Abstract
Purpose
Resistance training (RT) induces muscle growth at varying rates across RT phases, and evidence suggests that the muscle-molecular responses to training bouts become refined or attenuated in the trained state. This study examined how proteolysis-related biomarkers and extracellular matrix (ECM) remodeling factors respond to a bout of RT in the untrained (UT) and trained (T) state.
Methods
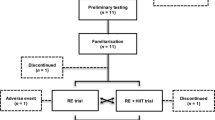
Participants (19 women and 19 men) underwent 10 weeks of RT. Biopsies of vastus lateralis were collected before and after (24 h) the first (UT) and last (T) sessions. Vastus lateralis cross-sectional area (CSA) was assessed before and after the experimental period.
Results
There were increases in muscle and type II fiber CSAs. In both the UT and T states, calpain activity was upregulated and calpain-1/-2 protein expression was downregulated from Pre to 24 h. Calpain-2 was higher in the T state. Proteasome activity and 20S proteasome protein expression were upregulated from Pre to 24 h in both the UT and T. However, proteasome activity levels were lower in the T state. The expression of poly-ubiquitinated proteins was unchanged. MMP activity was downregulated, and MMP-9 protein expression was elevated from Pre to 24 h in UT and T. Although MMP-14 protein expression was acutely unchanged, this marker was lower in T state. TIMP-1 protein levels were reduced Pre to 24 h in UT and T, while TIMP-2 protein levels were unchanged.
Conclusion
Our results are the first to show that RT does not attenuate the acute-induced response of proteolysis and ECM remodeling-related biomarkers.





Similar content being viewed by others
Data availability
Source data are available from the corresponding authors upon reasonable request.
Abbreviations
- ANCOVA:
-
Analysis of covariance
- AU:
-
Arbitray units
- BCA:
-
Bicinchoninic acid
- BSA:
-
Bovine serum albumin
- CI:
-
Confidence interval
- CSA:
-
Cross-sectional area
- ECM:
-
Extracellular matrix
- ES:
-
Effect size
- fCSA:
-
Fiber cross-sectional area
- FOXO1:
-
Forkhead box O1 protein
- FOXO3:
-
Forkhead box O3 protein
- mCSA:
-
Muscle cross-sectional area
- MMP:
-
Matrix metalloproteinase
- MuRF-1:
-
Muscle ring finger protein-1
- OCT:
-
Optimal cutting temperature
- PBS:
-
Phosphate-buffered saline
- RFU:
-
Relative fluorescence unit
- RLU:
-
Relative luminescence unit
- RM:
-
Repetition maximum
- RT:
-
Resistance training
- SD:
-
Standard deviation
- T:
-
Trained
- TBST:
-
Tris-buffered saline with 0.1% Tween-20
- TIMP:
-
Tissue inhibitor of metalloproteinase
- UPP:
-
Ubiquitin–proteasome pathway
- US:
-
Ultrasound
- UT:
-
Untrained
References
Alameddine HS (2012) Matrix metalloproteinases in skeletal muscles: friends or foes? Neurobiol Dis 48(3):508–518. https://doi.org/10.1016/j.nbd.2012.07.023
Baehr L, Tunzi M, Bodine S (2014) Muscle hypertrophy is associated with increases in proteasome activity that is independent of MuRF1 and MAFbx expression. Front Physiol 5:69. https://doi.org/10.3389/fphys.2014.00069
Brew K, Nagase H (2010) The tissue inhibitors of metalloproteinases (TIMPs): an ancient family with structural and functional diversity. Biochim Biophys Acta 1:55–71. https://doi.org/10.1016/j.bbamcr.2010.01.003
Chaves TS, Scarpelli MC, Bergamasco JGA, Silva DGD, Medalha Junior RA, Dias NF, Bittencourt D, Carello Filho PC, Angleri V, Nóbrega SR, Roberts MD, Ugrinowitsch C, Libardi CA (2024) Effects of resistance training overload progression protocols on strength and muscle mass. Int J Sports Med. https://doi.org/10.1055/a-2256-5857
Cohen J (1988) Statistical power analysis for the behavioral sciences, 2nd edn. Lawrence Erlbaum, Hillsdale, NJ
Damas F, Libardi CA, Ugrinowitsch C (2018a) The development of skeletal muscle hypertrophy through resistance training: the role of muscle damage and muscle protein synthesis. Eur J Appl Physiol 118(3):485–500. https://doi.org/10.1007/s00421-017-3792-9
Damas F, Phillips SM, Libardi CA, Vechin FC, Lixandrao ME, Jannig PR, Costa LA, Bacurau AV, Snijders T, Parise G, Tricoli V, Roschel H, Ugrinowitsch C (2016a) Resistance training-induced changes in integrated myofibrillar protein synthesis are related to hypertrophy only after attenuation of muscle damage. J Physiol 594(18):5209–5222. https://doi.org/10.1113/JP272472
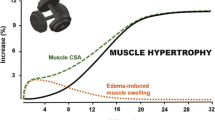
Damas F, Phillips SM, Lixandrão ME, Vechin FC, Libardi CA, Roschel H, Tricoli V, Ugrinowitsch C (2016b) Early resistance training-induced increases in muscle cross-sectional area are concomitant with edema-induced muscle swelling. Eur J Appl Physiol 116(1):49–56. https://doi.org/10.1007/s00421-015-3243-4
Damas F, Ugrinowitsch C, Libardi CA, Jannig PR, Hector AJ, McGlory C, Lixandrão ME, Vechin FC, Montenegro H, Tricoli V, Roschel H, Phillips SM (2018b) Resistance training in young men induces muscle transcriptome-wide changes associated with muscle structure and metabolism refining the response to exercise-induced stress. Eur J Appl Physiol 118(12):2607–2616. https://doi.org/10.1007/s00421-018-3984-y
Davis ME, Gumucio JP, Sugg KB, Bedi A, Mendias CL (2013) MMP inhibition as a potential method to augment the healing of skeletal muscle and tendon extracellular matrix. J Appl Physiol 115(6):884–891. https://doi.org/10.1152/japplphysiol.00137.2013
DeFreitas JM, Beck TW, Stock MS, Dillon MA, Kasishke PR 2nd (2011) An examination of the time course of training-induced skeletal muscle hypertrophy. Eur J Appl Physiol 111(11):2785–2790. https://doi.org/10.1007/s00421-011-1905-4
Féasson L, Stockholm D, Freyssenet D, Richard I, Duguez S, Beckmann JS, Denis C (2002) Molecular adaptations of neuromuscular disease-associated proteins in response to eccentric exercise in human skeletal muscle. J Physiol 543(Pt 1):297–306. https://doi.org/10.1113/jphysiol.2002.018689
Gillies AR, Lieber RL (2011) Structure and function of the skeletal muscle extracellular matrix. Muscle Nerve 44(3):318–331. https://doi.org/10.1002/mus.22094
Godwin JS, Sexton CL, Kontos NJ, Ruple BA, Willoughby DS, Young KC, Mobley CB, Roberts MD (2023a) Extracellular matrix content and remodeling markers do not differ in college-aged men classified as higher and lower responders to resistance training. J Appl Physiol 134:731–741
Godwin JS, Telles GD, Vechin FC, Conceição MS, Ugrinowitsch C, Roberts MD, Libardi CA (2023b) Time course of proteolysis biomarker responses to resistance, high-intensity interval, and concurrent exercise bouts. J Strength Cond Res. https://doi.org/10.1519/jsc.0000000000004550
Goll DE, Thompson VF, Li H, Wei W, Cong J (2003) The calpain system. Physiol Rev 83(3):731–801. https://doi.org/10.1152/physrev.00029.2002
Hedges LV, Olkin I (1985) Estimation of a Single Effect Size: Parametric and Nonparametric Methods. In: Hedges LV, Olkin I (eds) Statistical Methods for Meta-Analysis. Academic Press, San Diego. https://doi.org/10.1016/B978-0-08-057065-5.50010-5
Höhfeld J, Benzing T, Bloch W, Fürst DO, Gehlert S, Hesse M, Hoffmann B, Hoppe T, Huesgen PF, Köhn M, Kolanus W, Merkel R, Niessen CM, Pokrzywa W, Rinschen MM, Wachten D, Warscheid B (2021) Maintaining proteostasis under mechanical stress. EMBO Rep 22(8):52507
Hoier B, Nordsborg N, Andersen S, Jensen L, Nybo L, Bangsbo J, Hellsten Y (2012) Pro- and anti-angiogenic factors in human skeletal muscle in response to acute exercise and training. J Physiol 590(3):595–606. https://doi.org/10.1113/jphysiol.2011.216135
Hyatt HW, powers SK (2020) The role of calpains in skeletal muscle remodeling with exercise and inactivity-induced atrophy. Int J Sports Med 41(14):994–1008. https://doi.org/10.1055/a-1199-7662
Kim J, Lee J (2020) Plasma MMP-9, TIMP-1, and TGF-beta1 responses to exercise-induced muscle injury. Int J Environ Res Public Health 17(2):566. https://doi.org/10.3390/ijerph17020566
Kitajima Y, Tashiro Y, Suzuki N, Warita H, Kato M, Tateyama M, Ando R, Izumi R, Yamazaki M, Abe M, Sakimura K, Ito H, Urushitani M, Nagatomi R, Takahashi R, Aoki M (2014) Proteasome dysfunction induces muscle growth defects and protein aggregation. J Cell Sci 127(Pt 24):5204–5217. https://doi.org/10.1242/jcs.150961
Kitajima Y, Yoshioka K, Suzuki N (2020) The ubiquitin–proteasome system in regulation of the skeletal muscle homeostasis and atrophy: from basic science to disorders. J Physiol Sci 70(1):40. https://doi.org/10.1186/s12576-020-00768-9
Kjaer M (2004) Role of extracellular matrix in adaptation of tendon and skeletal muscle to mechanical loading. Physiol Rev 84(2):649–698. https://doi.org/10.1152/physrev.00031.2003
Kulkarni AS, Peck BD, Walton RG, Kern PA, Mar JC, Windham ST, Bamman MM, Barzilai N, Peterson CA (2020) Metformin alters skeletal muscle transcriptome adaptations to resistance training in older adults. Aging 12(20):104096
Lo Presti R, Hopps E, Caimi G (2017) Gelatinases and physical exercise: a systematic review of evidence from human studies. Medicine 96(37):e8072. https://doi.org/10.1097/md.0000000000008072
Mackey AL, Kjaer M (2017) Connective tissue regeneration in skeletal muscle after eccentric contraction-induced injury. J Appl Physiol 122(3):533–540. https://doi.org/10.1152/japplphysiol.00577.2016
Mackey AL, Kjaer M, Charifi N, Henriksson J, Bojsen-Moller J, Holm L, Kadi F (2009) Assessment of satellite cell number and activity status in human skeletal muscle biopsies. Muscle Nerve 40(3):455–465. https://doi.org/10.1002/mus.21369
Mascher H, Tannerstedt J, Brink-Elfegoun T, Ekblom B, Gustafsson T, Blomstrand E (2008) Repeated resistance exercise training induces different changes in mRNA expression of MAFbx and MuRF-1 in human skeletal muscle. Am J Physiol Endocrinol Metab 294(1):E43-51. https://doi.org/10.1152/ajpendo.00504.2007
Mellgren RL, Miyake K, Kramerova I, Spencer MJ, Bourg N, Bartoli M, Richard I, Greer PA, McNeil PL (2009) Calcium-dependent plasma membrane repair requires m- or mu-calpain, but not calpain-3, the proteasome, or caspases. Biochim Biophys Acta 1793(12):1886–1893. https://doi.org/10.1016/j.bbamcr.2009.09.013
Millward DJ (1995) A protein-stat mechanism for regulation of growth and maintenance of the lean body mass. Nutr Res Rev 8(1):93–120. https://doi.org/10.1079/nrr19950008
Murton AJ, Constantin D, Greenhaff PL (2008) The involvement of the ubiquitin proteasome system in human skeletal muscle remodelling and atrophy. Biochim Biophys Acta 1782(12):730–743. https://doi.org/10.1016/j.bbadis.2008.10.011
Naba A, Clauser KR, Ding H, Whittaker CA, Carr SA, Hynes RO (2016) The extracellular matrix: Tools and insights for the “omics” era. Matrix Biol 49:10–24. https://doi.org/10.1016/j.matbio.2015.06.003
Nakagawa S, Cuthill IC (2007) Effect size, confidence interval and statistical significance: a practical guide for biologists. Biol Rev Camb Philos Soc 82(4):591–605. https://doi.org/10.1111/j.1469-185X.2007.00027.x
Nedergaard A, Vissing K, Overgaard K, Kjaer M, Schjerling P (2007) Expression patterns of atrogenic and ubiquitin proteasome component genes with exercise: effect of different loading patterns and repeated exercise bouts. J Appl Physiol 103(5):1513–1522. https://doi.org/10.1152/japplphysiol.01445.2006
Ogasawara R, Thiebaud RS, Loenneke JP, Loftin M, Abe T (2012) Time course for arm and chest muscle thickness changes following bench press training. Interv Med & App Sci 4(4):217–220. https://doi.org/10.1556/imas.4.2012.4.7
Ojima K, Ono Y, Ottenheijm C, Hata S, Suzuki H, Granzier H, Sorimachi H (2011) Non-proteolytic functions of calpain-3 in sarcoplasmic reticulum in skeletal muscles. J Mol Biol 407(3):439–449. https://doi.org/10.1016/j.jmb.2011.01.057
Osburn SC, Vann CG, Church DD, Ferrando AA, Roberts MD (2021) Proteasome- and calpain-mediated proteolysis, but not autophagy, is required for leucine-induced protein synthesis in C2C12 Myotubes. Physiologia 1(1):22–33. https://doi.org/10.3390/physiologia1010005
Pandit A, Tran TB, Letton M, Cowley E, Gibbs M, Wewege MA, Hagstrom AD (2023) Data informing governing body resistance-training guidelines exhibit sex bias: an audit-based review. Sports Med 53(9):1681–1691. https://doi.org/10.1007/s40279-023-01878-1
Raastad T, Owe SG, Paulsen G, Enns D, Overgaard K, Crameri R, Kiil S, Belcastro A, Bergersen L, Hallen J (2010) Changes in calpain activity, muscle structure, and function after eccentric exercise. Med Sci Sports Exerc 42(1):86–95. https://doi.org/10.1249/MSS.0b013e3181ac7afa
Reeves ND, Maganaris CN, Narici MV (2004) Ultrasonographic assessment of human skeletal muscle size. Eur J Appl Physiol 91(1):116–118
Roberts MD, Haun CT, Mobley CB, Mumford PW, Romero MA, Roberson PA, Vann CG, McCarthy JJ (2018) Physiological differences between low versus high skeletal muscle hypertrophic responders to resistance exercise training: current perspectives and future research directions. Front Physiol 9:834. https://doi.org/10.3389/fphys.2018.00834
Roberts MD, McCarthy JJ, Hornberger TA, Phillips SM, Mackey AL, Nader GA, Boppart MD, Kavazis AN, Reidy PT, Ogasawara R, Libardi CA, Ugrinowitsch C, Booth FW, Esser KA (2023) Mechanisms of mechanical overload-induced skeletal muscle hypertrophy: current understanding and future directions. Physiol Rev 103(4):2679–2757. https://doi.org/10.1152/physrev.00039.2022
Rullman E, Norrbom J, Strömberg A, Wågsäter D, Rundqvist H, Haas T, Gustafsson T (2009) Endurance exercise activates matrix metalloproteinases in human skeletal muscle. J Appl Physiol 106(3):804–812. https://doi.org/10.1152/japplphysiol.90872.2008
Rullman E, Rundqvist H, Wågsäter D, Fischer H, Eriksson P, Sundberg CJ, Jansson E, Gustafsson T (2007) A single bout of exercise activates matrix metalloproteinase in human skeletal muscle. J Appl Physiol 102(6):2346–2351. https://doi.org/10.1152/japplphysiol.00822.2006
Scarpelli MC, Nóbrega SR, Santanielo N, Alvarez IF, Otoboni GB, Ugrinowitsch C, Libardi CA (2022) Muscle hypertrophy response is affected by previous resistance training volume in trained individuals. J Strength and Cond Res 36(4):1153–1157. https://doi.org/10.1519/jsc.0000000000003558
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9(7):671–675. https://doi.org/10.1038/nmeth.2089
Schwanhäusser B, Busse D, Li N, Dittmar G, Schuchhardt J, Wolf J, Chen W, Selbach M (2011) Global quantification of mammalian gene expression control. Nature 473(7347):337–342. https://doi.org/10.1038/nature10098
Stefanetti RJ, Lamon S, Wallace M, Vendelbo MH, Russell AP, Vissing K (2015) Regulation of ubiquitin proteasome pathway molecular markers in response to endurance and resistance exercise and training. Pflugers Arch 467(7):1523–1537. https://doi.org/10.1007/s00424-014-1587-y
Stupka N, Tarnopolsky MA, Yardley NJ, Phillips SM (2001) Cellular adaptation to repeated eccentric exercise-induced muscle damage. J Appl Physiol 91(4):1669–1678. https://doi.org/10.1152/jappl.2001.91.4.1669
Wen Y, Murach KA, Vechetti IJ Jr, Fry CS, VickeryPeterson CCA, McCarthy JJ, Campbell KS (2018) MyoVision: software for automated high-content analysis of skeletal muscle immunohistochemistry. J Appl Physiol 124(1):40–51. https://doi.org/10.1152/japplphysiol.00762.2017
Willoughby DS, Taylor M, Taylor L (2003) Glucocorticoid receptor and ubiquitin expression after repeated eccentric exercise. Med Sci Sports Exerc 35(12):2023–2031. https://doi.org/10.1249/01.MSS.0000099100.83796.77
Wolfe RR (2006) The underappreciated role of muscle in health and disease. Am J Clin Nutr 84(3):475–482. https://doi.org/10.1093/ajcn/84.3.475
Acknowledgements
We greatly acknowledge all participants of this study. We also appreciate the technical assistance of the staff of the Nutrabolt Applied and Molecular Laboratory at Auburn University and the assistance of Dr. Felipe Cassaro Vechin in muscle biopsy procedures.
Funding
This work was supported by the Coordination for the Improvement of Higher Education Personnel (CAPES) (#88887.634296/2021–00 and #88887.717127/2022–00 to M.C.S.; #88887.717126/2022–00 to J.G.B.; 88,887.634297/2021–00 to T.S.C.), by the National Council for Scientific and Technological Development (CNPq) (#311387/2021–7 to C.A.L.; #425917/2018–5 to C.U.; #140753/2020–6 to J.G.B.) and by discretionary laboratory funds of C.A.L. and M.D.R.
Author information
Authors and Affiliations
Contributions
M.C.S., C.U., M.D.R., and C.A.L. conceived and designed this research. M.C.S, J.G.B., and T.S.C recruited participants and coordinated training sessions and data collection. D.G.S, D.F.B., J.G.B, N.F.D., P.C.C.F., R.M.J., T.S.C., V.A. and L.A.R.C. performed training sessions and data collection. M.C.S., J.G.B., J.S.G., and P.H.C.M. conducted laboratory assays and experiments. M.C.S., C.A.L., and M.D.R. analyzed and interpreted the data. M.C.S. primarily drafted the manuscript, and C.A.L. and M.D.R. edited it. All authors revised the manuscript and provided intellectual contributions. All authors approved the final version of the manuscript. All authors qualify for authorship, and all those who qualify are listed.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no perceived or potential conflicts of interest to declare, financial or otherwise.
Ethical approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Ethics Committee of the Federal University of São Carlos (no. 56259622.0.0000.5504).
Consent to participate
Informed consent was obtained from all individual participants included in the study.
Additional information
Communicated by Mark Elisabeth Willems.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Scarpelli, M.C., Bergamasco, J.G.A., Godwin, J.S. et al. Resistance training-induced changes in muscle proteolysis and extracellular matrix remodeling biomarkers in the untrained and trained states. Eur J Appl Physiol (2024). https://doi.org/10.1007/s00421-024-05484-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00421-024-05484-5