Abstract
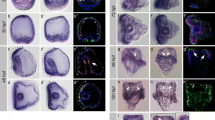
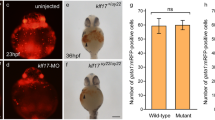
Otospiralin (OTOSP) is a small protein of unknown function, expressed in fibrocytes of the inner ear and required for normal cochlear auditory function. Despite its conservation from fish to mammals, expression of otospiralin was only investigated in mammals. Here, we report for the first time the expression profile of OTOS orthologous genes in zebrafish (Danio rerio): otospiralin and si:ch73-23l24.1 (designated otospiralin-like). In situ hybridization analyses in zebrafish embryos showed a specific expression of otospiralin-like in notochord (from 14 to 48 hpf) and similar expression patterns for otospiralin and otospiralin-like in gut (from 72 to 120 hpf), swim bladder (from 96 to 120 hpf) and inner ear (at 120 hpf). Morpholino knockdown of otospiralin and otospiralin-like showed no strong change of the body structure of the embryos at 5 dpf and the inner ear was normally formed. Nevertheless, knockdown embryos showed a reduced number of kinocilia in the lateral crista, indicating that these genes play an important role in kinocilium formation. RT-qPCR revealed that otospiralin is highly expressed in adult zebrafish inner ear comparing to the others analyzed tissues as previously shown for mice. Interestingly, otospiralin-like was not detected in the inner ear which suggests that otospiralin have a more important function in hearing than otospiralin-like. Phylogenetic analysis of otospiralin proteins in vertebrates indicated the presence of two subgroups and supported the functional divergence observed in zebrafish for otospiralin and otospiralin-like genes. This study offers the first insight into the expression of otospiralin and otospiralin-like in zebrafish. Expression data point to an important role for otospiralin in zebrafish hearing and a specific role for otospiralin-like in notochord vacuolization.






Similar content being viewed by others
Change history
27 January 2020
In the originally published article, the first names and family names of the authors were interchanged, hence not correct. The correct presentation of names is presented above.
References
Abbas L, Whitfield TT (2010) The zebrafish inner ear. Fish Physiol 29:123–171. https://doi.org/10.1016/S1546-5098(10)02904-3
Asai Y, Chan DK, Starr CJ et al (2006) Mutation of the atrophin2 gene in the zebrafish disrupts signaling by fibroblast growth factor during development of the inner ear. Proc Natl Acad Sci U S A 103:9069–9074. https://doi.org/10.1073/pnas.0603453103
Decourt B, Hillman D, Bouleau Y et al (2009) Is otospiralin inner ear specific? Evidence for its expression in mouse brain. Int J Dev Neurosci 27:87–96. https://doi.org/10.1016/j.ijdevneu.2008.09.001
Delprat B, Boulanger A, Wang J et al (2002) Downregulation of otospiralin, a novel inner ear protein, causes hair cell degeneration and deafness. J NeurosciJ Neurosci Off J Soc Neurosci 22:1718–1725
Delprat B, Ruel J, Guitton MJ et al (2005) Deafness and cochlear fibrocyte alterations in mice deficient for the inner ear protein otospiralin. Mol Cell Biol 25:847–853. https://doi.org/10.1128/MCB.25.2.847-853.2005
Edgar RC (2004) MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5:113. https://doi.org/10.1186/1471-2105-5-113
Fritzsch B, Signore M, Simeone A (2001) Otx1 null mutant mice show partial segregation of sensory epithelia comparable to lamprey ears. Dev Genes Evol 211:388–396. https://doi.org/10.1007/s004270100166
Hammond KL, Whitfield TT (2006) The developing lamprey ear closely resembles the zebrafish otic vesicle: otx1 expression can account for all major patterning differences. Dev Camb Engl 133:1347–1357. https://doi.org/10.1242/dev.02306
Kappler JA, Starr CJ, Chan DK et al (2004) A nonsense mutation in the gene encoding a zebrafish myosin VI isoform causes defects in hair-cell mechanotransduction. Proc Natl Acad Sci U S A 101:13056–13061. https://doi.org/10.1073/pnas.0405224101
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular Evolutionary Genetics Analysis vVersion 7.0 for bBigger dDatasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Lavigne-Rebillard M, Delprat B, Surget M-O et al (2003) Gene structure, chromosomal localization, and mutation screening of the human gene for the inner ear protein otospiralin. Neurogenetics 4:137–140. https://doi.org/10.1007/s10048-003-0145-0
Minowa O, Ikeda K, Sugitani Y et al (1999) Altered cochlear fibrocytes in a mouse model of DFN3 nonsyndromic deafness. Science 285:1408–1411
Morsli H, Tuorto F, Choo D et al (1999) Otx1 and Otx2 activities are required for the normal development of the mouse inner ear. Dev Camb Engl 126:2335–2343
Nielsen H (2017) Predicting sSecretory pProteins with SignalP. Methods Mol Biol 1611:59–73. https://doi.org/10.1007/978-1-4939-7015-5_6
Pace AJ, Madden VJ, Henson OW et al (2001) Ultrastructure of the inner ear of NKCC1-deficient mice. Hear Res 156:17–30
Spracklen TF, Whitehorn H, Vorster AA et al (2014) Genetic variation in Otos is associated with cisplatin-induced ototoxicity. Pharmacogenomics 15:1667–1676. https://doi.org/10.2217/pgs.14.112
Starr CJ, Kappler JA, Chan DK et al (2004) Mutation of the zebrafish choroideremia gene encoding Rab escort protein 1 devastates hair cells. Proc Natl Acad Sci U S A 101:2572–2577
Takamiya M, Weger BD, Schindler S et al (2015) Molecular description of eye defects in the zebrafish Pax6b mutant, sunrise, reveals a Pax6b-dependent genetic network in the developing anterior chamber. PLoS OnePloS One 10:e0117645. https://doi.org/10.1371/journal.pone.0117645
Thisse C, Thisse B (2008) High-resolution in situ hybridization to whole-mount zebrafish embryos. Nat Protoc 3:59–69. https://doi.org/10.1038/nprot.2007.514
Westerfield M (2000) The zebrafish book. A guide for the laboratory use of zebrafish (Danio rerio), 4th edn. Univ. of Oregon Press, Eugene
Xia A-P, Kikuchi T, Minowa O et al (2002) Late-onset hearing loss in a mouse model of DFN3 non-syndromic deafness: morphologic and immunohistochemical analyses. Hear Res 166:150–158
Zhuo X-L, Wang Y, Zhuo W-L et al (2008) Adenoviral-mediated up-regulation of Otos, a novel specific cochlear gene, decreases cisplatin-induced apoptosis of cultured spiral ligament fibrocytes via MAPK/mitochondrial pathway. Toxicology 248:33–38. https://doi.org/10.1016/j.tox.2008.03.004
Acknowledgments
We would like to thank Christelle Etard and Volker Middel for their kind advice and Nathalie Decker for technical assistance. We also thank Pr. Hela Azaiez for proofreading the English language manuscript. We acknowledge PASRI/MOBIDOC program (funded by EU) and PAQ-PAES (T1-PAS7) program for the financial support to Dr. Aissette Baanannou.
Funding
This study was co-funded by the Tunisian Ministry for Higher Education and Scientific Research and the German Federal Ministry of Education and Research (TUNGER15-046 project).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The care and use of animals were in accordance with the international guidelines on the protection of experimental animals. Because embryos used in this work were no more than 5 days old, no license was required by Council of Europe (Directive 2010/63/EU) or the local authority.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Matthias Hammerschmidt
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original version of this article was revised: In the originally published article, the first names and family names of the authors were interchanged, hence not correct. The correct presentation of names is presented above.
Rights and permissions
About this article
Cite this article
Baanannou, A., Rastegar, S., Bouzid, A. et al. Gene duplication and functional divergence of the zebrafish otospiralin genes. Dev Genes Evol 230, 27–36 (2020). https://doi.org/10.1007/s00427-019-00642-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-019-00642-8