Abstract
Main conclusion
Regulation of a gene encoding coniferaldehyde 5-hydroxylase leads to substantial alterations in lignin structure in rice cell walls, identifying a promising genetic engineering target for improving grass biomass utilization.
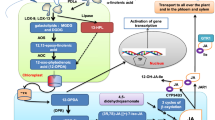
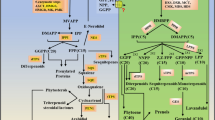
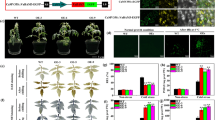
The aromatic composition of lignin greatly affects utilization characteristics of lignocellulosic biomass and, therefore, has been one of the primary targets of cell wall engineering studies. Limited information is, however, available regarding lignin modifications in monocotyledonous grasses, despite the fact that grass lignocelluloses have a great potential for feedstocks of biofuel production and various biorefinery applications. Here, we report that manipulation of a gene encoding coniferaldehyde 5-hydroxylase (CAld5H, or ferulate 5-hydroxylase, F5H) leads to substantial alterations in syringyl (S)/guaiacyl (G) lignin aromatic composition in rice (Oryza sativa), a major model grass and commercially important crop. Among three CAld5H genes identified in rice, OsCAld5H1 (CYP84A5) appeared to be predominantly expressed in lignin-producing rice vegetative tissues. Down-regulation of OsCAld5H1 produced altered lignins largely enriched in G units, whereas up-regulation of OsCAld5H1 resulted in lignins enriched in S units, as revealed by a series of wet-chemical and NMR structural analyses. Our data collectively demonstrate that OsCAld5H1 expression is a major factor controlling S/G lignin composition in rice cell walls. Given that S/G lignin composition affects various biomass properties, we contemplate that manipulation of CAld5H gene expression represents a promising strategy to upgrade grass biomass for biorefinery applications.






Similar content being viewed by others
Abbreviations
- CAld5H:
-
Coniferaldehyde 5-hydroxylase
- CWR:
-
Cell wall residue
- FA:
-
Ferulate
- G:
-
Guaiacyl
- HSQC:
-
Heteronuclear single-quantum coherence
- H:
-
p-Hydroxyphenyl
- NMR:
-
Nuclear magnetic resonance
- pCA:
-
p-Coumarate
- S:
-
Syringyl
References
Anderson NA, Tobimatsu Y, Ciesielski PN, Ximenes E, Ralph J, Donohoe BS, Ladisch M, Chapple C (2015) Manipulation of guaiacyl and syringyl monomer biosynthesis in an Arabidopsis cinnamyl alcohol dehydrogenase mutant results in atypical lignin biosynthesis and modified cell wall structure. Plant Cell 27:2195–2209
Beckham GT, Johnson CW, Karp EM, Salvachúa D, Vardon DR (2016) Opportunities and challenges in biological lignin valorization. Curr Opin Biotechnol 42:40–53
Boerjan W, Ralph J, Baucher M (2003) Lignin biosynthesis. Annu Rev Plant Biol 54:519–546
Bonawitz ND, Chapple C (2010) The genetics of lignin biosynthesis: connecting genotype to phenotype. Annu Rev Genet 44:337–363
Bonawitz ND, Chapple C (2013) Can genetic engineering of lignin deposition be accomplished without an unacceptable yield penalty? Curr Opin Biotechnol 24:336–343
Chapple CCS, Vogt T, Ellis BE, Somerville CR (1992) An Arabidopsis mutant defective in the general phenylpropanoid pathway. Plant Cell 4:1413–1424
Chen CL (1992) Nitrobenzene and cupric oxide oxidations. In: Lin SY, Dence CW (eds) Methods in lignin chemistry. Springer, Berlin, pp 301–321
Ciesielski PN, Resch MG, Hewetson B, Killgore JP, Curtin A, Anderson N, Chiaramonti AN, Hurley DC, Sanders A, Himmel ME et al (2014) Engineering plant cell walls: tuning lignin monomer composition for deconstructable biofuel feedstocks or resilient biomaterials. Green Chem 16:2627–2635
del Río JC, Rencoret J, Prinsen P, Martínez ÁT, Ralph J, Gutiérrez A (2012) Structural characterization of wheat straw lignin as revealed by analytical pyrolysis, 2D-NMR, and reductive cleavage methods. J Agric Food Chem 60:5922–5935
Ding J, Jia J, Yang L, Wen H, Zhang C, Liu W, Zhang D (2004) Validation of a rice specific gene, sucrose phosphate synthase, used as the endogenous reference gene for qualitative and real-time quantitative PCR detection of transgenes. J Agric Food Chem 52:3372–3377
Franke R, McMichael CM, Meyer K, Shirley AM, Cusumano JC, Chapple C (2000) Modified lignin in tabacco and popular plants over-expressing the Arabidopsis gene encoding ferulate 5-hydroxylase. Plant J 22:223–234
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N et al (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40:1178–1186
Grabber JH, Quideau S, Ralph J (1996) p-Coumaroylated syringyl units in maize lignin: implications for β-ether cleavage by thioacidolysis. Phytochemistry 43:1189–1194
Grand C (1984) Ferulic acid 5-hydroxylase: a new cytochrome P-450-dependent enzyme from higher plant microsomes involved in lignin synthesis. FEBS Lett 169:7–11
Gui J, Shen J, Li L (2011) Functional characterization of evolutionarily divergent 4-coumarate: coenzyme A ligases in rice. Plant Physiol 157:574–586
Guillaumie S, San-Clemente H, Deswarte C, Martinez Y, Lapierre C, Murigneux A, Barriere Y, Pichon M, Goffner D (2006) MAIZEWALL. Database and developmental gene expression profiling of cell wall biosynthesis and assembly in maize. Plant Physiol 143:339–363
Hattori T, Murakami S, Mukai M, Yamada T, Hirochika H, Ike M, Tokuyasu K, Suzuki S, Sakamoto M, Umezawa T (2012) Rapid analysis of transgenic rice straw using near-infrared spectroscopy. Plant Biotechnol 29:359–366
Hiei Y, Ohta S, Komari T, Kumashiro T (1994) Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J 6:271–282
Ho SN, Hunt HD, Horton RM, Pullen JK, Pease LR (1989) Site-directed mutagenesis by overlap extension using the polymerase chain reaction. Gene 77:51–59
Humphreys JM, Hemm MR, Chapple C (1999) New routes for lignin biosynthesis defined by biochemical characterization of recombinant ferulate 5-hydroxylase, a multifunctional cytochrome P450-dependent monooxygenase. Proc Natl Acad Sci USA 96:10045–10050
Huntley SK, Ellis D, Gilbert M, Chapple C, Mansfield SD (2003) Significant increases in pulping efficiency in C4H-F5H-transformed poplars: improved chemical savings and reduced environmental toxins. J Agric Food Chem 51:6178–6183
Kim H, Ralph J (2010) Solution-state 2D NMR of ball-milled plant cell wall gels in DMSO-d 6 /pyridine-d 5 . Org Biomol Chem 8:576–591
Koshiba T, Hirose N, Mukai M, Yamamura M, Hattori T, Suzuki S, Sakamoto M, Umezawa T (2013a) Characterization of 5-hydroxyconiferaldehyde O-methyltransferase in Oryza sativa. Plant Biotechnol 30:157–167
Koshiba T, Murakami S, Hattori T, Mukai M, Takahashi A, Miyao A, Hirochika H, Suzuki S, Sakamoto M, Umezawa T (2013b) CAD2 deficiency causes both brown midrib and gold hull and internode phenotypes in Oryza sativa L. cv. Nipponbare. Plant Biotechnol 30:365–373
Koshiba T, Yamamoto N, Tobimatsu Y, Yamamura M, Suzuki S, Hattori T, Mukai M, Noda S, Shibata D, Sakamoto M, Umezawa T (2017) MYB-mediated upregulation of lignin biosynthesis in Oryza sativa towards biomass refinery. Plant Biotechnol 34:7–15
Kuroda M, Kimizy M, Mikami C (2010) A simple set of plasmids for the production of transgenic plants. Biosci Biotechnol Biochem 74:2348–2351
Lam PY, Tobimatsu Y, Takeda Y, Suzuki S, Yamamura M, Umezawa T, Lo C (2017) Disrupting flavone synthase II alters lignin and improves biomass digestibility. Plant Physiol. doi:10.1104/pp.16.01973
Lan W, Lu F, Regner M, Zhu Y, Rencoret J, Ralph SA, Zakai UI, Morreel K, Boerjan W, Ralph J (2015) Tricin, a flavonoid monomer in monocot lignification. Plant Physiol 167:1284–1295
Lan W, Morreel K, Lu F, Rencoret J, del Río JC, Voorend W, Vermerris W, Boerjan W, Ralph J (2016) Maize tricin-oligolignol metabolites and their implications for monocot lignification. Plant Physiol 171:810–820
Lapierre C, Monties B, Roland C (1986) Preparative thioacidolysis of spruce lignin: isolation and identification of main monomeric products. Holzforschung 40:47–50
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R et al (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Li L, Popko JL, Umezawa T, Chiang VL (2000) 5-Hydroxyconiferyl aldehyde modulates enzymatic methylation for syringyl monolignol formation, a new view of monolignol biosynthesis in angiosperms. J Biol Chem 275:6537–6545
Li L, Zhou Y, Cheng X, Sun J, Marita JM, Ralph J, Chiang VL (2003) Combinatorial modification of multiple lignin traits in trees through multigene cotransformation. Proc Natl Acad Sci USA 100:4939–4944
Li X, Ximenes E, Kim Y, Slininger M, Meilan R, Ladisch M, Chapple C (2010) Lignin monomer composition affects Arabidopsis cell-wall degradability after liquid hot water pretreatment. Biotechnol Biofuels 3:27–34
Mansfield SD, Kim H, Lu F, Ralph J (2012) Whole plant cell wall characterization using solution-state 2D NMR. Nat Protoc 7:1579–1589
Marita JM, Hatfield RD, Rancour DM, Frost KE (2014) Identification and suppression of the p-coumaroyl CoA: hydroxycinnamyl alcohol transferase in Zea mays L. Plant J 78:850–864
Meyer K, Shirley AM, Cusumano JC, Bell-Lelong DA, Chapple C (1998) Lignin monomer composition is determined by the expression of a cytochrome P450-dependent monooxygenase in Arabidopsis. Proc Natl Acad Sci USA 95:6619–6623
Miki D, Shimamoto K (2004) Simple RNAi vectors for stable and transient suppression of gene function in rice. Plant Cell Physiol 45:490–495
Miki D, Itoh R, Shimamoto K (2005) RNA silencing of single and multiple members in a gene family of rice. Plant Physiol 138:1903–1913
Moldenhauer KAK, Gibbson JH (2003) Rice morphology and development. In: Smith CW, Dilday RH (eds) Rice: origin, history, technology, and production. Wiley, Hoboken, pp 103–127
Nelson DR (2009) The cytochrome p450 homepage. Hum Genom 4:59–65
Noda S, Koshiba T, Hattori T, Yamaguchi M, Suzuki S, Umezawa T (2015) The expression of a rice secondary wall-specific cellulose synthase gene, OsCesA7, is directly regulated by a rice transcription factor, OsMYB58/63. Planta 242:589–600
Osakabe K, Tsao CC, Li L, Popko JL, Umezawa T, Carraway DT, Smeltzer RH, Joshi CP, Chiang VL (1999) Coniferyl aldehyde 5-hydroxylation and methylation direct syringyl lignin biosynthesis in angiosperms. Proc Natl Acad Sci USA 96:8955–8960
Perrière G, Gouy M (1996) WWW-query: an on-line retrieval system for biological sequence banks. Biochimie 78:364–369
Petrik DL, Karlen SD, Cass CL, Padmakshan D, Lu F, Liu S, Le Bris P, Antelme S, Santoro N, Wilkerson CG et al (2014) p-Coumaroyl-CoA:monolignol transferase (PMT) acts specifically in the lignin biosynthetic pathway in Brachypodium distachyon. Plant J 77:713–726
Ragauskas AJ, Beckham GT, Biddy MJ, Chandra R, Chen F, Davis MF, Davison BH, Dixon RA, Gilna P, Keller M et al (2014) Lignin valorization: improving lignin processing in the biorefinery. Science 344:1246843–1246843
Ralph J, Lundquist K, Brunow G, Lu F, Kim H, Schatz PF, Marita JM, Hatfield RD, Ralph SA, Christensen JH et al (2004) Lignins: natural polymers from oxidative coupling of 4-hydroxyphenyl-propanoids. Phytochem Rev 3:29–60
Reddy MS, Chen F, Shadle G, Jackson L, Aljoe H, Dixon RA (2005) Targeted down-regulation of cytochrome P450 enzymes for forage quality improvement in alfalfa (Medicago sativa L.). Proc Natl Acad Sci USA 102:16573–16578
Rice P, Longden I, Bleasby A (2000) EMBOSS: the European molecular biology open software suite. Trends Genet 16:276–277
Rinaldi R, Jastrzebski R, Clough T, Ralph J, Kennema M, Bruijnincx PCA, Weckhuysen BM (2016) Paving the way for lignin valorisation: recent advances in bioengineering, biorefining and catalysis. Angew Chem Int Ed 55:8164–8215
Saidi M, Samimi F, Karimipourfard D, Nimmanwudipong T, Gates BC, Rahimpour MR (2014) Upgrading of lignin-derived bio-oils by catalytic hydrodeoxygenation. Energy Environ Sci 7:103–129
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sato Y, Takehisa H, Kamatsuki K, Minami H, Namiki N, Ikawa H, Ohyanagi H, Sugimoto K, Antonio BA, Nagamura Y (2013) RiceXPro version 3.0: expanding the informatics resource for rice transcriptome. Nucleic Acids Res 41:1206–1213
Shen H, Mazarei M, Hisano H, Escamilla-Trevino L, Fu C, Pu Y, Rudis MR, Tang Y, Xiao X, Jackson L et al (2013) A genomics approach to deciphering lignin biosynthesis in switchgrass. Plant Cell 25:4342–4361
Shi J, Pattathil S, Ramakrishnan P, Anderson NA, Kim JI, Venketachalam S, Hahn MG, Chapple C, Simmons BA, Singh S (2016) Impact of engineered lignin composition on biomass recalcitrance and ionic liquid pretreatment efficiency. Green Chem 18:4884–4895
Shigeto J, Ueda Y, Sasaki S, Fujita K, Tsutsumi Y (2017) Enzymatic activities for lignin monomer intermediates highlight the biosynthetic pathway of syringyl monomers in Robinia pseudoacacia. J Plant Res 130:203–210
Sibout R, Baucher M, Gatineau M, Doorsselaere JV, Mila I, Pollet B, Maba B, Pilate G, Lapierre C, Boerjan W, Jouanin L (2002) Expression of a popular cDNA encoding a ferulate-5-hydroxylase/coniferaldehyde 5-hydroxylase increases S lignin deposition on Arabidopsis thaliana. Plant Physiol Biochem 40:1087–1096
Silva TCF, Santos RB, Jameel H, Colodette JL, Lucia LA (2012) Quantitative molecular structure-pyrolytic energy correlation for hardwood lignins. Energy Fuels 26:1315–1322
Stewart JJ, Akiyama T, Chapple C, Ralph J, Mansfield SD (2009) The effects on lignin structure of overexpression of ferulate 5-hydroxylase in hybrid poplar. Plant Physiol 150:621–635
Studer MH, DeMartini JD, Davis MF, Sykes RW, Davison B, Keller M, Tuskan GA, Wyman CE (2011) Lignin content in natural Populus variants affects sugar release. Proc Natl Acad Sci USA 108:6300–6305
Sun JX, Sun XF, Sun RC, Fowler P, Baird MS (2003) Inhomogeneities in the chemical structure of sugarcane bagasse lignin. J Agric Food Chem 51:6719–6725
Sun SL, Wen JL, Ma MG, Li MF, Sun RC (2013) Revealing the structural inhomogeneity of lignins from sweet sorghum stem by successive alkali extractions. J Agric Food Chem 61:4226–4235
Suzuki S, Suzuki Y, Yamamoto N, Hattori T, Sakamoto M, Umezawa T (2009) High-throughput determination of thioglycolic acid lignin from rice. Plant Biotechnol 26:337–340
Tobimatsu Y, Chen F, Nakashima J, Escamilla-Trevino LL, Jackson L, Dixon RA, Ralph J (2013) Coexistence but independent biosynthesis of catechyl and guaiacyl/syringyl lignin polymers in seed coats. Plant Cell 25:2587–2600
Umezawa T (2010) The cinnamate/monolignol pathway. Phytochem Rev 9:1–17
Vanholme R, Demedts B, Morreel K, Ralph J, Boerjan W (2010) Lignin biosynthesis and structure. Plant Physiol 153:895–905
Wagner A, Tobimatsu Y, Phillips L, Flint H, Geddes B, Lu F, Ralph J (2015) Syringyl lignin production in conifers: proof of concept in a pine tracheary element system. Proc Natl Acad Sci USA 112:6218–6223
Wang JP, Shuford CM, Li Q, Song J, Lin YC, Sun YH, Chen HC, Williams CM, Muddiman DC, Sederoff RR et al (2012) Functional redundancy of the two 5-hydroxylases in monolignol biosynthesis of Populus trichocarpa: LC–MS/MS based protein quantification and metabolic flux analysis. Planta 236:795–808
Wang P, Dudareva N, Morgan JA, Chapple C (2015) Genetic manipulation of lignocellulosic biomass for bioenergy. Curr Opin Chem Biol 29:32–39
Wen JL, Sun SL, Xue BL, Sun RC (2015) Structural elucidation of inhomogeneous lignins from bamboo. Int J Biol Macromol 77:250–259
Weng JK, Mo H, Chapple C (2010) Over-expression of F5H in COMT-deficient Arabidopsis leads to enrichment of an unusual lignin and disruption of pollen wall formation: lignin modification leads to male sterility. Plant J 64:898–911
Weng JK, Li Y, Mo H, Chapple C (2012) Assembly of an evolutionarily new pathway for α-pyrone biosynthesis in Arabidopsis. Science 337:960–964
White RH (1986) Effect of lignin content and extractives on the higher heating value of wood. Wood Fiber Sci 19:446–452
Withers S, Lu F, Kim H, Zhu Y, Ralph J, Wilkerson CG (2012) Identification of grass-specific enzyme that acylates monolignols with p-coumarate. J Biol Chem 287:8347–8355
Wu M, Pang J, Lu F, Zhang X, Che L, Xu F, Sun R (2013) Application of new expansion pretreatment method on agricultural waste. Part I: Influence of pretreatment on the properties of lignin. Ind Crops Prod 50:887–895
Yamamura M, Hattori T, Suzuki S, Shibata D, Umezawa T (2010) Microscale alkaline nitrobenzene oxidation method for high-throughput determination of lignin aromatic components. Plant Biotechnol 27:305–310
Yamamura M, Wada S, Sakakibara N, Nakatsubo T, Suzuki S, Hattori T, Takeda M, Sakurai N, Suzuki H, Shibata D et al (2011) Occurrence of guaiacyl/p-hydroxyphenyl lignin in Arabidopsis thaliana T87 cells. Plant Biotechnol 28:1–8
Yamamura M, Hattori T, Suzuki S, Shibata D, Umezawa T (2012) Microscale thioacidolysis method for the rapid analysis of β-O-4 substructures in lignin. Plant Biotechnol 29:419–423
Yamamura M, Noda S, Hattori T, Shino A, Kikuchi J, Takabe K, Tagane S, Gau M, Uwatoko N, Mii M et al (2013) Characterization of lignocellulose of Erianthus arundinaceus in relation to enzymatic saccharification efficiency. Plant Biotechnol 30:25–35
Yang L, Ding J, Zhang C, Jia J, Weng H, Liu W, Zhang D (2005) Estimating the copy number of transgenes in transformed rice by real-time quantitative PCR. Plant Cell Rep 23:759–763
Yue F, Lu F, Sun RC, Ralph J (2012) Syntheses of lignin-derived thioacidolysis monomers and their uses as quantitation standards. J Agric Food Chem 60:922–928
Zhang K, Qian Q, Huang Z, Wang Y, Li M, Hong L, Zeng D, Gu M, Chu C, Cheng Z (2006) GOLD HULL AND INTERNODE2 encodes a primarily multifunctional cinnamyl-alcohol dehydrogenase in rice. Plant Physiol 140:972–983
Zhao Q, Tobimatsu Y, Zhou R, Pattathil S, Gallego-Giraldo L, Fu C, Jackson LA, Hahn MG, Kim H, Chen F et al (2013) Loss of function of cinnamyl alcohol dehydrogenase 1 leads to unconventional lignin and a temperature-sensitive growth defect in Medicago truncatula. Proc Natl Acad Sci USA 110:13660–13665
Acknowledgements
We thank Ms. Aiko Morita, Ms. Mai Mukai, Ms. Sawako Otsu, Dr. Daisuke Kabusaki, Ms. Masami Tanigawa, Ms. Megumi Urano, and Ms. Kumiko Murata (Research Institute for Sustainable Humanosphere, Kyoto University) for assisting in the development and characterization of rice transgenic lines, Dr. Hironori Kaji and Ms. Ayaka Maeno (Institute for Chemical Research, Kyoto University) for their assistance in NMR analysis, and Dr. Junji Sugiyama and Dr. Tomoya Imai (Research Institute for Sustainable Humanosphere, Kyoto University) for their assistance in microscopy. We also thank Dr. Ko Shimamoto (Nara Institute of Science and Technology) and Dr. Masaharu Kuroda (National Agricultural Research Center) for providing pANDA and pZH2B-mUP vectors. This work was supported in part by grants from the Japan Science and Technology Agency/Japan International Cooperation Agency (Science and Technology Research Partnership for Sustainable Development, SATREPS), the Ministry of Agriculture, Forestry and Fisheries of Japan (Genomics for Agricultural Innovation, GMA-0006), the Japan Society for the Promotion of Science (Grants-in-aid for Scientific Research, KAKENHI, #25292104 and #25450241), and the New Energy and Industrial Technology Development Organization (NEDO), Japan. A part of this study was conducted using the facilities in the Development and Assessment of Sustainable Humanosphere/Forest Biomass Analytical System (DASH/FBAS) at Research Institute for Sustainable Humanosphere, and the NMR spectrometer in the Joint Usage/Research Center (JURC) at Institute for Chemical Research, Kyoto University, Japan.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Takeda, Y., Koshiba, T., Tobimatsu, Y. et al. Regulation of CONIFERALDEHYDE 5-HYDROXYLASE expression to modulate cell wall lignin structure in rice. Planta 246, 337–349 (2017). https://doi.org/10.1007/s00425-017-2692-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-017-2692-x