Abstract.
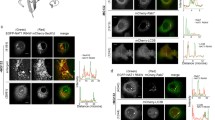
To analyze the cellular response caused by the overexpression of proteins in subcellular compartments, we constructed four expression clones encoding wild-type peroxisomal urate oxidase (UO), truncated UO lacking the peroxisomal targeting signal (UOdC), and chimeric UOs with a mitochondrial targeting signal (MTS) at the N-terminus of UOdC (MUOdC) or UO (MUO). After transfection, we examined COS-1 and HEK293 cells by immunofluorescence and immunoelectron microscopy, transmission electron microscopy, and pulse-chase experiments. The overexpressed UO and UOdC formed large electron-dense aggregates with no limiting membrane in both the cytoplasm and the nucleus. The UO aggregates exhibited the crystalloid structure quite similar to that of rat liver peroxisomal cores, whereas the UOdC aggregates formed a loose mass consisting of small dense substructures. The overexpressed MUOdC and MUO, on the other hand, formed other types of aggregates which were distributed in the cytoplasm. They consisted of tubular and circular membrane structures, which were morphologically confirmed to be derived from the endoplasmic reticulum (ER). No immunolabeling signals for MUOdC and MUO were present free in the cytoplasm and most of them were associated with membrane structures, suggesting that overexpressed UO containing the MTS attached to the ER membranes soon after synthesis and segregated from the cytosolic compartment. All the UO aggregates were stained for ubiquitin antigen. Pulse-chase experiments in combination with proteasome inhibitors suggested that proteasomes did not contribute to the degradation of these products.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Electronic Publication
Rights and permissions
About this article
Cite this article
Yokota, S., Kamijo, K. & Oda, T. Aggregate formation and degradation of overexpressed wild-type and mutant urate oxidase proteins. Quality control of organelle-destined proteins by the endoplasmic reticulum. Histochem Cell Biol 114, 433–446 (2000). https://doi.org/10.1007/s004180000208
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s004180000208