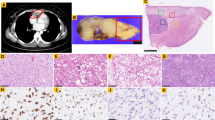
Abstract
The major cytogenetic subgroup of lipomas is characterized by aberrations of chromosome segment 12q13–15, which recombines with a large number of other chromosomal regions. The gene HMGA2 is the main target in these aberrations. For some recurrent rearrangements, chimeric transcripts, including the 5′ part of HMGA2, have been described. The 3′ partners identified are LPP, LHFP, CMKOR1, and EBF. In addition, subsets of other benign solid tumors show aberrations of 12q13–15. Among pleomorphic adenomas of the salivary glands, where the preferred recombination partner with 12q13–15 is 9p22–24, an HMGA2/NFIB fusion gene has been reported. In the present study, two cases of lipoma with rearrangements of 9p22–24 and 12q15 were analyzed by reverse transcription polymerase chain reaction to find out if HMGA2/NFIB was also present in lipoma. An in-frame fusion transcript, combining the four first exons of HMGA2 with exon 8 of NFIB, was detected in one case. It was identical to a transcript that was previously described in salivary gland adenoma and contained a stop codon shortly 3′ of the fusion point. The finding of the same fusion gene in different tumors is not unique. For example, HMGA2/LPP has been reported in lipoma, pulmonary chondroid hamartoma, and soft tissue chondroma. Since similar 9;12 translocations have been described also in rare cases of hamartoma and uterine leiomyoma, the occurrence of HMGA2/NFIB could be postulated in these tumors as well.

Similar content being viewed by others
Reference
Ashar HR, Tkachenko A, Shah P, Chada K (2003) HMGA2 is expressed in an allele-specific manner in human lipomas. Cancer Genet Cytogenet 143:160–168
Broberg K, Zhang M, Strömbeck B, Isaksson M, Nilsson M, Mertens F, Mandahl N, Panagopoulos I (2002) Fusion of RDC1 with HMGA2 in lipomas as the result of chromosome aberrations involving 2q35–37 and 12q13–15. Int J Oncol 21:321–326
Dahlén A, Mertens F, Rydholm A, Brosjö O, Wejde J, Mandahl N, Panagopoulos I (2003) Fusion, disruption and expression of HMGA2 in bone and soft tissue chondromas. Mod Pathol 16:1132–1140
Fedele M, Battista S, Manfioletti G, Croce CM, Giancotti V, Fusco A (2001) Role of the high mobility group A protein in human lipomas. Carcinogenesis 22:1583–1591
Geurts JMW, Schoenmakers EFPM, Röijer E, Aström A-K, Stenman G, van de Ven WJM (1998) Identification of NFIB as recurrent translocation partner gene of HMGIC in pleomorphic adenomas. Oncogene 16:865–872
Kataoka Yamada H, Hoshi N, Kudo M, Hareyama H, Sakuragi N, Fujimoto S (2003) Cytogenetic analysis of uterine leiomyoma: the size, histopathology and GnRHa-response in relation to chromosome karyotype. Eur J Obstet Gynecol Reprod Biol 110:58–62
Kazmierczak B, Meyer-Bolte K, Tran KH, Wockel W, Breightman I, Rosigkeit J, Bartnitzke S, Bullerdiek J (1999) A high frequency of tumors with rearrangements of genes of the HMGI(Y) family in a series of 191 pulmonary chondroid hamartomas. Genes Chromosomes Cancer 26:125–133
Kiechle-Schwarz M, Sreekantaiah C, Berger CS, Pedron S, Medchill MT, Surti U, Sandberg AA (1991) Nonrandom cytogenetic changes in leiomyomas of the female genitourinary tract. A report of 35 cases. Cancer Genet Cytogenet 53:125–136
Mandahl N, Höglund M, Mertens F, Rydholm A, Willén H, Brosjö O, Mitelman F (1994) Cytogenetic aberrations in 188 benign and borderline adipose tissue tumors. Genes Chromosomes Cancer 9:207–215
Mandahl N (2001) Methods in solid tumor cytogenetics. In: Rooney DE (ed) Human cytogenetics: malignancy and acquired abnormalities, 3rd edn. Oxford University Press, New York, pp 165–203
Mark J, Dahlenfors R, Wedell B (1997) Impact of the in vitro technique used on the cytogenetic patterns in pleomorphic adenomas. Cancer Genet Cytogenet 95:9–15
Mitelman F (1995) ISCN (1995). An international system for human cytogenetic nomenclature. Karger, Basel, Switzerland
Mitelman Database of Chromosome Aberrations in Cancer. In: Mitelman F, Johansson B, Mertens F (eds). http://cgap.nci.nih.gov/Chromosomes/Mitelman
Nielsen GP, Mandahl N (2002) Lipoma. In: Fletcher CDM, Unni KK, Mertens F (eds) World Health Organization classification of tumours. Pathology and genetics of tumours of soft tissue and bone. IARC Press, Lyon, pp 19–46
Nilsson M, Mertens F, Höglund M, Mandahl N, Panagopoulos I (2005) Truncation and fusion of HMGA2 in lipomas with rearrangements of 5q32–33 and 12q14–15. Cytogenet Genome Res (in press)
Panagopoulos I, Mertens F, Domanski HA, Isaksson M, Brosjö O, Gustafson P, Mandahl N (2001) No EWS/FLI1 fusion transcripts in giant-cell tumors of bone. Int J Cancer 93:769–772
Pandis N, Heim S, Bardi G, Flodérus UM, Willén H, Mandahl N, Mitelman F (1991) Chromosome analysis of 96 uterine leiomyomas. Cancer Genet Cytogenet 55:11–18
Petit MMR, Mols R, Schoenmakers EFPM, Mandahl N, Van de Ven WJM (1996) LPP, the preferred fusion partner gene of HMGIC in lipomas, is a novel member of the LIM protein gene family. Genomics 36:118–129
Petit MMR, Schoenmakers EFPM, Huysman C, Geurts JMW, Mandahl N, Van de Ven WJM (1999) LHFP, a novel translocation partner gene of HMGIC in a lipoma, is a member of a new family of LHFP-like genes. Genomics 57:438–441
Rogalla P, Lemke I, Kazmierczak B, Bullerdiek J (2000) An identical HMGIC–LPP fusion transcript is consistently expressed in pulmonary chondroid hamartoma with t(3;12)(q27–28;q14–15). Genes Chromosomes Cancer 29:363–366
Acknowledgements
This work was supported by the Swedish Cancer Society.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nilsson, M., Panagopoulos, I., Mertens, F. et al. Fusion of the HMGA2 and NFIB genes in lipoma. Virchows Arch 447, 855–858 (2005). https://doi.org/10.1007/s00428-005-0037-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-005-0037-9