Abstract
Main conclusion
Overexpression of forage sorghum oleosin genes in Arabidopsis oleosin-deficient mutant and yeast showed increased germination rate, triacylglycerol content, and protection against lipase-mediated TAG degradation.
Abstract
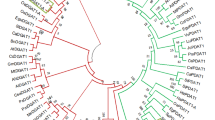
Plant lipids are an important source of ration for cattle or other livestock animals to fulfil their energy needs. Poor energy containing green forages are still one of the major sources of food for livestock animals, leaving the animals undernourished. This lowers the milk and meat production efficiency, thereby affecting human consumption. Oleosin, an essential oil body surface protein, is capable of enhancing and stabilizing the lipid content in plants. We identified and functionally characterized three forage sorghum oleosin genes (SbOle1, SbOle2, and SbOle3) in Arabidopsis and yeast. Phylogenetic analysis of SbOle proteins showed a close relationship with rice and maize oleosins. Expression analysis of SbOle genes determined a higher expression pattern in embryo followed by endosperm, while its expression in the non-seed tissues remained negligible. Overexpression of SbOle genes in Arabidopsis ole1-deficient mutants showed restoration of normal germination whereas control mutant seeds showed lower germination rates. Heterologous overexpression of SbOle in yeast cells resulted in increased TAG accumulation. Additionally, the TAG turnover assay showed the effectiveness of SbOle genes in reducing the yeast endogenous and rumen bacterial lipase-mediated TAG degradation. Taken together, our findings not only provide insights into forage sorghum oleosin for increasing the energy content in non-seed organs but also opened up the direction towards implication of oleosin in rumen protection of fodders.







Similar content being viewed by others
References
Abell BM, Holbrook LA, Abenes M et al (1997) Role of the proline knot motif in oleosin endoplasmic reticulum topology and oil body targeting. Plant Cell 9:1481–1493. https://doi.org/10.1105/tpc.9.8.1481
Bailey TL, Johnson J, Grant CE, Noble WS (2015) The MEME suite. Nucleic Acids Res 43:W39–W49. https://doi.org/10.1093/nar/gkv416
Beaudoin F, Wilkinson BM, Stirling CJ, Napier JA (2000) In vivo targeting of a sunflower oil body protein in yeast secretory (sec) mutants. Plant J 23:159–170. https://doi.org/10.1046/j.1365-313X.2000.00769.x
Beechey-Gradwell Z, Cooney L, Winichayakul S et al (2020) Storing carbon in leaf lipid sinks enhances perennial ryegrass carbon capture especially under high N and elevated CO2. J Exp Bot 71:2351–2361. https://doi.org/10.1093/jxb/erz494
Bhunia RK, Chakraborty A, Kaur R et al (2014) Seed-specific increased expression of 2S albumin promoter of sesame qualifies it as a useful genetic tool for fatty acid metabolic engineering and related transgenic intervention in sesame and other oil seed crops. Plant Mol. Biol 86:351–365. https://doi.org/10.1007/s11103-014-0233-6
Bhunia RK, Sinha K, Chawla K et al (2021a) Functional characterization of two type-1 diacylglycerol acyltransferase (DGAT1) genes from rice (Oryza sativa) embryo restoring the triacylglycerol accumulation in yeast. Plant Mol Biol 105:247–262. https://doi.org/10.1007/s11103-020-01085-w
Bhunia RK, Sinha K, Kaur R et al (2021b) A holistic view of the genetic factors involved in triggering hydrolytic and oxidative rancidity of rice bran lipids. Food Rev Int. https://doi.org/10.1080/87559129.2021.1915328
Buccioni A, Decandia M, Minieri S et al (2012) Lipid metabolism in the rumen: new insights on lipolysis and biohydrogenation with an emphasis on the role of endogenous plant factors. Anim Feed Sci Technol 174:1–25
Chapman KD, Dyer JM, Mullen RT (2012) Biogenesis and functions of lipid droplets in plants: thematic review series: lipid droplet synthesis and metabolism: from yeast to man. J Lipid Res 53:215–226
Chawla K, Kaur S, Kaur R, Bhunia RK (2020a) Metabolic engineering of oleaginous yeasts to enhance single cell oil production. J Food Process Eng. https://doi.org/10.1111/jfpe.13634
Chawla K, Sinha K, Neelam et al (2020b) Identification and functional characterization of two acyl CoA:diacylglycerol acyltransferase 1 (DGAT1) genes from forage sorghum (Sorghum bicolor) embryo. Phytochemistry 176:112405. https://doi.org/10.1016/j.phytochem.2020.112405
Chen K, Yin Y, Liu S et al (2019) Genome-wide identification and functional analysis of oleosin genes in Brassica napus L. BMC Plant Biol. https://doi.org/10.1186/s12870-019-1891-y
Durrett TP, Benning C, Ohlrogge J (2008) Plant triacylglycerols as feedstocks for the production of biofuels. Plant J 54:593–607
Gadeyne F, De Neve N, Vlaeminck B, Fievez V (2017) State of the art in rumen lipid protection technologies and emerging interfacial protein cross-linking methods. Eur J Lipid Sci Technol 119:2
Hall BG (2013) Building phylogenetic trees from molecular data with MEGA. Mol Biol Evol 30:1229–1235. https://doi.org/10.1093/molbev/mst012
Harrison SJ, Mott EK, Parsley K et al (2006) A rapid and robust method of identifying transformed Arabidopsis thaliana seedlings following floral dip transformation. Plant Methods. https://doi.org/10.1186/1746-4811-2-19
Huang AHC (2018) Plant lipid droplets and their associated proteins: potential for rapid advances. Plant Physiol 176:1894–1918
Huang CU, Huang AHC (2017) Unique motifs and length of hairpin in oleosin target the cytosolic side of endoplasmic reticulum and budding lipid droplet. Plant Physiol 174:2248–2260. https://doi.org/10.1104/pp.17.00366
Hsieh K, Huang AHC (2004) Endoplasmic Reticulum, Oleosins, and Oils in Seeds and Tapetum Cells. Plant Physiol. 136: 3427–3434. https://doi.org/10.1104/pp.104.051060
Jacquier N, Mishra S, Choudhary V, Schneiter R (2013) Expression of oleosin and perilipins in yeast promotes formation of lipid droplets from the endoplasmic reticulum. J Cell Sci 126:5198–5209. https://doi.org/10.1242/jcs.131896
Jenkins TC, Bridges WC (2007) Protection of fatty acids against ruminal biohydrogenation in cattle. Eur J Lipid Sci Technol 109:778–789
Johnson M, Zaretskaya I, Raytselis Y et al (2008) NCBI BLAST: a better web interface. Nucleic Acids Res. https://doi.org/10.1093/nar/gkn201
Klug L, Daum G (2014) Yeast lipid metabolism at a glance. FEMS Yeast Res 14:369–388
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Liu WX, Liu HL, Qu LQ (2013) Embryo-specific expression of soybean oleosin altered oil body morphogenesis and increased lipid content in transgenic rice seeds. Theor Appl Genet 126:2289–2297. https://doi.org/10.1007/s00122-013-2135-4
Olson SN, Ritter K, Rooney W et al (2012) High biomass yield energy sorghum: developing a genetic model for C4 grass bioenergy crops. Biofuels Bioprod Biorefining 6:640–655. https://doi.org/10.1002/bbb.1357
Shimada TL, Shimada T, Takahashi H et al (2008) A novel role for oleosins in freezing tolerance of oilseeds in Arabidopsis thaliana. Plant J 55:798–809. https://doi.org/10.1111/j.1365-313X.2008.03553.x
Siloto RMP, Findlay K, Lopez-Villalobos A et al (2006) The accumulation of oleosins determines the size of seed oilbodies in Arabidopsis. Plant Cell 18:1961–1974. https://doi.org/10.1105/tpc.106.041269
Sinha K, Kaur R, Bhunia RK (2019) Tailoring triacylglycerol biosynthetic pathway in plants for biofuel production. Sustain Biofuel Biomass 2:41–60
Sinha K, Kaur R, Singh N et al (2020) Mobilization of storage lipid reserve and expression analysis of lipase and lipoxygenase genes in rice (Oryza sativa var. Pusa Basmati 1) bran during germination. Phytochemistry. https://doi.org/10.1016/j.phytochem.2020.112538
Slattery RA, Ort DR (2015) Photosynthetic energy conversion efficiency: setting a baseline for gauging future improvements in important food and biofuel crops. Plant Physiol 168:383–392. https://doi.org/10.1104/pp.15.00066
Song Y, Wang XD, Rose RJ (2017) Oil body biogenesis and biotechnology in legume seeds. Plant Cell Rep 36:1519–1532
Ting JTL, Balsamo RA, Ratnayake C, Huang AHC (1997) Oleosin of plant seed oil bodies is correctly targeted to the lipid bodies in transformed yeast. J Biol Chem 272:3699–3706. https://doi.org/10.1074/jbc.272.6.3699
Tzen JTC (2012) Integral proteins in plant oil bodies. ISRN Bot 2012:1–16. https://doi.org/10.5402/2012/173954
Vanhercke T, El Tahchy A, Liu Q et al (2014) Metabolic engineering of biomass for high energy density: oilseed-like triacylglycerol yields from plant leaves. Plant Biotechnol J 12:231–239. https://doi.org/10.1111/pbi.12131
Vanhercke T, Belide S, Taylor MC et al (2019) Up-regulation of lipid biosynthesis increases the oil content in leaves of Sorghum bicolor. Plant Biotechnol J 17:220–232. https://doi.org/10.1111/pbi.12959
Winichayakul S, William Scott R, Roldan M et al (2013) In vivo packaging of triacylglycerols enhances Arabidopsis leaf biomass and energy density. Plant Physiol 162:626–639. https://doi.org/10.1104/pp.113.216820
Winichayakul S, Beechey-Gradwell Z, Muetzel S et al (2020) In vitro gas production and rumen fermentation profile of fresh and ensiled genetically modified high–metabolizable energy ryegrass. J Dairy Sci 103:2405–2418. https://doi.org/10.3168/jds.2019-16781
Xu XY, Yang HK, Singh SP et al (2018) Genetic manipulation of non-classic oilseed plants for enhancement of their potential as a biofactory for triacylglycerol production. Engineering 4:523–533. https://doi.org/10.1016/j.eng.2018.07.002
Xu XY, Akbar S, Shrestha P et al (2020) A synergistic genetic engineering strategy induced triacylglycerol accumulation in potato (Solanum tuberosum) leaf. Front Plant Sci. https://doi.org/10.3389/fpls.2020.00215
Zale J, Jung JH, Kim JY et al (2016) Metabolic engineering of sugarcane to accumulate energy-dense triacylglycerols in vegetative biomass. Plant Biotechnol J 14:661–669. https://doi.org/10.1111/pbi.12411
Zhang X, Henriques R, Lin SS et al (2006) Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat Protoc 1:641–646. https://doi.org/10.1038/nprot.2006.97
Acknowledgements
RKB acknowledge for the research funding from SERB Core Grant (CRG/2019/001154) and Department of Science and Technology in the form of INSPIRE Faculty award (DST/INSPIRE/04/2017/000484). Authors also thank Central Instrumentation Facility (CIF) for providing key instruments used during the study. Authors thank Professor Günther Daum, Graz University of Technology for kindly providing the yeast lipase-mutant strain. Authors also would like to extend their thanks to Dr Venkatesh Bhat, Indian Institute of Millets Research (IIMR), India for kindly providing the forage sorghum (CSV30F) grains.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All the authors declare that they have no financial or competing interests.
Data availability
The data used to support the findings of this study are included within the article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ojha, R., Kaur, S., Sinha, K. et al. Characterization of oleosin genes from forage sorghum in Arabidopsis and yeast reveals their role in storage lipid stability. Planta 254, 97 (2021). https://doi.org/10.1007/s00425-021-03744-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-021-03744-8