Abstract.
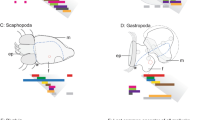
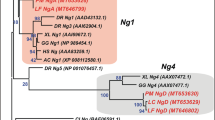
We report the identification and characterisation of five different Nkx5-related genes in medaka fish (Oryzias latipes). They constitute homologues of genes previously isolated in higher vertebrates, Nkx5–1, Nkx5–2, Hmx1/Nkx5–3 and SOHo-1, and were named accordingly: OlNkx5–1.1, OlNkx5–2, OlNkx5–3 and OlSOHo. For the Nkx5–1 gene a new, second homologue, OlNkx5–1.2, was isolated. In medaka, Nkx5 and SOHo genes are differentially expressed in three developing sensory organs: eye, ear and lateral line and later in defined brain regions. Phylogenetic analyses of the entire Nkx5 family revealed that four paralogous Nkx5 groups, Nkx5–1, Nkx5–2, Hmx1/Nkx5–3/GH6 and SOHo, are present in vertebrates. Only some of the Nkx5 family members have been identified in singular vertebrate species so far. Here we present, for the first time, the isolation of representatives of each Nkx5 subgroup in one species, the medaka fish. Based on similarities in sequence and expression patterns, and genomic organisation we propose a model of the evolutionary history of the Nkx5 family. The model predicts that the four vertebrate Nkx5 genes arose by a tandem duplication, followed by chromosomal duplication. The two Nkx5–1 genes identified so far exclusively in medaka most probably result from an additional genome duplication in the fish lineage.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Electronic Publication
Rights and permissions
About this article
Cite this article
Adamska, M., Wolff, A., Kreusler, M. et al. Five Nkx5 genes show differential expression patterns in anlagen of sensory organs in medaka: insight into the evolution of the gene family. Dev Genes Evol 211, 338–349 (2001). https://doi.org/10.1007/s004270100162
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s004270100162