF, Gammon ST, Kaye J, LeBleu VS, Mittendorf EA, Weitz J, Rahbari N, Reissfelder C, Pilarsky C, Fraga MF, Piwnica-Worms D, Kalluri R (2015) Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature 523:177–182
Huber V, Filipazzi P, Iero M, Fais S, Rivoltini L (2008) More insights into the immunosuppressive potential of tumor exosomes. J Transl Med 6:63-5876-6-63
Chen DS, Mellman I (2013) Oncology meets immunology: the cancer-immunity cycle. Immunity 39:1–10
Veglia F, Perego M, Gabrilovich D (2018) Myeloid-derived suppressor cells coming of age. Nat Immunol 19:108–119
Marvel D, Gabrilovich DI (2015) Myeloid-derived suppressor cells in the tumor microenvironment: expect the unexpected. J Clin Invest 125:3356–3364
Li B, Pozzi A, Young PP (2011) TNFalpha accelerates monocyte to endothelial transdifferentiation in tumors by the induction of integrin alpha5 expression and adhesion to fibronectin. Mol Cancer Res 9:702–711
Zhang H, Maric I, DiPrima MJ, Khan J, Orentas RJ, Kaplan RN, Mackall CL (2013) Fibrocytes represent a novel MDSC subset circulating in patients with metastatic cancer. Blood 122:1105–1113
Kumar V, Patel S, Tcyganov E, Gabrilovich DI (2016) The nature of myeloid-derived suppressor cells in the tumor microenvironment. Trends Immunol 37:208–220
Zeng Q, Fu J, Korrer M, Gorbounov M, Murray PJ, Pardoll D, Masica DL, Kim YJ (2018) Caspase-1 from human myeloid-derived suppressor cells can promote T cell-independent tumor proliferation. Cancer Immunol Res 6:566–577
Awad RM, De Vlaeminck Y, Maebe J, Goyvaerts C, Breckpot K (2018) Turn Back the TIMe: targeting tumor infiltrating myeloid cells to revert cancer progression. Front Immunol 9:1977
Rigoni A, Colombo MP, Pucillo C (2018) Mast cells, basophils and eosinophils: from allergy to cancer. Semin Immunol 35:29–34
Sharma P, Hu-Lieskovan S, Wargo JA, Ribas A (2017) Primary, adaptive, and acquired resistance to cancer immunotherapy. Cell 168:707–723
De Henau O, Rausch M, Winkler D, Campesato LF, Liu C, Cymerman DH, Budhu S, Ghosh A, Pink M, Tchaicha J, Douglas M, Tibbitts T, Sharma S, Proctor J, Kosmider N, White K, Stern H, Soglia J, Adams J, Palombella VJ, McGovern K, Kutok JL, Wolchok JD, Merghoub T (2016) Overcoming resistance to checkpoint blockade therapy by targeting PI3Kgamma in myeloid cells. Nature 539:443–447
Condamine T, Dominguez GA, Youn JI, Kossenkov AV, Mony S, Alicea-Torres K, Tcyganov E, Hashimoto A, Nefedova Y, Lin C, Partlova S, Garfall A, Vogl DT, Xu X, Knight SC, Malietzis G, Lee GH, Eruslanov E, Albelda SM, Wang X, Mehta JL, Bewtra M, Rustgi A, Hockstein N, Witt R, Masters G, Nam B, Smirnov D, Sepulveda MA, Gabrilovich DI (2016) Lectin-type oxidized LDL receptor-1 distinguishes population of human polymorphonuclear myeloid-derived suppressor cells in cancer patients. Sci Immunol 1:aaf8943.
Mezquita L, Auclin E, Ferrara R, Charrier M, Remon J, Planchard D, Ponce S, Ares LP, Leroy L, Audigier-Valette C, Felip E, Zeron-Medina J, Garrido P, Brosseau S, Zalcman G, Mazieres J, Caramela C, Lahmar J, Adam J, Chaput N, Soria JC, Besse B (2018) Association of the lung immune prognostic index with immune checkpoint inhibitor outcomes in patients with advanced non-small cell lung cancer. JAMA Oncol 4:351–357
Gooden MJ, de Bock GH, Leffers N, Daemen T, Nijman HW (2011) The prognostic influence of tumour-infiltrating lymphocytes in cancer: a systematic review with meta-analysis. Br J Cancer 105:93–103
Negorev D, Beier UH, Zhang T, Quatromoni JG, Bhojnagarwala P, Albelda SM, Singhal S, Eruslanov E, Lohoff FW, Levine MH, Diamond JM, Christie JD, Hancock WW, Akimova T (2018) Human neutrophils can mimic myeloid-derived suppressor cells (PMN-MDSC) and suppress microbead or lectin-induced T cell proliferation through artefactual mechanisms. Sci Rep 8:3135-018-21450-6
Fleming V, Hu X, Weber R, Nagibin V, Groth C, Altevogt P, Utikal J, Umansky V (2018) Targeting myeloid-derived suppressor cells to bypass tumor-induced immunosuppression. Front Immunol 9:398
Dominguez GA, Condamine T, Mony S, Hashimoto A, Wang F, Liu Q, Forero A, Bendell J, Witt R, Hockstein N, Kumar P, Gabrilovich DI (2017) Selective targeting of myeloid-derived suppressor cells in cancer patients using DS-8273a, an agonistic TRAIL-R2 antibody. Clin Cancer Res 23:2942–2950
Koinis F, Vetsika EK, Aggouraki D, Skalidaki E, Koutoulaki A, Gkioulmpasani M, Georgoulias V, Kotsakis A (2016) Effect of first-line treatment on myeloid-derived suppressor cells’ subpopulations in the peripheral blood of patients with non-small cell lung cancer. J Thorac Oncol 11:1263–1272
Annels NE, Shaw VE, Gabitass RF, Billingham L, Corrie P, Eatock M, Valle J, Smith D, Wadsley J, Cunningham D, Pandha H, Neoptolemos JP, Middleton G (2014) The effects of gemcitabine and capecitabine combination chemotherapy and of low-dose adjuvant GM-CSF on the levels of myeloid-derived suppressor cells in patients with advanced pancreatic cancer. Cancer Immunol Immunother 63:175–183
Ahn MJ (2018) Consolidation of immunotherapy becomes new standard of care in unresectable stage III non-small cell lung cancer. J Thorac Dis 10:1205–1206
Motzer RJ, Powles T, Atkins MB, Escudier B, McDermott DF, Suarez C, Bracarda S, Stadler WM, Donskov F, Lee JL, Hawkins RE, Ravaud A, Alekseev BY, Staehler MD, Uemura M, Donaldson F, Li S, Huseni MA, Schiff C, Rini BI (2018) IMmotion151: a randomized phase III study of atezolizumab plus bevacizumab vs sunitinib in untreated metastatic renal cell carcinoma (mRCC). J Clin Oncol 36:578
Acknowledgments
The authors are grateful to AIRC (5xMille 12162 to LR, IG-10727 and IG-14285 to LR, IG-17462 to MR; IG-15192 to CC), Cariplo Foundation (2015-0911 to VV), Harry J. Lloyd Charitable Trust for human studies (MR and LR), the European Union’s Horizon 2020 Research and Innovation Programme grant agreement No. 686089 (PRECIOUS), and the Italian Ministry of Health grant 52/RF-2010-2312620 to support all their studies.
Contribution statement
All the authors contributed in conception and design, writing and discussion of the present review.
Funding
This work was supported by grants from AIRC (5xMille 12162 to LR, IG-10727 and IG-14285 to LR, IG-17462 to MR; IG-15192 to CC) and from Cariplo Foundation (2015-0911 to VV), by awards from the Harry J. Lloyd Charitable Trust for human studies (MR and LR), by the European Union’s Horizon 2020 Research and Innovation Programme grant agreement No. 686089 (PRECIOUS), and by the Italian Ministry of Health grant 52/RF-2010-2312620.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Tuccitto, A., Shahaj, E., Vergani, E. et al. Immunosuppressive circuits in tumor microenvironment and their influence on cancer treatment efficacy. Virchows Arch 474, 407–420 (2019). https://doi.org/10.1007/s00428-018-2477-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-018-2477-z