Abstract
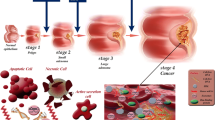
The field of genomics has shifted our view on disease development by providing insights in the molecular and functional processes encoded in the genome. In the case of cancer, many alterations in the DNA accumulate that enable tumor growth or even metastatic dissemination. Identification of molecular signatures that define different stages of progression towards cancer can enable early tumor detection. In this review, the impact of genomics will be addressed using early detection of colorectal cancer (CRC) as an example. Increased understanding of the adenoma-to-carcinoma progression has led to the discovery of several diagnostic biomarkers. This combined with technical advancements, has facilitated the development of molecular tests for non-invasive early CRC detection in stool and blood samples. Even though several tests have already made it to clinical practice, sensitivity and specificity for the detection of precancerous lesions still need improvement. Besides the diagnostic qualities, also the accuracy of the intermediate endpoint is an important issue on how the effectiveness of a novel test is perceived. Here, progression biomarkers may provide a more precise measure than the currently used morphologically based features. Similar developments in biomarker use for early detection have taken place in other cancer types.

Similar content being viewed by others
References
Forbes SA, Bhamra G, Bamford S et al (2008) The catalogue of somatic mutations in cancer (COSMIC). Curr Protoc Hum Genet Chapter 10:10.11
Muzny DM, Bainbridge MN, Chang K et al (2012) Comprehensive molecular characterization of human colon and rectal cancer. Nature 487:330–337
Hanahan D, Weinberg RA, Adams JM et al (2011) Hallmarks of cancer: the next generation. Cell 144:646–674
Wilson JMG, Jungner G (1968) Principles and practice of screening for disease. WHO. Available from http://www.who.int/bulletin/volumes/86/4/07-050112bp.pdf, Geneva Accessed 25 January 2017
Siegel R, Naishadham D, Jemal A (2012) Cancer statistics, 2012. CA Cancer J Clin 62:10–29
Gray JAM, Patnick J, Blanks RG (2008) Maximizing benefit and minimizing harm of screening. BMJ 336:480–483
IJspeert JEG, Vermeulen L, Meijer GA, Dekker E (2015) Serrated neoplasia-role in colorectal carcinogenesis and clinical implications. Nat Rev Gastroenterol Hepatol 12:401–409
Muto T, Bussey HJ, Morson BC (1975) The evolution of cancer of the colon and rectum. Cancer 36:2251–2270
Lieberman DA, Weiss DG, Bond JH et al (2000) Use of colonoscopy to screen asymptomatic adults for colorectal cancer. Veterans Affairs Cooperative Study Group 380. N Engl J Med 343:162–168
Imperiale TF, Wagner DR, Lin CY et al (2000) Risk of advanced proximal neoplasms in asymptomatic adults according to the distal colorectal findings. N Engl J Med 343:169–174
Shinya H, Wolff WI (1979) Morphology, anatomic distribution and cancer potential of colonic polyps. Ann Surg 190:679–683
Groden J, Thliveris A, Samowitz W et al (1991) Identification and characterization of the familial adenomatous polyposis coli gene. Cell 66:589–600
Drost J, van Jaarsveld RH, Ponsioen B et al (2015) Sequential cancer mutations in cultured human intestinal stem cells. Nature 521:43–47
Fearon ER (2011) Molecular genetics of colorectal cancer. Annu Rev Pathol Mech Dis 6:479–507
Fearon ER, Vogelstein B (1990) A genetic model for colorectal tumorigenesis. Cell 61:759–767
Derks S, Postma C, Moerkerk PTM et al (2006) Promoter methylation precedes chromosomal alterations in colorectal cancer development. Cell Oncol 28:247–257
Sook Kim M, Lee J, Sidransky D (2010) DNA methylation markers in colorectal cancer. Cancer Metastasis Rev 29:181–206
Lengauer C, Kinzler KW, Vogelstein B (1997) Genetic instability in colorectal cancers. Nature 386:623–627
Barber TD, McManus K, Yuen KWY et al (2008) Chromatid cohesion defects may underlie chromosome instability in human colorectal cancers. Proc Natl Acad Sci U S A 105:3443–3448
Rajagopalan H, Nowak MA, Vogelstein B, Lengauer C (2003) The significance of unstable chromosomes in colorectal cancer. Nat Rev Cancer 3:695–701
Ried T, Knutzen R, Steinbeck R et al (1996) Comparative genomic hybridization reveals a specific pattern of chromosomal gains and losses during the genesis of colorectal tumors. Genes Chromosom Cancer 15:234–245
Meijer GA, Hermsen MA, Baak JP et al Progression from colorectal adenoma to carcinoma is associated with non- random chromosomal gains as detected by comparative genomic hybridisation. J Clin Pathol 51:901–909
Douglas EJ, Fiegler H, Rowan A et al (2004) Array comparative genomic hybridization analysis of colorectal cancer cell lines and primary carcinomas. Cancer Res 64:4817–4825
Hermsen M, Postma C, Baak J et al (2002) Colorectal adenoma to carcinoma progression follows multiple pathways of chromosomal instability. Gastroenterology 123:1109–1119
Carvalho B, Postma C, Mongera S et al (2009) Multiple putative oncogenes at the chromosome 20q amplicon contribute to colorectal adenoma to carcinoma progression. Gut 58:79–89
de Groen FLM, Krijgsman O, Tijssen M et al (2014) Gene-dosage dependent overexpression at the 13q amplicon identifies DIS3 as candidate oncogene in colorectal cancer progression. Genes Chromosom Cancer 53:339–348
Camps J, Pitt JJ, Emons G et al (2013) Genetic amplification of the NOTCH modulator LNX2 upregulates the WNT/ -catenin pathway in colorectal cancer. Cancer Res 73:2003–2013
Firestein R, Bass AJ, Kim SY et al (2008) CDK8 is a colorectal cancer oncogene that regulates β-catenin activity. Nature 455:547–551
Sillars-Hardebol AH, Carvalho B, Tijssen M et al (2012) TPX2 and AURKA promote 20q amplicon-driven colorectal adenoma to carcinoma progression. Gut 61:1568–1575
Esquela-Kerscher A, Slack FJ (2006) Oncomirs—microRNAs with a role in cancer. Nat Rev Cancer 6:259–269
Ivanovska I, Ball AS, Diaz RL et al (2008) MicroRNAs in the miR-106b family regulate p21/CDKN1A and promote cell cycle progression. Mol Cell Biol 28:2167–2174
Petrocca F, Visone R, Onelli MR et al (2008) E2F1-regulated microRNAs impair TGFβ-dependent cell-cycle arrest and apoptosis in gastric cancer. Cancer Cell 13:272–286
Diosdado B, van de Wiel MA, Terhaar Sive Droste JS et al (2009) MiR-17-92 cluster is associated with 13q gain and c-myc expression during colorectal adenoma to adenocarcinoma progression. Br J Cancer 101:707–714
Nagel R, le Sage C, Diosdado B et al (2008) Regulation of the adenomatous polyposis coli gene by the miR-135 family in colorectal cancer. Cancer Res 68:5795–5802
Jass JR (2007) Classification of colorectal cancer based on correlation of clinical, morphological and molecular features. Histopathology 50:113–130
Guinney J, Dienstmann R, Wang X et al (2015) The consensus molecular subtypes of colorectal cancer. Nat Med 21:1350–1356
Dienstmann R, Vermeulen L, Guinney J et al (2017) Consensus molecular subtypes and the evolution of precision medicine in colorectal cancer. Nat Rev Cancer 17:79–92
Ahlquist DA (2010) Molecular detection of colorectal neoplasia. Gastroenterology 138:2127–2139
Lee JK, Liles EG, Bent S, et al. Accuracy of fecal immunochemical tests for colorectal cancer: systematic review and meta-analysis
Ahlquist DA, Harrington JJ, Burgart LJ, Roche PC (2000) Morphometric analysis of the “mucocellular layer” overlying colorectal cancer and normal mucosa: relevance to exfoliation and stool screening. Hum Pathol 31:51–57
Zou H, Taylor WR, Harrington JJ et al (2009) High detection rates of colorectal neoplasia by stool DNA testing with a novel digital melt curve assay. Gastroenterology 136:459–470
Sidransky D, Tokino T, Hamilton SR et al (1992) Identification of ras oncogene mutations in the stool of patients with curable colorectal tumors. Science 256:102–105
Deuter R, Müller O (1998) Detection of APC mutations in stool DNA of patients with colorectal cancer by HD-PCR. Hum Mutat 11:84–89
Eguchi S, Kohara N, Komuta K, Kanematsu T (1996) Mutations of the p53 gene in the stool of patients with resectable colorectal cancer. Cancer 77:1707–1710
Müller HM, Oberwalder M, Fiegl H et al (2004) Methylation changes in fecal DNA: a marker for colorectal cancer screening? Lancet 363:1283–1285
Bosch LJW, Carvalho B, Fijneman RJA et al (2011) Molecular tests for colorectal cancer screening. Clin Colorectal Cancer 10:8–23
Imperiale TF, Ransohoff DF, Itzkowitz SH et al (2014) Multitarget stool DNA testing for colorectal-cancer screening. N Engl J Med 370:1287–1297
Heigh RI, Yab TC, Taylor WR et al (2014) Detection of colorectal serrated polyps by stool DNA testing: comparison with fecal immunochemical testing for occult blood (FIT). PLoS One 9:e85659
de Wijkerslooth TR, Stoop EM, Bossuyt PM et al (2012) Immunochemical fecal occult blood testing is equally sensitive for proximal and distal advanced eoplasia. Am J Gastroenterol 107:1570–1578
Hoff G, Grotmol T, Thiis-Evensen E et al (2004) Testing for fecal calprotectin (PhiCal) in the Norwegian Colorectal Cancer Prevention trial on flexible sigmoidoscopy screening: comparison with an immunochemical test for occult blood (FlexSure OBT). Gut 53:1329–1333
Uppara M, Adaba F, Askari A et al (2015) A systematic review and meta-analysis of the diagnostic accuracy of pyruvate kinase M2 isoenzymatic assay in diagnosing colorectal cancer. World J Surg Oncol 13:48
Yajima S, Ishii M, Matsushita H et al (2007) Expression profiling of fecal colonocytes for RNA-based screening of colorectal cancer. Int J Oncol 31:1029–1037
Beaulieu J-F, Herring E, Kanaoka S, Tremblay É (2016) Use of integrin alpha 6 transcripts in a stool mRNA assay for the detection of colorectal cancers at curable stages. Oncotarget 7:14684–14692
Link A, Balaguer F, Shen Y et al (2010) Fecal microRNAs as novel biomarkers for colon cancer screening. Cancer Epidemiol Biomark Prev 19:1766–1774
Schwarzenbach H, Hoon DSB, Pantel K (2011) Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer 11:426–437
Kopreski MS, Benko FA, Kwee C et al (1997) Detection of mutant K-ras DNA in plasma or serum of patients with colorectal cancer. Br J Cancer 76:1293–1299
Church TR, Wandell M, Lofton-Day C et al (2014) Prospective evaluation of methylated SEPT9 in plasma for detection of asymptomatic colorectal cancer. Gut 63:317–325
FDA US Food & Drug Administration (2013) Letter of Approval Epi ProColon P130001 Available from: http://www.accessdata.fda.gov/scripts/cdrh/cfdocs/cfpma/pma_template.cfm?id=p130001. Accessed 31 Jan 2017
Heichman KA (2014) Blood-based testing for colorectal cancer screening. Mol Diagn Ther 18:127–135
Thomson DM, Krupey J, Freedman SO, Gold P (1969) The radioimmunoassay of circulating carcinoembryonic antigen of the human digestive system. Proc Natl Acad Sci U S A 64:161–167
Ma H, Chen G, Guo M (2016) Mass spectrometry based translational proteomics for biomarker discovery and application in colorectal cancer. PROTEOMICS - Clin Appl 10:503–515. doi:10.1002/prca.201500082
Kraus S, Shapira S, Kazanov D, et al. (2015) Predictive levels of CD24 in peripheral blood leukocytes for the early detection of colorectal adenomas and adenocarcinomas. Dis Markers 2015:916098.
García JM, García V, Peña C et al (2008) Extracellular plasma RNA from colon cancer patients is confined in a vesicle-like structure and is mRNA-enriched. RNA 14:1424–1432
Chao S, Ying J, Liew G et al (2013) Blood RNA biomarker panel detects both left- and right-sided colorectal neoplasms: a case-control study. J Exp Clin Cancer Res 32:44
Ganepola GA, Nizin J, Rutledge JR, Chang DH (2014) Use of blood-based biomarkers for early diagnosis and surveillance of colorectal cancer. World J Gastrointest Oncol 6:83–97
Young GP, Senore C, Mandel JS et al (2016) Recommendations for a step-wise comparative approach to the evaluation of new screening tests for colorectal cancer. Cancer 122:826–839
Bailey VJ, Zhang Y, Keeley BP et al (2010) Single-tube analysis of DNA methylation with silica superparamagnetic beads. Clin Chem 56:1022–1025
Huttenhain R, Soste M, Selevsek N, et al. (2012) Reproducible quantification of cancer-associated proteins in body fluids using targeted proteomics. Sci Transl Med 4:142ra94-142ra94
Schwenk JM, Igel U, Kato BS et al (2010) Comparative protein profiling of serum and plasma using an antibody suspension bead array approach. Proteomics 10:532–540
Winawer SJ, Zauber AG, Ho MN et al (1993) Prevention of colorectal cancer by colonoscopic polypectomy. The National Polyp Study Workgroup. N Engl J Med 329:1977–1981
Sillars-Hardebol AH, Carvalho B, Van Engeland M et al (2012) The adenoma hunt in colorectal cancer screening: defining the target. J Pathol 226:1–6
Matano M, Date S, Shimokawa M et al (2015) Modeling colorectal cancer using CRISPR-Cas9–mediated engineering of human intestinal organoids. Nat Med 21:256
Sahasrabuddhe VV, Luhn P, Wentzensen N (2011) Human papillomavirus and cervical cancer: biomarkers for improved prevention efforts. Future Microbiol 6:1083–1098
Van Der Veen N Uitvoeringskader Bevolkingsonderzoek Baarmoederhalskanker
Paulson TG, Maley CC, Li X et al (2009) Chromosomal instability and copy number alterations in Barrett’s esophagus and esophageal adenocarcinoma. Clin Cancer Res 15:3305–3314
Acknowledgements
The authors were supported by the Dutch Cancer Society (reference numbers: KWF Fellowship 2013-5885 and NKI 2013-6338) and a SU2C-DCS International Translational Cancer Research Dream Team Grant (MEDOCC). Stand Up To Cancer is a program of the Entertainment Industry Foundation administered by the American Association for Cancer Research.
Ethical responsibilities of authors
All authors conform to the ethical responsibilities as outlined in the ICMJE recommendation for qualification of authorship. The ICMJE recommends that authorship be based on the following four criteria:
-
Substantial contributions to the conception or design of the work or the acquisition, analysis, or interpretation of data for the work
-
Drafting the work or revising it critically for important intellectual content
-
Final approval of the version to be published
-
Agreement to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
Not applicable.
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
van Lanschot, M.C.J., Bosch, L.J.W., de Wit, M. et al. Early detection: the impact of genomics. Virchows Arch 471, 165–173 (2017). https://doi.org/10.1007/s00428-017-2159-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-017-2159-2