Abstract
During Drosophila oogenesis the expression of the sulfotransferase Pipe in ventral follicle cells is crucial for dorsoventral axis formation. Pipe modifies proteins that are incorporated in the ventral eggshell and activate Toll signaling which in turn initiates embryonic dorsoventral patterning. Ventral pipe expression is the result of an oocyte-derived EGF signal which down-regulates pipe in dorsal follicle cells. The analysis of mutant follicle cell clones reveals that none of the transcription factors known to act downstream of EGF signaling in Drosophila is required or sufficient for pipe regulation. However, the pipe cis-regulatory region harbors a 31-bp element which is essential for pipe repression, and ovarian extracts contain a protein that binds this element. Thus, EGF signaling does not act by down-regulating an activator of pipe as previously suggested but rather by activating a repressor. Surprisingly, this repressor acts independent of the common co-repressors Groucho or CtBP.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dorsoventral (DV) axis formation in Drosophila is a result of the localized activation of a serine protease cascade in the perivitelline space surrounding the developing embryo (Morisato and Anderson 1995; Moussian and Roth 2005). This protease cascade leads to a ventral-to-dorsal gradient of Toll receptor activation in the embryonic plasma membrane which governs the patterning of the embryo along the DV axis. The spatially limited activation of the protease cascade at the ventral side of the egg depends on cues contained in the vitelline membrane, which is a product of somatic follicle cells which surround the growing oocyte during oogenesis. The activity of the pipe gene is required within the follicle cells to produce these ventral eggshell cues (Sen et al. 1998; Nilson and Schupbach 1998). The pipe locus is genetically complex. It codes for ten different protein isoforms (Sen et al. 1998; Sergeev et al. 2001). Seven of these are expressed in the follicular epithelium, but only one, namely Pip-PA (also called Pipe-ST2), has been shown to be essential for the polarization of the embryonic DV axis (Zhang et al. 2009b). The expression of this isoform is restricted to the ventral side of the follicular epithelium, explaining the spatial restriction of the eggshell cues.
All pipe isoforms contain a specific domain which is homologous to vertebrate glycosaminoglycan (GAG) sulfotransferases (Sen et al. 1998; Kobayashi et al. 1997; Kobayashi et al. 1999). It has been shown recently that pipe sulfates several structural components of the vitelline membrane (Zhang et al. 2009a). Being stably embedded into the vitelline membrane, these components are unlikely to diffuse, explaining the local requirement of pipe that was demonstrated by clonal analysis (Nilson and Schupbach 1998). After fertilization and egg deposition, the sulfated vitelline membrane components on the ventral side lead to localized initiation of the proteolytic cascade, and thus to the initiation of embryonic DV axis formation (Dissing et al. 2001; Moussian and Roth 2005; LeMosy 2006; Cho et al. 2010). Since pipe is the only gene involved in the induction of the embryonic DV axis which is known to be expressed asymmetrically in the follicular epithelium, it is likely to be the key component responsible for the transfer of DV polarity from the egg chamber to the embryo.
The ventral restriction of pipe expression depends on the localized activation of the EGF receptor (EGFR) in the follicular epithelium. During mid-oogenesis, the TGFα-like signaling molecule Gurken (Grk) localizes to an anterior cortical position inside the oocyte which is defined by the position of the oocyte nucleus (Neuman-Silberberg and Schupbach 1993). From here Grk is secreted and activates the EGFR in the overlying follicle cells (Queenan et al. 1999; Peri et al. 1999; Ghiglione et al. 2002; Shmueli et al. 2002). It has been shown that Grk forms a long range morphogen gradient extending from the dorsal to the ventral side of the egg chamber (Chang et al. 2008; Pai et al. 2000). Mathematical modeling predicts a direct influence of the Grk morphogen gradient on pipe expression (Goentoro et al. 2006; Yakoby et al. 2008), an idea supported by follicle cell clones mutant for the EGF pathway components Ras and Raf (James et al. 2002; Peri et al. 2002). No other pathways, such as Dpp and Notch, have been found to contribute to pipe regulation so far ((Peri et al. 2002; Shravage et al. 2007) and unpublished data). Thus, EGF pathway activation by Grk is likely the sole cause of the ventral restriction of pipe. However, the mechanisms of pipe regulation by EGF signaling are largely unknown.
In this study, we show that transcription factors which have been suggested to act downstream of EGF signaling in Drosophila and transcription factors previously assumed to play a role in the control of pipe either lack detectable effects on pipe or are insufficient to account for critical aspects of pipe's spatial control. To gain access to potential transcriptional regulators we analyzed a cis-regulatory element of the pipe upstream genomic region which drives normal pipe expression. Using bioinformatic tools based on the evolutionary conservation of functional elements (phylogenetic footprinting) and reporter constructs, we identified a 31-bp cis-regulatory element which plays a crucial role in the dorsal repression of pipe and which is bound in vitro by proteins from ovarian extracts.
Materials and methods
Fly stocks and genetic mosaic analysis in the follicular epithelium
Transgenic flies were generated as described (Spradling and Rubin 1982). The following Drosophila melanogaster strains were used: Oregon R, w 1118 , pnt D88 FRT82B/SM6a-TM6B, yan XE18 FRT40A/CyO (gifts from Matthew Freeman), and ttk 1e11 FRT 82B/TM6B (gifts from Antonio Baonza), w[1118]; P{Ubi-GFP(S65T)nls}2L P{neoFRT}40A/CyO (Bloomington stock collection), FRT82B ubi-nls GFP, y w hsflp; If/CyO; FRT82B GFP/TM6 (gifts from Stefan Luschnig), FRT82B cic fetU6 /TM3, Sb (gift from Donald Morisato), FRT 82B gro E48/TM3 and FRT 82B CtBP P1590/TM3 (gifts from Ze’ev Paroush), FRT101 GFP (gift from Thomas Klein) and pipe-LacZ (Sen et al. 1998).
All loss-of-function clones genetically marked by the absence of GFP were generated by the FRT/FLP recombination technique (Xu and Rubin 1993). Clones were induced by the follicle-cell specific recombination cassette e22c-Gal4, UAS-FLP (Duffy et al. 1998), or by a heat-shock inducible Flipase (hs-FLP 12). In the latter case, adult females were heat shocked for 1 h at 37°C, and the ovaries dissected after 4 days. pipe expression was visualized using a reporter construct expressing β-galactosidase in a pipe-like pattern (pipe-LacZ, (Sen et al. 1998) and 3 kb-pipe-LacZ generated for this study). DNA was counterstained with DAPI (Roche).
Immunohistochemistry
For antibody staining of egg chambers, ovaries were dissected in ice cold Grace's insect medium, fixed for 20 min in 4% formaldehyde, washed several times, and after blocking for 1 h with PBST (containing 0.5% BSA and 1% Triton X-100) incubated over night at 4°C with the primary antibody diluted in PBST (containing 0.5% BSA and 0.3% Triton X-100). After washing two times with PBST (0.1% Triton X-100), ovaries were again blocked for 1 h in PBST (0.1% Triton X-100) with 10% goat serum and afterwards incubated with the secondary antibody in PBST (0.1% Triton X-100) for 2 h at room temperature. After washing, the ovaries were finally mounted in Vectashield (Linaris). The following antisera were used: rabbit anti-β-galactosidase (pre-adsorbed to fixed tissue; Cappel, 1:1,000), mouse anti-β-galactosidase (Promega, 1:1,000), mouse anti-GFP (Roche, 1:500), rabbit anti-GFP (pre-adsorbed to fixed tissue; Molecular Probes, 1:200), rat anti-Cic ((Roch et al. 2002), 1:500). Primary antibodies were detected using the following fluorescence-labeled secondary antibodies: Alexa 555-conjugated goat anti-mouse IgG, Alexa 488-conjugated goat anti-guinea pig IgG (Molecular Probes, 1:400).
Cloning of promoter constructs
pipe promoter fragments were amplified by the Expand Long Template or the Expand High Fidelity PCR System (Roche) using the pipe-LacZ plasmid (Sen et al. 1998) as a template. The fragments were cloned into C4PLZ (Wharton and Crews 1993) or pHstinger (Barolo et al. 2000) using BamHI and EcoRI restriction sites. For some constructs, internal deletions or internally modified sequences were introduced using a fusion-PCR strategy (Shevchuk et al. 2004): first, two separate PCR products were generated using for each of the two products a primer with overhangs complementary to the respective other construct's primer and a BamHI or EcoRI restriction site at the other side, respectively. These two constructs were combined in a subsequent fusion PCR step. In the case of the shifted 31-bp element, 5′-phosphorylated single-stranded sense and antisense fragments for the 31-bp sequence having overhangs complementary to BamHI or EcoRI restriction sites were ordered from Sigma. After hybridization, the double-stranded fragments were ligated into BamHI or EcoRI digested 1500-deltaB-C4PLZ. Thereby, the 31-bp element was positioned either to the distal or proximal end of the 1,500-bp fragment.
Bioinformatic-TFBS prediction
To analyze the conservation of the 1.5-kb upstream cis-regulatory region of pipe, we used the GenomeVISTA browser (Bray et al. 2003; Couronne et al. 2003; Kent 2002). In addition, the PhastCons tool (Siepel et al. 2005) of the UCSC Genome Browser (Kent et al. 2002), the software ClustalW (Thompson et al. 1994), and the bayes aligner (Zhu et al. 1998) were used, which gave similar results as compared to GenomeVISTA (data not shown). For the subsequent identification of potential transcription factor binding sites (TFBS), we used several TFBS prediction programs in parallel, which restricted the output to evolutionary conserved motifs. We started the analysis using the software platform CREDO (Hindemitt and Mayer 2005), which integrates the results of a variety of algorithms that are widely used for the detection of conserved sequence motifs: AlignACE (Hughes et al. 2000), DIALIGN (Morgenstern 1999, 2004), FootPrinter (Blanchette et al. 2002); (Blanchette and Tompa 2002); (Blanchette and Tompa 2003), MEME (Bailey and Elkan 1994, 1995) and MotifSampler (Thijs et al. 2001, 2002). The CREDO-implemented programs were also tested independently and directly from the original webpage. In addition, we used the programs MOST (Motifs Searching Tool (Pizzi et al. 2005)), SOMBRERO (Self-Organizing Map for Biological Regulatory Element Recognition and Ordering (Mahony et al. 2005a, b)), and WeederH (Pavesi et al. 2007).
Electrophoretic mobility shift assay
An EMSA with the 31-bp-pipe repressor element was performed according to (Benezra et al. 1990). The single-stranded 31 bp fragments were ordered from Sigma and labeled radioactively using T4-polynucleotidkinase. The labeled complementary single-stranded probes were subsequently hybridized to form a double-stranded probe for the EMSA. Ovarian crude extract was isolated according to (Chung et al. 1996) and incubated with the labeled probe for the EMSA.
Results
Clonal analysis with mutants of candidate transcription factors
To test the influence of potential regulators of pipe expression, we generated follicle cell clones mutant for candidate transcription factors using the FLP-FRT system (Xu and Rubin 1993). We monitored pipe expression using reporter constructs which drive LacZ expression in a pattern identical to the endogenous pipe transcript (Peri et al. 2002).
The most intensively studied EGFR regulated transcription factors in Drosophila are the ETS domain proteins Yan and Pointed (Pnt) (Tootle and Rebay 2005; Rohrbaugh et al. 2002; Gabay et al. 1996; Treisman 1996). Yan and Pnt bind to the same sites, and thus compete for access to cis-regulatory regions of common target genes. Yan acts as a transcriptional repressor and Pnt as an activator (O'Neill et al. 1994; Lai and Rubin 1992). Both proteins are phosphorylated by activated EGFR, which leads to the activation of Pnt, and inactivation of Yan (Brunner et al. 1994; Klambt 1993; O'Neill et al. 1994), while in the absence of EGFR signaling, Pnt target genes are repressed by Yan (Flores et al. 2000; Halfon et al. 2000; Xu et al. 2000).
Pnt is expressed during oogenesis in dorsal-anterior follicle cells of stage 9–10 egg chambers as a consequence of the Grk dependent activation of the EGFR and is involved in the specification of the dorsal appendages (Morimoto et al. 1996; Boisclair Lachance et al. 2009; Zartman et al. 2009). To check whether Pnt and Yan are involved in the transcriptional regulation of pipe, we generated follicle cell clones mutant for pnt Δ88 (a null allele for both Pnt-isoforms) and yan XE18 (Fig. 1a–i). pnt ∆88 or yan XE18 mutant clones did not alter the expression pattern of pipe, suggesting that these two EGFR-regulated transcription factors are not involved in the regulation of pipe expression.
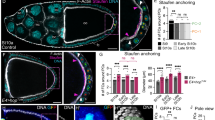
pipe repression is normal in follicle cell clones lacking the activity of transcription factors known to act downstream of EGF signaling. Stage 10 egg chambers are oriented with the anterior pole to the left. The dorsoventral orientation (lateral or ventral surface view) is indicated at the left side. Mutant cell clones are marked by the absence of GFP (green). pipe expression is monitored using a pipe-LacZ and anti-βGal antibody staining (red). a–f pnt ∆88 mutant follicle cell clones. a–c Ventral view; ventrally localized pnt ∆88 clone. d–f Lateral view; dorso-lateral pnt ∆88 clones. g–i yan XE18 clones extending from ventral to dorsal. j–o ttk 1E11 mutant follicle cell clones, nuclei are labeled with DAPI in (l). j–l Lateral view; dorsally localized ttk 1E11 clones. m–o Ventral view; ventral localized ttk 1E11 clones. l, o The nuclei are more densely packed in ttk 1E11 mutant tissue as compared to the surrounding wildtypic tissue. p–r Clones homozygous for Df(2L)γ27. The deficiency Df(2L)γ27 deletes the complete coding region and about 20 kb upstream of CF2. In all cases, the mutant follicle cell clones do not affect the proper expression pattern of pipe and thus all the tested candidate transcription factors are apparently not involved in the transcriptional regulation of pipe
A transcriptional repressor acting downstream of the EGFR pathway in several developmental contexts is the zinc finger protein Tramtrack, which exists in two isoforms known as Ttk69 and Ttk88 (Brown et al. 1991; Harrison and Travers 1990; Read and Manley 1992; Baonza et al. 2002; Lai et al. 1997; Li et al. 1997; Tang et al. 1997; Fairall et al. 1992; Lai and Li 1999; Wen et al. 2000; Xiong and Montell 1993). Ttk69 is expressed during all stages of oogenesis in all follicle cells, whereas the expression of Ttk88 can never be detected in ovaries (French et al. 2003; Read et al. 1992). Ttk69 is involved in the synthesis of the chorion and the morphogenesis of the dorsal appendages (French et al. 2003). To analyze the influence of Ttk69 on pipe expression, we generated follicle cell clones mutant for the loss-of-function allele ttk 1E11. The loss of Ttk69 on the dorsal side of the egg chamber does not lead to ectopic expression of the pipe-LacZ reporter (Fig. 1j–l). Also, clones localized ventrally do not affect the strength of pipe expression, as visualized by the LacZ reporter (Fig. 1m–o). However, the mutant cells are more closely packed than the adjacent wild-type cells, suggesting that the ttk 1E11 allele cell autonomously affects cellular density (visible in the DAPI staining in Fig. 1l as well as in the ventral pipe-LacZ expressing cells in Fig. 1n–o).
CF2, another zinc finger transcription factor, has been implicated in EGFR mediated follicle cell patterning on the basis of ectopic expression of sense and antisense constructs (Hsu et al. 1992, 1996; Shea et al. 1990). The embryonic phenotypes observed in these experiments suggest that CF2 regulates pipe. CF2 is expressed in all follicle cells starting at stage 8 and is post-translationally down-regulated from late stage 9 onwards as a direct consequence of EGFR activation in dorsal anterior follicle cells (Hsu et al. 1996, 2001; Mantrova and Hsu 1998). It is not known to date if CF2 acts as a transcriptional repressor or activator. If CF2 is indeed controlling the spatial expression pattern of pipe, it could either act directly as an essential activator of pipe or rather indirectly, for instance by the repression of a repressor.
To analyze if the loss of CF2 affects the expression of pipe, we generated CF2 mutant follicle cell clones. Since no specific CF2 allele was available, we used a deletion, in which the complete coding region and about 20 kb upstream of CF2 are missing (Df(2L)γ27, (Hsu et al. 1996)). Large Df(2L)γ27 homozygous cell clones were observed but did not modify the pipe-LacZ expression pattern (Fig. 1p–r). Thus, like all the other candidates tested so far, CF2 plays no role in the regulation of pipe expression. This negative result is surprising taking into consideration the published data regarding an involvement of CF2 in dorsoventral patterning of the embryo (Hsu et al. 1996). However, it is important to note that the prior analyses are based on heat shock induced ectopic expression of sense- and antisense constructs.
The only transcription factor which has been shown to be essential for pipe expression by standard genetic loss-of-function studies is the HMG-Box protein Capicua (Cic) (Goff et al. 2001). Cic acts as a repressor during embryogenesis, wing development, and follicle cell patterning (Jimenez et al. 2000; Roch et al. 2002; Atkey et al. 2006). Cic is downregulated by receptor tyrosine kinase signaling, resulting in the activation of target gene expression (Astigarraga et al. 2007; Cinnamon et al. 2004; Goff et al. 2001; Jimenez et al. 2000; Tseng et al. 2007). cic mutant females lay eggs with dorsalized eggshells in which dorsalized embryos develop (Atkey et al. 2006; Goff et al. 2001). It has been shown that pipe is not expressed in cic mutant egg chambers (Goff et al. 2001). To investigate whether the effect of cic on pipe expression is cell autonomous, we analyzed cic mutant follicle cell clones. We found that during stage 9, all regions of pipe expression are dependent on cic in a cell-autonomous manner (Fig. 2a–d). This also applies to the vast majority of cic mutant clones observed in stage ten egg chambers (Fig. 2e–g). In rare cases (less than 3%), non-autonomous effects were observed for anterior clones (data not shown). The predominantly cell-autonomous requirement of cic for pipe expression poses the question whether the spatial regulation of pipe can be explained via Grk-dependent down-regulation of cic. Earlier work has shown that Cic protein relocates from the nucleus to the cytoplasm in dorsal follicle cells in response to Grk signaling, so we re-investigated Cic protein distribution together with pipe expression using pipe-LacZ.
Clonal analysis and protein distribution of the HMG-Box transcription factor Capicua (Cic). Stage 9 (a–d) and 10 (e–k) egg chambers oriented with the anterior pole to the left and the dorsal side upwards. a–g cic fetU6 mutant follicle cell clones are marked by the absence of GFP (green). pipe expression is monitored using a pipe-LacZ and anti-βGal antibody staining (red). The nuclei are stained with DAPI (white). Early (a–d) and late (e–g) pipe expression depends on cic activity in a cell-autonomous fashion. h–k Cic protein distribution in stage 10 egg chambers. Cic shows uniform concentrations in lateral follicle cell nuclei where the boundary of the pipe expression domain is positioned. k Cic protein amount is reduced in a small patch of nuclei at the dorsal side of stage 10 egg chambers. The panel shows a different focal plane of the same egg chamber depicted in (h–j). The nuclei are stained with DAPI (blue)
In stage 9 egg chambers, Cic uniformly accumulates in the nuclei of all follicle cells and only during stage 10 becomes cytoplasmic in anterior-dorsal follicle cells (data not shown and Fig. 2k) (Astigarraga et al. 2007). pipe, however, is never expressed dorsally; even at the onset of its expression during early stage 9, it is only detectable on the ventral side ((Peri et al. 2002; Sen et al. 1998), data not shown). Cic protein might form a nuclear concentration gradient delimiting pipe expression via a concentration dependent mechanism. To test this hypothesis, we monitored Cic protein distribution in lateral regions of stage 10 egg chambers where pipe expression sharply drops from maximal to undetectable levels. No changes in nuclear concentrations of Cic were observed across the region of the pipe border (Fig. 2h–j). Thus, the temporal and spatial dynamics of nuclear Cic distribution do not match the pattern of pipe expression, suggesting that Cic is not a major factor responsible for the spatial regulation of pipe. Cic rather appears to be constitutively required for pipe expression by acting either as an activator or as the repressor of a repressor while the spatial control of pipe depends on other factors.
In summary, our results demonstrate that the transcription factors which so far have been discussed as mediators of EGF signaling in Drosophila are either not involved in (Yan, Pnt, Ttk, CF2) or not sufficient (Cic) to explain the spatial control of the pipe expression.
pipe promoter analysis
To identify the minimal cis-regulatory region of pipe required for proper expression in ventral follicle cells, we generated a series of LacZ reporter constructs. It has been shown previously that 8 kb of the pipe cis-regulatory region surrounding the pipe promoter drive LacZ expression in a pattern identical to the endogenous pipe transcript (Sen et al. 1998; Peri et al. 2002). This 8 kb fragment contains roughly 3 kb of the cis-regulatory region upstream of the transcription start site plus the first exon of pipe (which is common to all pipe isoforms and which contains the translation start site) and about 5 kb of the first intron.
We found that 3 kb of the sequence upstream of the transcription start site are sufficient to drive LacZ expression in a wildtype-like pattern. This 3 kb fragment was used to generate a series of proximal and distal deletions (Fig. 3). Figure 4a–e shows the expression patterns observed for promoter-distal deletions. Of particular interest are the constructs between 1,100 and 1,000 bp, which give rise to uniform expression of the reporter gene in the complete follicular epithelium (Fig. 4c, d). The ectopic expression of these constructs on the dorsal side of the egg chamber suggests that important repressor binding sites are deleted, which are responsible for EGFR-mediated inhibition of expression. The 1,100-bp construct is the largest showing clear de-repression. In contrast, the slightly longer 1,135 bp construct is ventrally expressed like the endogenous pipe transcript (Fig. 4b). Thus, by comparing these two constructs, the repressor binding can be narrowed down to 35 bp.
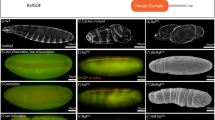
Reporter constructs identify a repressor element within the minimal cis-regulatory region driving normal pipe expression. The schematic drawing at the top represents the sequence region at the transcription start site of the pipe locus, extending 3,000 bp upstream and 1,000 bp downstream. The chromosomal orientation is inverted. The cloned reporter constructs are depicted below. The black lines mark the extent of the promoter fragments driving the expression of the LacZ reporter gene (LacZ). The position of the motifs A–C identified by TFBS prediction software (see Fig. 6) is marked with colored bars. The effects on pipe repression are indicated on the right
Expression patterns of terminally deleted pipe reporter constructs. Stage 10 egg chambers oriented with the anterior pole to the left and the dorsal side upwards. The expression pattern of the LacZ reporter gene is visualized by anti-βGal antibody staining (red, left column). The DAPI staining (white, right column) demonstrates the integrity of the follicular epithelium. a–e Reporter constructs comprising distally deleted promoter fragments. f–j Reporter constructs comprising proximally deleted promoter fragments. The expression patterns are discussed in the text. See also Fig. 3
Figure 4f–j presents the promoter constructs that are deleted from the promoter proximal side. The expression patterns of these constructs show that the region directly upstream of the transcription start site is not essential for the ventral expression in the follicular epithelium. Up to 985 bp can be deleted from the proximal site without obvious change in the expression pattern of the reporter gene. A proximal deletion of 1,000 bp leads to an attenuation of expression in most of the ventral follicle cells. Only expression at the posterior of the egg chamber remains at normal levels (Fig. 4h). This effect is even stronger when 50 additional bp are deleted, leading to nearly exclusive expression at the posterior (Fig. 4i). A truncation of 1,100 bp leads to the complete absence of expression suggesting that the pipe cis-regulatory region harbors binding sites for essential activators between 985 and 1,100 bp upstream of the start site (Fig. 4j). In all proximally deleted constructs which still show expression, the repressing influence of the EGF pathway at the dorsal side remains unaffected. Thus, it is likely that all essential repressor sites are located between 1,100 and 1,135 bp upstream of the transcription start site.
The loss of the global co-repressors Groucho or CtBP does not affect the expression of pipe
The results of the promoter analysis strongly suggest that an essential repressor binding site is located inside the pipe CRM. Most transcriptional repressors require one (or several) co-repressors to effectively inhibit the expression of their target genes (Courey and Jia 2001; Gray and Levine 1996b; Payankaulam et al. 2010). Transcriptional repressors have been subdivided into short-range repressors inhibiting expression over a distance of less than 150 bp, and long-range repressors acting from a distance of more than 500 bp (Courey and Jia 2001; Gray and Levine 1996b). These two classes of repressors mostly interact with specific long-range or short-range co-repressors.
Groucho (Gro) (Buscarlet and Stifani 2007; Chen and Courey 2000; Fisher and Caudy 1998; Mannervik et al. 1999; Parkhurst 1998) is an intensively analyzed and evolutionary highly conserved long-range co-repressor, interacting with a multitude of different repressors. Some of the well-characterized Gro-dependent repressors as well as Gro itself are regulated by the EGFR pathway (Hasson et al. 2005; Hasson and Paroush 2006; Price et al. 1997). To analyze if Gro is involved in the regulation of pipe expression, we generated follicle cell clones mutant for the loss of function allele gro E48 and analyzed the influence on the expression of the pipe-LacZ reporter gene. Neither ventrally nor dorsally located follicle cell clones mutant for gro E48 had any effect on the pipe expression pattern (Fig. 5a–c). Thus, Gro has no essential function for the transcriptional regulation of pipe.
pipe repression does not require the co-repressors Groucho or CtBP. a–i Stage 10 egg chambers oriented with the anterior pole to the left and the dorsal side upwards. Mutant follicle cell clones are marked by the absence of GFP (green). A pipe-LacZ construct reflecting the expression of pipe is visualized using an anti-βGal antibody (red). a–c Ventrally and dorsally localized gro E48 clones, d–f large CtBP P1590 clone extending from ventral to dorsal. To reveal the ventral border of the clone, the egg chamber is shown from a ventrolateral view. Therefore, the pipe domain appears to expand more to the dorsal side as compared to lateral views. g–i Dorsally localized CtBP P1590 clone
The most thoroughly studied co-repressor with regard to short-range repression is CtBP (C-terminal binding protein). CtBP is highly conserved and many repressors depend at least in part on CtBP (Chinnadurai 2002; Mannervik et al. 1999; Nibu et al. 1998; Poortinga et al. 1998). It has been shown recently that CtBP can sometimes act as a context dependent co-activator, different regions of the protein being responsible for its repressing or activating function (Fang et al. 2006). Several isoforms of CtBP are expressed in Drosophila, but to date, no unique functions have been identified for these isoforms (Fang et al. 2006; Mani-Telang and Arnosti 2007; Sutrias-Grau and Arnosti 2004). We analyzed the expression of pipe-LacZ in egg chambers bearing CtBP mutant follicle cell clones. As depicted in Fig. 5d–i, no alteration of the expression pattern could be detected, showing that pipe repression does not depend on co-repression by CtBP.
Bioinformatic identification of potential transcription factor binding sites by ab initio prediction programs
Parallel to the in vivo expression analysis of promoter reporter constructs, bioinformatic methods were used to identify potential cis-regulatory sequences. With this analysis, we firstly wanted to see whether the experimentally obtained data were in agreement with the pattern of phylogenetic conservation. Secondly, we wanted to uncover potentially redundant regulatory sites which might have been missed using deletion constructs. Thirdly, we hoped to identify potential binding sites of known TFs. We started by analyzing the conservation in the 1.5-kb upstream sequence of pipe, which drives normal expression of the reporter gene (see Fig. 4a). We compared several algorithms which all gave similar results. Figure 6a shows the pairwise alignments of the D.mel. reference genome and the homologous sequences of seven Drosophilids, which have been generated by GenomeVISTA (Bray et al. 2003; Couronne et al. 2003; Kent 2002).
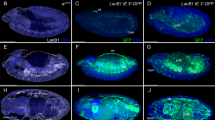
Evolutionary conservation and identification of potential TFBS in the cis-regulatory region upstream of pipe. a Plots of pair-wise alignments of the upstream region of pipe generated by GenomeVISTA. The Drosophila species used for the alignment with D. melanogaster is depicted on the left. The schematic drawing of the pipe upstream region at the top shows the coordinates of the upstream sequence, the transcription start site is marked by an arrow; the coding region of pipe is illustrated by the thick black bar. Because pipe is encoded on the reverse complement, the core promoter and the transcription start side are located at the right side and the upstream cis-regulatory sequence extends to the left. The graphs represent the conservation above 50%, the upper line marks 100% conservation. The colored shadings of the graphs depict conserved regions (70% minimal identity within at least 100 bp). Conserved translated regions are dark blue, non-coding transcribed regions are light blue, non-transcribed regions are red. The blue open box marks the ~500 bp sequence region used in the analysis with the TFBS prediction software. The position of the motifs A–C identified by this software is depicted by the vertical bars (red, blue, and green). b Graphical representation of the CREDO results. The Motif Overview at the top illustrates the position of all detected motifs. The individual hits for the different consensus motifs are depicted in one individual block for each program as colored arrows. Each individual color represents one specific consensus motif. The Summary View below shows the motifs found by all programs for each nucleotide position. The color code represent the number of different programs that detected a motif at that specific position (purple: one out of five; light blue: two; green: three; yellow: four; red: five). The number of total motif hits at each position is indicated by the height of the bar (some programs detect several overlapping motifs at the same position). The colored open boxes mark the location of the motifs A–C that are predicted by most programs, including the results of the programs MOST, SOMBRERO, and WeederH which are not implemented in CREDO. c Sequence of the motifs A, B, and C predicted by at least five different algorithms
Besides the sequence of the first exon and the cis-regulatory sequence directly upstream, only the interval located approximately 1,035–1,125 bp upstream of pipe is conserved up to D.moj. and D.vir., which diverged from D.mel. about 40 million years ago. For the D.moj. alignment, the region around 1,100 bp upstream of the transcription start site is actually the only conserved non-coding element, corroborating the analysis of the pipe-promoter constructs which shows that essential repressor binding sites are located in this section of the upstream cis-regulatory region (Figs. 3 and 4). To identify potential transcription factor binding sites (TFBS) or cis-regulatory motifs in this sequence, we used multiple bioinformatic programs. Due to the results of the promoter constructs and the determination of the position of conserved cis-regulatory sequences, we concentrated on the sequence between 1,000 and 1,500 bp upstream of the transcription start site to detect putative TFBSs (exact location of sequence: −936 to −1,453 bp, see blue box in Fig. 6a). We used eight different TFBS prediction programs in parallel, which restricted the output to potential cis-regulatory motifs conserved among different Drosophilids (phylogenetic footprinting).
The starting point for the evaluation of the results of the different prediction programs was the output of the CREDO software, which shows the correlations of five different algorithms (Fig. 6b). We concentrated on the sequence sections that show overlapping predictions for at least three of the CREDO linked programs (yellow and green color code in Fig. 6b, Summary View) and analyzed, which of the other three programs (see “Materials and methods”) identifies TFBSs at the same location. Thus, we finally discovered three putative cis-regulatory motifs that were independently predicted by at least five different programs in total. In the following, we will refer to these motifs as motif A, B, and C. The motif hits in the sequence of Drosophila melanogaster are depicted in Fig. 6c. Only motif B was identified by all eight programs. Motif A was recognized by seven and motif C by only five programs.
None of the motifs harbored obvious consensus binding sites for known transcriptions factors.
The positions of the predicted motifs A–C are depicted in the alignments generated by GenomeVISTA (Fig. 6a, red, blue, and green line). Only motif B is located in the highly conserved region. The rather low conservation of motif C demonstrates that the prediction of a motif by several programs is not simply based on the degree of conservation because in some more strongly conserved parts of the sequence, no motifs can be found that are predicted with the same confidence.
Motif B represents an essential repressor binding site
To test the relevance of the predicted cis-regulatory motifs A–C in vivo, we generated internally deleted reporter constructs. The cis-regulatory region used in these constructs comprised the complete 1.5 kb upstream sequence with just one of the predicted motifs and a few of the remaining nucleotides deleted in each of the constructs (altogether 20 bp were deleted in each of the constructs). The deletion of either motif A or motif C did not affect the expression pattern of the LacZ reporter gene (Figs. 3 and 7a, c). These results are in accordance with the expression patterns observed for terminally truncated promoter fragments that affect these motifs (Fig. 3). The constructs 1150-LacZ und 1135-LacZ, which both lack the motif C, are expressed normally in ventral follicle cells (Figs. 3 and 4b). Also, the proximally deleted construct 985-1300-LacZ in which motif A is partially deleted (only five nucleotides remaining) is normally expressed (Fig. 4g), suggesting that this motif is not representing an essential TFBS. Because motif A and C are quite similar (some of the programs even regarded them as two different versions of one consensus motif), we also generated another construct in which both motifs are deleted simultaneously. This construct was also normally expressed (data not shown). An important function for these motifs can thus be ruled out.
Expression patterns of internally deleted or mutated pipe promoter constructs. Stage 10 egg chambers oriented with the anterior pole to the left and the dorsal side upwards. The expression pattern of the LacZ reporter gene is visualized by anti-β-Gal antibody staining (red). a–c Internally deleted promoter constructs. The constructs include 1,500 bp of the pipe upstream region. In each case, one of the predicted motifs A–C (see Fig. 6c) is deleted. Only the deletion of motif B leads to a clear de-repression (uniform expression). d The sequences surrounding motif B (marked by an open blue box) which are modified (highlighted in red) in the reporter constructs mutA–mutD. The effects on pipe repression are indicated on the right. e–k Expression patterns of the reporter constructs mutA–mutD. l Expression pattern of a construct in which the 31-bp repressor element was shifted to a more distal position (see Fig. 3)
In contrast, the deletion of motif B alters the expression pattern of the reporter gene, which is evident from the uniform expression of the construct 1500-∆B-LacZ (Fig. 7b). The striking de-repression as a consequence of the deletion of motif B strongly suggests that this region harbors essential TFBSs. However, the complete internal deletion of the sequence will also affect the spacing of nearby binding sites outside the deleted section. This can compromise necessary interactions between different transcription factors and thus the observed misexpression of the 1500-∆B-LacZ construct might also result from factors that bind upstream or downstream of motif B.
To confirm the importance of the determined repressor binding site (motif B), we generated internally modified (mutated) reporter constructs (Fig. 7d–k). Seven distinct constructs were generated, in each case, 20 bp were altered, and the modified sequence was overlapping for 10 bp as compared to adjacent construct (McKnight and Kingsbury 1982). Figure 7e–k shows the results for these internally mutated reporter constructs. The observed expression patterns highlight the importance of the predicted motif B, but in addition, they show that in total, a larger sequence is essential for proper binding of repressing factors. In particular, the construct mutC that does not affect motif B at all, is also ectopically expressed dorsally. In some cases, the expression of the constructs is weaker at the very dorsal side of the egg chamber as compared to the rest of the follicle cells. This dorsal region of weaker de-repression is not located directly above the oocyte nucleus, but slightly more posterior. These results are corroborated by the deletion constructs shown in Figs. 3 and 4. The 1,100 bp construct, which contains the complete motif B, nevertheless shows clear de-repression (Fig. 4c). Thus, we can conclude that some sequences surrounding the predicted motif B also contribute to proper EGFR mediated transcriptional repression of pipe. In summary, comparing the expression patterns of all constructs, the complete region containing essential repressor binding sites could be narrowed down to 31 bp.
In order to investigate whether the function of the 31-bp repressor element is dependent on its position within the pipe CRM, we produced constructs in which the endogenous element was deleted and inserted either more proximal or more distal to the original position (Fig. 7l and data not shown). Both constructs lacked pipe repression suggesting that the 31-bp element does not represent an autonomous repressor element and that its function depends on the context of the pipe promoter.
Proteins extracted from ovaries bind in vitro specifically to the identified repressor binding element
We performed electrophoretic mobility shift assays (EMSA) to analyze if proteins from ovarian extracts bind in vitro to the 31-bp repressor element. Incubation of the radioactively labeled 31 bp fragment with crude ovarian tissue extracts led to a shift in the mobility of the fragment (Fig. 8). This bandshift was specific, since adding unlabeled fragment diminished the intensity of the shifted band (lanes 4–7), and no shift was seen with a mutated fragment (lane 1). These results suggest that ovarian extracts contain one or more proteins which specifically bind to the repressor element. The activity of transcription factors is often regulated at the level of DNA-binding capacity (Whitmarsh and Davis 2000). Since the transcriptional repressor we are interested in is directly regulated via the EGFR pathway, this factor is expected to be inactive in gurken mutant egg chambers. Thus, we analyzed whether the binding capacity of the factor is affected when proteins were extracted from gurken mutant ovaries (grk HF48 /grk 2B6 females). No change in position or quantity of the shifted fragment was observed under this condition (lane 2). However, because the EMSA is an in vitro experiment, this result does not exclude that the observed bandshift results from the binding of an EGFR regulated transcription factor. Many transcription factors are regulated at the level of nuclear translocation (Hill and Treisman 1995; Hunter and Karin 1992; Lin et al. 1998). As the EMSA is conducted with protein extract from homogenized tissue, proteins which are not localized to the nucleus in vivo are able to bind to the labeled DNA during the incubation of the probe with the protein extract.
EMSA with ovarian protein extract using a radioactively labeled 31-bp repressor element probe. Lane 1: mutated probe incubated with protein extract from wild-type ovaries; lane 2: normal probe (31-bp pipe promoter fragment) incubated with protein extract from grk HF48 /grk 2B6 ovaries; lane 3: normal probe incubated with protein extract from wild-type ovaries; lanes 4–7: normal probe incubated with protein extract from wild-type ovaries plus increasing amounts of non-labeled probe; lane 8: normal probe only. 0.1 ng of radioactively labeled probe (mutated or normal) were used. The arrow marks the bandshift that occurs after incubation with protein extracts from grk HF48 /grk 2B6 or wild-type ovaries. The asterisk marks a band which is also observed in the case of the mutated probe due to unspecific binding. The strong band at the bottom results from unbound probe. The addition of increasing amount of unlabeled probe as cold competitor reduces the strength of the band-shift (lanes 2–4). The mutated probe does not result in a bandshift when incubated (lane 1). Thus, the bandshift is caused by specific binding of one or more proteins to the 31-bp element
Discussion
The transcriptional regulation of pipe is crucial for establishing the dorsoventral axis of the embryo (Sen et al. 1998). pipe is down-regulated in dorsal follicle cells by Grk, a TGFα-like ligand which is localized close to germinal vesicle at a dorsal-anterior position of the oocyte (Cheung et al. 2011). Local Grk secretion from the oocyte leads to an anteroposterior (AP) and dorsoventral (DV) gradient of Grk uptake in the follicular epithelium (Chang et al. 2008). Quantitative analyses have suggested that the resulting two-dimensional profile of EGF signaling is sufficient to explain the spatial pattern of pipe expression, which in turn determines the DV axis of the embryo (Goentoro et al. 2006). This is a remarkable result since the DV axis is established simultaneously along the entire length of the embryo in Drosophila. Thus, all positions along the AP axis of the embryo require precise DV patterning information prior to gastrulation. This places high demands on the accuracy of the system providing DV spatial information, i.e., on the transcriptional regulation of pipe. Although we know from previous experiments that some variation in width of the pipe domain is compatible with normal DV axis formation in the embryo, even slight variations along the AP axis cause severe embryonic defects (Roth and Schupbach 1994; Roth et al. 1999).
Besides its accuracy with regard to the future embryonic axis, there is another remarkable feature of pipe regulation. The transcription of pipe shows a sharp on-off pattern in lateral regions of the follicular epithelium (Sen et al. 1998; Roth 2003). Although it has been shown that EGF signaling extends even to the ventral side of the egg chamber the distribution of active MAPK indicates that the signaling levels are very low in the lateral regions of the follicular epithelium where the border of the pipe domain resides (Pai et al. 2000; Peri et al. 1999; Wasserman and Freeman 1998). This poses the additional question of how low levels of EGF signaling resulting in a shallow gradient of MAPK activation lead to a sharp transcriptional response. To address this point, a signaling relay had been suggested (Jordan et al. 2000; Zhao et al. 2000). However, later work showed that EGF signaling controls pipe in cell-autonomous way (James et al. 2002; Peri et al. 2002). This might occur either through transcription factors which are a direct target of MAPK phosphorylation or through another tier of transcriptional regulation. According to the latter, alternative EGF signaling could either activate the transcription of a repressor or repress that of an essential activator.
To approach these questions we followed two experimental strategies. (1) We performed clonal analyses with mutants for transcription factors which have been implicated in EGF signaling in Drosophila or which have been suggested to be specifically involved in pipe regulation. (2) We analyzed cis-regulatory sequences responsible for pipe regulation in the follicular epithelium.
Candidate transcription factors involved in pipe regulation
Using clonal analysis, we first re-investigated the role of the two Ets domain proteins, Pointed and Yan, which are targets of MAPK phosphorylation in many tissues (Hsu and Schulz 2000). pnt has been shown to play a role in follicle cell patterning (Morimoto et al. 1996; Atkey et al. 2006), where it is required to establish the dorsal midline cell fate, which separates the two dorsal appendages. Eggs carrying pnt clones support embryonic development leading to larvae with no obvious DV patterning defects (Morimoto et al. 1996). This suggests that pnt has no major influence on pipe. However, previous studies did not analyze pipe expression and thus could not rule out subtle effects on pipe, e.g., on the precision of pipe repression in lateral regions. Our clonal analysis shows that despite its effect on dorsal follicle cell patterning, pnt lacks any detectable influence on pipe (Fig. 1a–f). The same applies to the Ets domain protein Yan which normally acts as a repressor in conjunction with Pnt (Fig. 1g–i).
An influence on embryonic DV patterning had been proposed for the zinc finger transcription factor CF2 (Hsu et al. 1996). Follicle cell expression of antisense or sense CF2 constructs apparently resulted in DV patterning defects in the embryo, making CF2 a likely candidate for a transcription factor controlling pipe (Hsu et al. 1996). In dorsal follicle cells, EGF signaling leads to cytoplasmic retention and degradation of CF2, while CF2 accumulates in the nuclei of lateral and ventral follicle cells (Mantrova and Hsu 1998). Accordingly, CF2 might be an activator of pipe, and the down-regulation of this factor would determine the lateral pipe border. This assumption, however, was never rigorously tested. Our clonal analysis reveals that CF2 is not involved in pipe regulation (Fig. 1p–r). The same applies for the zinc finger transcription factor Ttk, which is expressed in the follicular epithelium and has been implicated in EGF signaling in other tissues (Fig. 1j–o) (French et al. 2003) (Baonza et al. 2002).
The most likely candidate for EGF-mediated pipe regulation is the HMG-box protein Cic (Jimenez et al. 2000). cic mutant flies produce egg chambers with an anterior ring of dorsal follicle cells and lack of pipe expression (Goff et al. 2001). The expansion of dorsal follicle cells in cic mutant egg chamber is accompanied by ectopic expression of mirror in the anterior half of the follicular epithelium (Atkey et al. 2006). Clonal analysis shows that cic represses mirror in a cell-autonomous manner. Cic function is down-regulated by EGF signaling through the prevention of nuclear accumulation of Cic in dorsal follicle cells (Astigarraga et al. 2007). Thus, the dorsal follicle cell fate is established by EGF-dependent repression of repressor. Our clonal analysis shows that cic is also required for pipe expression in a cell-autonomous manner (Fig. 2a–g). Thus, one could imagine that EGF signaling-dependent down-regulation of cic in dorsal follicle cells accounts for spatial regulation of pipe. Although this cannot be strictly excluded, the temporal and spatial profile of nuclear Cic accumulation are not in agreement with this suggestion. In particular, Cic is present uniformly in the nuclei of lateral follicle cells spanning the region where the sharp on-off boundary of pipe expression resides (Fig. 2h–j).
The pipe CRM contains a repressor element
Since the clonal analysis of candidate genes presented in this paper together with previous work (Peri et al. 2002; Shravage et al. 2007) did not lead to the identification of the crucial pipe regulators, we embarked on a promoter analysis of the pipe gene. The main result of this analysis is the finding that the spatial regulation of pipe is due to transcriptional repression rather than to the down-regulation of an activator. The cis-regulatory module (CRM) driving pipe expression consists of a repressor element of about 30 bp followed by approximately 100 bp which harbor essential activator binding sites (Figs. 3, 6, and 7). Ovarian extracts contain a protein which binds to the repressor element (Fig. 8).
The constructs affecting the repressor element resulted in global de-repression along the entire AP axis of the egg chamber, suggesting that a single repressor binds to the element (Figs. 4a–e, 7a–k). However, the constructs affecting the activator binding sites showed region-specific effects. For example, partial pipe expression at the posterior of the egg chamber was observed for some constructs (proximal deletions) reducing the size of the activator domain (Fig. 4h–i). Other constructs resulted in loss of medial expression while anterior and posterior expression was maintained (data not shown). These findings indicate that the part of the pipe CRM which harbors the activator binding sites has a modular structure with separate binding sites and distinct transcription factors being responsible for the anterior, medial, and posterior subregions of pipe expression. Similar results have been described previously for the cis-regulatory region of the chorion gene s36 (Tolias and Kafatos 1990; Tolias et al. 1993).
Regarding the function of the pipe repressor element, several alternatives can be envisaged. Transcriptional repressors have been subdivided into long-range and short-range repressors (Gray and Levine 1996b). Long-range repressors function over distances of at least 500 bp by inhibiting activators bound to CRMs or by directly blocking the basal transcription machinery. The factor binding to the pipe repressor element is unlikely to work as long-range repressor since the repressor element loses its function when it is separated from the activation domain. Both a distal and proximal shift of the element by 400 and 1,000 bp, respectively, led to complete de-repression of pipe (Figs. 3, 7l). Thus, the pipe repressor appears to act in short-range manner.
Different modes of short-range repression have been described, which can be distinguished on the basis of the spatial organization of activator and repressor binding sites (Gaston and Jayaraman 2003; Kulkarni and Arnosti 2005). In one scenario, repressors and activators directly compete for overlapping binding sites (Gray and Levine 1996a, b). In this case, the deletion of the common binding sites leads to a complete loss of expression. We can exclude this mode of competitive binding, due to the uniform expression in all follicle cells which arises as a consequence of the mutation or deletion of the repressor element in the pipe CRM (Figs. 4a–e, 7).
Two other modes of short-range repression are known as quenching and direct repression. In the case of quenching, the repressors and activators bind simultaneously at independent binding sites and the repressors inhibit the interaction of the activators with the general transcription machinery (Gray et al. 1994; Gray and Levine 1996a; Arnosti et al. 1996). In contrast to this, direct or active repression involves repressors, which directly target the general transcription machinery (Gray and Levine 1996a; Arnosti et al. 1996; Latchman 1996). Both of these mechanisms could apply for the repression of pipe. However, it has been shown that in the case of direct repression, the repressor has to bind in close proximity (~100 bp) to the basal promoter (Gray and Levine 1996a; Arnosti et al. 1996; Kulkarni and Arnosti 2005). The distance of the identified pipe repressor element to the core promoter, however, exceeds 1,000 bp making direct repression an unlikely mechanism. In addition, positioning the element next to the transcription start site did not lead to repression (Fig. 3).
For short-range repressors acting during embryogenesis, quenching has been reported to be the most prevalent mode (Kulkarni and Arnosti 2003, 2005). Quenching leads to the inhibition of every activator bound in a distance of up to 100 bp surrounding the repressor binding site (Gray and Levine 1996a). In addition, the specificity of the repression depends mainly on the position of the bound repressor and not on the type of activator. This fits to the results of the pipe promoter analysis, as we detect extensive de-repression affecting all follicle cells, although the observed partial expression patterns suggest that independent activators are required for different subdomains of the follicular epithelium.
Quenching requires the presence of co-repressors which mediate the interaction between the repressors and the activators bound to independent sites. Groucho and CtBP are among the most widely studied co-repressors (Courey and Jia 2001; Payankaulam et al. 2010). While Groucho mediates both long-range and short-range repression, CtBP locally interferes with neighboring activators. Surprisingly, neither Groucho nor CtBP are involved in pipe regulation (Fig. 5). Thus, we predict that the molecular mechanisms of short-range repression functioning within the pipe CRM are different from those cases which have been studied most intensively in the early embryo (Payankaulam et al. 2010).
In summary, our analysis of pipe regulation provides a solid basis for future studies on the molecular mechanisms of EGF signaling-dependent transcriptional repression. In addition, the systematic manipulation of the pipe CRM allows the generation of transgenes which change the expression pattern of pipe independent from EGF signaling. In the past, ectopic expression of pipe has been achieved only with the help of heat-shock constructs or the GAL4-UAS system (Sen et al. 1998; Zhang et al. 2009b). These experiments had the disadvantage that they did not reproduce endogenous levels of pipe expression. For example, no experiment has been reported so far which shows the consequences of uniform expression of endogenous pipe on the embryonic DV patterning. However, such an experiment would be of pivotal importance for understanding the self-organizing processes which occur downstream of pipe and lead to the formation of the embryonic nuclear dorsal gradient that establishes the pattern of cell-fates along the embryonic DV axis (Moussian and Roth 2005).
References
Arnosti DN, Gray S, Barolo S, Zhou J, Levine M (1996) The gap protein knirps mediates both quenching and direct repression in the Drosophila embryo. Embo J 15:3659–3666
Astigarraga S, Grossman R, Diaz-Delfin J, Caelles C, Paroush Z, Jimenez G (2007) A MAPK docking site is critical for downregulation of Capicua by Torso and EGFR RTK signaling. Embo J 26:668–677
Atkey MR, Lachance JF, Walczak M, Rebello T, Nilson LA (2006) Capicua regulates follicle cell fate in the Drosophila ovary through repression of mirror. Development 133:2115–2123
Bailey TL, Elkan C (1994) Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc Int Conf Intell Syst Mol Biol 2:28–36
Bailey TL, Elkan C (1995) The value of prior knowledge in discovering motifs with MEME. Proc Int Conf Intell Syst Mol Biol 3:21–29
Baonza A, Murawsky CM, Travers AA, Freeman M (2002) Pointed and Tramtrack69 establish an EGFR-dependent transcriptional switch to regulate mitosis. Nat Cell Biol 4:976–980
Barolo S, Carver LA, Posakony JW (2000) GFP and β-galactosidase transformation vectors for promoter/enhancer analysis in Drosophila. Biotechniques 29:726–732
Benezra R, Davis RL, Lockshon D, Turner DL, Weintraub H (1990) The protein Id: a negative regulator of helix-loop-helix DNA binding proteins. Cell 61:49–59
Blanchette M, Tompa M (2002) Discovery of regulatory elements by a computational method for phylogenetic footprinting. Genome Res 12:739–748
Blanchette M, Tompa M (2003) FootPrinter: a program designed for phylogenetic footprinting. Nucleic Acids Res 31:3840–3842
Blanchette M, Schwikowski B, Tompa M (2002) Algorithms for phylogenetic footprinting. J Comput Biol 9:211–223
Boisclair Lachance JF, Fregoso Lomas M, Eleiche A, Bouchard Kerr P, Nilson LA (2009) Graded Egfr activity patterns the Drosophila eggshell independently of autocrine feedback. Development 136:2893–2902
Bray N, Dubchak I, Pachter L (2003) AVID: a global alignment program. Genome Res 13:97–102
Brown JL, Sonoda S, Ueda H, Scott MP, Wu C (1991) Repression of the Drosophila fushi tarazu (ftz) segmentation gene. Embo J 10:665–674
Brunner D, Ducker K, Oellers N, Hafen E, Scholz H, Klambt C (1994) The ETS domain protein pointed-P2 is a target of MAP kinase in the sevenless signal transduction pathway. Nature 370:386–389
Buscarlet M, Stifani S (2007) The ‘Marx’ of Groucho on development and disease. Trends Cell Biol 17:353–361
Chang WL, Liou W, Pen HC, Chou HY, Chang YW, Li WH, Chiang W, Pai LM (2008) The gradient of Gurken, a long-range morphogen, is directly regulated by Cbl-mediated endocytosis. Development 135:1923–1933
Chen G, Courey AJ (2000) Groucho/TLE family proteins and transcriptional repression. Gene 249:1–16
Cheung LS, Schupbach T, Shvartsman SY (2011) Pattern formation by receptor tyrosine kinases: analysis of the Gurken gradient in Drosophila oogenesis. Curr Opin Genet Dev (in press)
Chinnadurai G (2002) CtBP, an unconventional transcriptional corepressor in development and oncogenesis. Mol Cell 9:213–224
Cho YS, Stevens LM, Stein D (2010) Pipe-dependent ventral processing of Easter by Snake is the defining step in Drosophila embryo DV axis formation. Curr Biol 20:1133–1137
Chung YD, Kwon HC, Chung KW, Kim SJ, Kim K, Lee CC (1996) Identification of ovarian enhancer-binding factors which bind to ovarian enhancer 1 of the Drosophila genes yp1 and yp2. Mol Gen Genet 251:347–351
Cinnamon E, Gur-Wahnon D, Helman A, St Johnston D, Jimenez G, Paroush Z (2004) Capicua integrates input from two maternal systems in Drosophila terminal patterning. Embo J 23:4571–4582
Courey AJ, Jia S (2001) Transcriptional repression: the long and the short of it. Genes Dev 15:2786–2796
Couronne O, Poliakov A, Bray N, Ishkhanov T, Ryaboy D, Rubin E, Pachter L, Dubchak I (2003) Strategies and tools for whole-genome alignments. Genome Res 13:73–80
Dissing M, Giordano H, DeLotto R (2001) Autoproteolysis and feedback in a protease cascade directing Drosophila dorsal–ventral cell fate. Embo J 20:2387–2393
Duffy JB, Harrison DA, Perrimon N (1998) Identifying loci required for follicular patterning using directed mosaics. Development 125:2263–2271
Fairall L, Harrison SD, Travers AA, Rhodes D (1992) Sequence-specific DNA binding by a two zinc-finger peptide from the Drosophila melanogaster Tramtrack protein. J Mol Biol 226:349–366
Fang M, Li J, Blauwkamp T, Bhambhani C, Campbell N, Cadigan KM (2006) C-terminal-binding protein directly activates and represses Wnt transcriptional targets in Drosophila. Embo J 25:2735–2745
Fisher AL, Caudy M (1998) Groucho proteins: transcriptional corepressors for specific subsets of DNA-binding transcription factors in vertebrates and invertebrates. Genes Dev 12:1931–1940
Flores GV, Duan H, Yan H, Nagaraj R, Fu W, Zou Y, Noll M, Banerjee U (2000) Combinatorial signaling in the specification of unique cell fates. Cell 103:75–85
French RL, Cosand KA, Berg CA (2003) The Drosophila female sterile mutation twin peaks is a novel allele of tramtrack and reveals a requirement for Ttk69 in epithelial morphogenesis. Dev Biol 253:18–35
Gabay L, Scholz H, Golembo M, Klaes A, Shilo BZ, Klambt C (1996) EGF receptor signaling induces pointed P1 transcription and inactivates Yan protein in the Drosophila embryonic ventral ectoderm. Development 122:3355–3362
Gaston K, Jayaraman PS (2003) Transcriptional repression in eukaryotes: repressors and repression mechanisms. Cell Mol Life Sci 60:721–741
Ghiglione C, Bach EA, Paraiso Y, Carraway KL 3rd, Noselli S, Perrimon N (2002) Mechanism of activation of the Drosophila EGF receptor by the TGF alpha ligand gurken during oogenesis. Development 129:175–186
Goentoro LA, Reeves GT, Kowal CP, Martinelli L, Schupbach T, Shvartsman SY (2006) Quantifying the Gurken morphogen gradient in Drosophila oogenesis. Dev Cell 11:263–272
Goff DJ, Nilson LA, Morisato D (2001) Establishment of dorsal-ventral polarity of the Drosophila egg requires capicua action in ovarian follicle cells. Development 128:4553–4562
Gray S, Levine M (1996a) Short-range transcriptional repressors mediate both quenching and direct repression within complex loci in Drosophila. Genes Dev 10:700–710
Gray S, Levine M (1996b) Transcriptional repression in development. Curr Opin Cell Biol 8:358–364
Gray S, Szymanski P, Levine M (1994) Short-range repression permits multiple enhancers to function autonomously within a complex promoter. Genes Dev 8:1829–1838
Halfon MS, Carmena A, Gisselbrecht S, Sackerson CM, Jimenez F, Baylies MK, Michelson AM (2000) Ras pathway specificity is determined by the integration of multiple signal-activated and tissue-restricted transcription factors. Cell 103:63–74
Harrison SD, Travers AA (1990) The tramtrack gene encodes a Drosophila finger protein that interacts with the ftz transcriptional regulatory region and shows a novel embryonic expression pattern. Embo J 9:207–216
Hasson P, Paroush Z (2006) Crosstalk between the EGFR and other signalling pathways at the level of the global transcriptional corepressor Groucho/TLE. Br J Cancer 94:771–775
Hasson P, Egoz N, Winkler C, Volohonsky G, Jia S, Dinur T, Volk T, Courey AJ, Paroush Z (2005) EGFR signaling attenuates Groucho-dependent repression to antagonize Notch transcriptional output. Nat Genet 37:101–105
Hill CS, Treisman R (1995) Transcriptional regulation by extracellular signals: mechanisms and specificity. Cell 80:199–211
Hindemitt T, Mayer KF (2005) CREDO: a web-based tool for computational detection of conserved sequence motifs in noncoding sequences. Bioinformatics 21:4304–4306
Hsu T, Schulz RA (2000) Sequence and functional properties of Ets genes in the model organism Drosophila. Oncogene 19:6409–6416
Hsu T, Gogos JA, Kirsh SA, Kafatos FC (1992) Multiple zinc finger forms resulting from developmentally regulated alternative splicing of a transcription factor gene. Science 257:1946–1950
Hsu T, Bagni C, Sutherland JD, Kafatos FC (1996) The transcriptional factor CF2 is a mediator of EGF-R-activated dorsoventral patterning in Drosophila oogenesis. Genes Dev 10:1411–1421
Hsu T, McRackan D, Vincent TS, Gert de Couet H (2001) Drosophila Pin1 prolyl isomerase Dodo is a MAP kinase signal responder during oogenesis. Nat Cell Biol 3:538–543
Hughes JD, Estep PW, Tavazoie S, Church GM (2000) Computational identification of cis-regulatory elements associated with groups of functionally related genes in Saccharomyces cerevisiae. J Mol Biol 296:1205–1214
Hunter T, Karin M (1992) The regulation of transcription by phosphorylation. Cell 70:375–387
James KE, Dorman JB, Berg CA (2002) Mosaic analyses reveal the function of Drosophila Ras in embryonic dorsoventral patterning and dorsal follicle cell morphogenesis. Development 129:2209–2222
Jimenez G, Guichet A, Ephrussi A, Casanova J (2000) Relief of gene repression by torso RTK signaling: role of capicua in Drosophila terminal and dorsoventral patterning. Genes Dev 14:224–231
Jordan KC, Clegg NJ, Blasi JA, Morimoto AM, Sen J, Stein D, McNeill H, Deng WM, Tworoger M, Ruohola-Baker H (2000) The homeobox gene mirror links EGF signalling to embryonic dorso-ventral axis formation through notch activation. Nat Genet 24:429–433
Kent WJ (2002) BLAT—the BLAST-like alignment tool. Genome Res 12:656–664
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D (2002) The human genome browser at UCSC. Genome Res 12:996–1006
Klambt C (1993) The Drosophila gene pointed encodes two ETS-like proteins which are involved in the development of the midline glial cells. Development 117:163–176
Kobayashi M, Habuchi H, Yoneda M, Habuchi O, Kimata K (1997) Molecular cloning and expression of Chinese hamster ovary cell heparan-sulfate 2-sulfotransferase. J Biol Chem 272:13980–13985
Kobayashi M, Sugumaran G, Liu J, Shworak NW, Silbert JE, Rosenberg RD (1999) Molecular cloning and characterization of a human uronyl 2-sulfotransferase that sulfates iduronyl and glucuronyl residues in dermatan/chondroitin sulfate. J Biol Chem 274:10474–10480
Kulkarni MM, Arnosti DN (2003) Information display by transcriptional enhancers. Development 130:6569–6575
Kulkarni MM, Arnosti DN (2005) cis-regulatory logic of short-range transcriptional repression in Drosophila melanogaster. Mol Cell Biol 25:3411–3420
Lai ZC, Li Y (1999) Tramtrack69 is positively and autonomously required for Drosophila photoreceptor development. Genetics 152:299–305
Lai ZC, Rubin GM (1992) Negative control of photoreceptor development in Drosophila by the product of the yan gene, an ETS domain protein. Cell 70:609–620
Lai ZC, Fetchko M, Li Y (1997) Repression of Drosophila photoreceptor cell fate through cooperative action of two transcriptional repressors Yan and Tramtrack. Genetics 147:1131–1137
Latchman DS (1996) Inhibitory transcription factors. Int J Biochem Cell Biol 28:965–974
LeMosy EK (2006) Spatially dependent activation of the patterning protease, Easter. FEBS Lett 580:2269–2272
Li S, Li Y, Carthew RW, Lai ZC (1997) Photoreceptor cell differentiation requires regulated proteolysis of the transcriptional repressor Tramtrack. Cell 90:469–478
Lin R, Heylbroeck C, Pitha PM, Hiscott J (1998) Virus-dependent phosphorylation of the IRF-3 transcription factor regulates nuclear translocation, transactivation potential, and proteasome-mediated degradation. Mol Cell Biol 18:2986–2996
Mahony S, Golden A, Smith TJ, Benos PV (2005a) Improved detection of DNA motifs using a self-organized clustering of familial binding profiles. Bioinformatics 21(Suppl 1):i283–i291
Mahony S, Hendrix D, Golden A, Smith TJ, Rokhsar DS (2005b) Transcription factor binding site identification using the self-organizing map. Bioinformatics 21:1807–1814
Mani-Telang P, Arnosti DN (2007) Developmental expression and phylogenetic conservation of alternatively spliced forms of the C-terminal binding protein corepressor. Dev Genes Evol 217:127–135
Mannervik M, Nibu Y, Zhang H, Levine M (1999) Transcriptional coregulators in development. Science 284:606–609
Mantrova EY, Hsu T (1998) Down-regulation of transcription factor CF2 by Drosophila Ras/MAP kinase signaling in oogenesis: cytoplasmic retention and degradation. Genes Dev 12:1166–1175
McKnight SL, Kingsbury R (1982) Transcriptional control signals of a eukaryotic protein-coding gene. Science 217:316–324
Morgenstern B (1999) DIALIGN 2: improvement of the segment-to-segment approach to multiple sequence alignment. Bioinformatics 15:211–218
Morgenstern B (2004) DIALIGN: multiple DNA and protein sequence alignment at BiBiServ. Nucleic Acids Res 32:W33–W36
Morimoto AM, Jordan KC, Tietze K, Britton JS, O'Neill EM, Ruohola-Baker H (1996) Pointed, an ETS domain transcription factor, negatively regulates the EGF receptor pathway in Drosophila oogenesis. Development 122:3745–3754
Morisato D, Anderson KV (1995) Signaling pathways that establish the dorsal-ventral pattern of the Drosophila embryo. Annu Rev Genet 29:371–399
Moussian B, Roth S (2005) Dorsoventral axis formation in the Drosophila embryo—shaping and transducing a morphogen gradient. Curr Biol 15:R887–R899
Neuman-Silberberg FS, Schupbach T (1993) The Drosophila dorsoventral patterning gene gurken produces a dorsally localized RNA and encodes a TGF alpha-like protein. Cell 75:165–174
Nibu Y, Zhang H, Levine M (1998) Interaction of short-range repressors with Drosophila CtBP in the embryo. Science 280:101–104
Nilson LA, Schupbach T (1998) Localized requirements for windbeutel and pipe reveal a dorsoventral prepattern within the follicular epithelium of the Drosophila ovary. Cell 93:253–262
O'Neill EM, Rebay I, Tjian R, Rubin GM (1994) The activities of two Ets-related transcription factors required for Drosophila eye development are modulated by the Ras/MAPK pathway. Cell 78:137–147
Pai LM, Barcelo G, Schupbach T (2000) D-cbl, a negative regulator of the Egfr pathway, is required for dorsoventral patterning in Drosophila oogenesis. Cell 103:51–61
Parkhurst SM (1998) Groucho: making its Marx as a transcriptional co-repressor. Trends Genet 14:130–132
Pavesi G, Zambelli F, Pesole G (2007) WeederH: an algorithm for finding conserved regulatory motifs and regions in homologous sequences. BMC Bioinformatics 8:46
Payankaulam S, Li LM, Arnosti DN (2010) Transcriptional repression: conserved and evolved features. Curr Biol 20:R764–R771
Peri F, Bokel C, Roth S (1999) Local Gurken signaling and dynamic MAPK activation during Drosophila oogenesis. Mech Dev 81:75–88
Peri F, Technau M, Roth S (2002) Mechanisms of Gurken-dependent pipe regulation and the robustness of dorsoventral patterning in Drosophila. Development 129:2965–2975
Pizzi C, Bortoluzzi S, Bisognin A, Coppe A, Danieli GA (2005) Detecting seeded motifs in DNA sequences. Nucleic Acids Res 33:e135
Poortinga G, Watanabe M, Parkhurst SM (1998) Drosophila CtBP: a hairy-interacting protein required for embryonic segmentation and hairy-mediated transcriptional repression. Embo J 17:2067–2078
Price JV, Savenye ED, Lum D, Breitkreutz A (1997) Dominant enhancers of Egfr in Drosophila melanogaster: genetic links between the Notch and Egfr signaling pathways. Genetics 147:1139–1153
Queenan AM, Barcelo G, Van Buskirk C, Schupbach T (1999) The transmembrane region of Gurken is not required for biological activity but is necessary for transport to the oocyte membrane in Drosophila. Mech Dev 89:35–42
Read D, Manley JL (1992) Alternatively spliced transcripts of the Drosophila tramtrack gene encode zinc finger proteins with distinct DNA binding specificities. Embo J 11:1035–1044
Read D, Levine M, Manley JL (1992) Ectopic expression of the Drosophila tramtrack gene results in multiple embryonic defects, including repression of even-skipped and fushi tarazu. Mech Dev 38:183–195
Roch F, Jimenez G, Casanova J (2002) EGFR signalling inhibits Capicua-dependent repression during specification of Drosophila wing veins. Development 129:993–1002
Rohrbaugh M, Ramos E, Nguyen D, Price M, Wen Y, Lai ZC (2002) Notch activation of yan expression is antagonized by RTK/pointed signaling in the Drosophila eye. Curr Biol 12:576–581
Roth S (2003) The origin of dorsoventral polarity in Drosophila. Philos Trans R Soc Lond B Biol Sci 358:1317–1329
Roth S, Schupbach T (1994) The relationship between ovarian and embryonic dorsoventral patterning in Drosophila. Development 120:2245–2257
Roth S, Jordan P, Karess R (1999) Binuclear Drosophila oocytes: consequences and implications for dorsal-ventral patterning in oogenesis and embryogenesis. Development 126:927–934
Sen J, Goltz JS, Stevens L, Stein D (1998) Spatially restricted expression of pipe in the Drosophila egg chamber defines embryonic dorsal-ventral polarity. Cell 95:471–481
Sergeev P, Streit A, Heller A, Steinmann-Zwicky M (2001) The Drosophila dorsoventral determinant PIPE contains ten copies of a variable domain homologous to mammalian heparan sulfate 2-sulfotransferase. Dev Dyn 220:122–132
Shea MJ, King DL, Conboy MJ, Mariani BD, Kafatos FC (1990) Proteins that bind to Drosophila chorion cis-regulatory elements: a new C2H2 zinc finger protein and a C2C2 steroid receptor-like component. Genes Dev 4:1128–1140
Shevchuk NA, Bryksin AV, Nusinovich YA, Cabello FC, Sutherland M, Ladisch S (2004) Construction of long DNA molecules using long PCR-based fusion of several fragments simultaneously. Nucleic Acids Res 32:e19
Shmueli A, Cohen-Gazala O, Neuman-Silberberg FS (2002) Gurken, a TGF-alpha-like protein involved in axis determination in Drosophila, directly binds to the EGF-receptor homolog Egfr. Biochem Biophys Res Commun 291:732–737
Shravage BV, Altmann G, Technau M, Roth S (2007) The role of Dpp and its inhibitors during eggshell patterning in Drosophila. Development 134:2261–2271
Siepel A, Bejerano G, Pedersen JS, Hinrichs AS, Hou M, Rosenbloom K, Clawson H, Spieth J, Hillier LW, Richards S, Weinstock GM, Wilson RK, Gibbs RA, Kent WJ, Miller W, Haussler D (2005) Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res 15:1034–1050
Spradling AC, Rubin GM (1982) Transposition of cloned P elements into Drosophila germ line chromosomes. Science 218:341–347
Sutrias-Grau M, Arnosti DN (2004) CtBP contributes quantitatively to Knirps repression activity in an NAD binding-dependent manner. Mol Cell Biol 24:5953–5966
Tang AH, Neufeld TP, Kwan E, Rubin GM (1997) PHYL acts to down-regulate TTK88, a transcriptional repressor of neuronal cell fates, by a SINA-dependent mechanism. Cell 90:459–467
Thijs G, Lescot M, Marchal K, Rombauts S, De Moor B, Rouze P, Moreau Y (2001) A higher-order background model improves the detection of promoter regulatory elements by Gibbs sampling. Bioinformatics 17:1113–1122
Thijs G, Marchal K, Lescot M, Rombauts S, De Moor B, Rouze P, Moreau Y (2002) A Gibbs sampling method to detect overrepresented motifs in the upstream regions of coexpressed genes. J Comput Biol 9:447–464
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tolias PP, Kafatos FC (1990) Functional dissection of an early Drosophila chorion gene promoter: expression throughout the follicular epithelium is under spatially composite regulation. Embo J 9:1457–1464
Tolias PP, Konsolaki M, Halfon MS, Stroumbakis ND, Kafatos FC (1993) Elements controlling follicular expression of the s36 chorion gene during Drosophila oogenesis. Mol Cell Biol 13:5898–5906
Tootle TL, Rebay I (2005) Post-translational modifications influence transcription factor activity: a view from the ETS superfamily. Bioessays 27:285–298
Treisman R (1996) Regulation of transcription by MAP kinase cascades. Curr Opin Cell Biol 8:205–215
Tseng AS, Tapon N, Kanda H, Cigizoglu S, Edelmann L, Pellock B, White K, Hariharan IK (2007) Capicua regulates cell proliferation downstream of the receptor tyrosine kinase/ras signaling pathway. Curr Biol 17:728–733
Wasserman JD, Freeman M (1998) An autoregulatory cascade of EGF receptor signaling patterns the Drosophila egg. Cell 95:355–364
Wen Y, Nguyen D, Li Y, Lai ZC (2000) The N-terminal BTB/POZ domain and C-terminal sequences are essential for Tramtrack69 to specify cell fate in the developing Drosophila eye. Genetics 156:195–203
Wharton KA Jr, Crews ST (1993) CNS midline enhancers of the Drosophila slit and Toll genes. Mech Dev 40:141–154
Whitmarsh AJ, Davis RJ (2000) Regulation of transcription factor function by phosphorylation. Cell Mol Life Sci 57:1172–1183
Xiong WC, Montell C (1993) Tramtrack is a transcriptional repressor required for cell fate determination in the Drosophila eye. Genes Dev 7:1085–1096
Xu T, Rubin GM (1993) Analysis of genetic mosaics in developing and adult Drosophila tissues. Development 117:1223–1237
Xu C, Kauffmann RC, Zhang J, Kladny S, Carthew RW (2000) Overlapping activators and repressors delimit transcriptional response to receptor tyrosine kinase signals in the Drosophila eye. Cell 103:87–97
Yakoby N, Bristow CA, Gong D, Schafer X, Lembong J, Zartman JJ, Halfon MS, Schupbach T, Shvartsman SY (2008) A combinatorial code for pattern formation in Drosophila oogenesis. Dev Cell 15:725–737
Zartman JJ, Kanodia JS, Cheung LS, Shvartsman SY (2009) Feedback control of the EGFR signaling gradient: superposition of domain-splitting events in Drosophila oogenesis. Development 136:2903–2911
Zhang Z, Stevens LM, Stein D (2009a) Sulfation of eggshell components by Pipe defines dorsal-ventral polarity in the Drosophila embryo. Curr Biol 19:1200–1205
Zhang Z, Zhu X, Stevens LM, Stein D (2009b) Distinct functional specificities are associated with protein isoforms encoded by the Drosophila dorsal–ventral patterning gene pipe. Development 136:2779–2789
Zhao D, Woolner S, Bownes M (2000) The mirror transcription factor links signalling pathways in Drosophila oogenesis. Dev Genes Evol 210:449–457
Zhu J, Liu JS, Lawrence CE (1998) Bayesian adaptive sequence alignment algorithms. Bioinformatics 14:25–39
Acknowledgments
We are grateful to David Stein, Matthew Freeman, Antonio Baonza, Stefan Luschnig, Donald Morisato, Ze'ev Paroush, and Thomas Klein for generously providing Drosophila stocks essential for this study. We thank Gerardo Jiménez for antibodies against Capicua. We are also grateful to Jeremy Lynch for critical reading of the manuscript. The work was supported by the grants from the DFG (CRC572).
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by C. Desplan
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Technau, M., Knispel, M. & Roth, S. Molecular mechanisms of EGF signaling-dependent regulation of pipe, a gene crucial for dorsoventral axis formation in Drosophila . Dev Genes Evol 222, 1–17 (2012). https://doi.org/10.1007/s00427-011-0384-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-011-0384-2