Abstract.
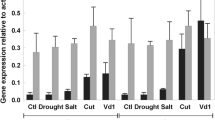
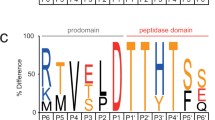
The wound-induced leucine aminopeptidase (EC 3.4.11.1) genes, LapA1 and LapA2, from tomato (Lycopersicon esculentum Mill.) were isolated and characterized. The genes were organized in a tandem array with approximately 6 kb separating their coding regions. Quantitation of LapA mRNA levels in conjunction with nuclear run-on experiments indicated that LapA genes were primarily under transcriptional control after wounding and infection with Pseudomonas syringae pv. tomato. In contrast, actin genes were down-regulated after pathogen infection. The sequences of the LapA1 and LapA2 5′-flanking regions were determined and several potential regulatory motifs were identified. Ribonuclease protection studies revealed that LapA1 and LapA2 had short 18-bp 5′-untranslated regions (UTR), both genes were expressed after wounding, and LapA1 mRNAs were 3.3-fold more abundant than LapA2 transcripts. While the region surrounding LapA1 was conserved, the 3′-UTRs and 3′-flanking regions of LapA2 had diverged in two inbred tomato lines. The accumulation of LapA mRNAs and of LAP-A (acidic pI), LAP-N (neutral pI) and LAP-related proteins were examined in two monocot and five dicot species. The LAP-N and 66- and 77-kDa LAP-related proteins were detected in healthy and wounded leaves of all plants examined. The LAP-A proteins were only detected in nightshade and their accumulation was distinct from that observed in tomato.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 8 July 1999 / Accepted: 8 August 1999
Rights and permissions
About this article
Cite this article
Chao, W., Pautot, V., Holzer, F. et al. Leucine aminopeptidases: the ubiquity of LAP-N and the specificity of LAP-A. Planta 210, 563–573 (2000). https://doi.org/10.1007/s004250050045
Issue Date:
DOI: https://doi.org/10.1007/s004250050045