Abstract
Purpose
To investigate the differences in the eyelid and buccal microbiomes between patients receiving long-term prostaglandin analogs for open-angle glaucoma (PG-OAG) and naïve-OAG patients by using metagenomics.
Methods
Eyelid and buccal samples were collected from 30 PG-OAG and 32 naïve-OAG patients. The taxonomic composition of the microbiome was obtained via 16S rRNA gene sequencing, operational taxonomic unit analysis, and diversity analysis. Differential gene expression analysis (DEG) and Bland–Altman (MA) plots were used to determine taxon differences between the microbiomes of PG-OAG and naïve-OAG patients.
Results
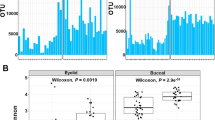
The eyelid microbiome showed marginally significant differences, while the alpha-diversity of the buccal microbiome showed significant differences between PG-OAG and naïve-OAG patients. However, the beta-diversity of both eyelid and buccal microbiomes was higher in PG-OAG patients than in naïve-OAG patients. The MA plot showed cluster differences in the eyelid microbiome. DEG analysis of the eyelid microbiome revealed various taxa differences, including enrichment of Azomonas, Pseudomonas, and Granulicatella in PG-OAG patients over naïve-OAG patients, as well as significant depletion of Delftia and Rothia. In the buccal microbiome in PG-OAG patients, taxa such as Rikenella and Stenotrophomonas were significantly enriched.
Conclusion
Our findings suggest that the eyelid microbiome differs between PG-OAG and naïve-OAG patients, raising concerns regarding the eyelid environment in patients receiving these drugs. The overexpressed microbiome in the eyelid area suggests that microbiota may change after the administration of glaucoma medications in OAG.





Similar content being viewed by others
Data availability
Data are available on reasonable request.
Code availability
Not applicable.
References
Quigley HA, Addicks EM, Green WR et al (1981) Optic nerve damage in human glaucoma. II. The site of injury and susceptibility to damage. Arch Ophthalmol 99:635–649
Musch DC, Gillespie BW, Niziol LM et al (2011) Intraocular pressure control and long-term visual field loss in the Collaborative Initial Glaucoma Treatment Study. Ophthalmology 118:1766–1773. https://doi.org/10.1016/j.ophtha.2011.01.047
Ohtani S, Shimizu K, Nejima R et al (2017) Conjunctival bacteria flora of glaucoma patients during long-term administration of prostaglandin analog drops. Invest Ophthalmol Vis Sci 58:3991–3996. https://doi.org/10.1167/iovs.16-20853
Qin J, Li R, Raes J et al (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464:59–65. https://doi.org/10.1038/nature08821
Team NIHHMPA (2019) A review of 10 years of human microbiome research activities at the US National Institutes of Health, Fiscal Years 2007–2016. Microbiome 7:31. https://doi.org/10.1186/s40168-019-0620-y
Li Z, Gong Y, Chen S et al (2019) Comparative portrayal of ocular surface microbe with and without dry eye. J Microbiol 57:1025–1032. https://doi.org/10.1007/s12275-019-9127-2
Dong X, Wang Y, Wang W et al (2019) Composition and diversity of bacterial community on the ocular surface of patients with meibomian gland dysfunction. Invest Ophthalmol Vis Sci 60:4774–4783. https://doi.org/10.1167/iovs.19-27719
Wen X, Hu X, Miao L et al (2018) Epigenetics, microbiota, and intraocular inflammation: new paradigms of immune regulation in the eye. Prog Retin Eye Res 64:84–95. https://doi.org/10.1016/j.preteyeres.2018.01.001
Nayyar A, Gindina S, Barron A et al (2020) Do epigenetic changes caused by commensal microbiota contribute to development of ocular disease? A review of evidence. Hum Genomics 14:11. https://doi.org/10.1186/s40246-020-00257-5
Kountouras J, Zavos C, Zeglinas C et al (2015) Helicobacter pylori-related impact on glaucoma pathophysiology. Invest Ophthalmol Vis Sci 56:8029–8030. https://doi.org/10.1167/iovs.15-17969
Zeng J, Liu H, Liu X et al (2015) The relationship between Helicobacter pylori infection and open-angle glaucoma: a meta-analysis. Invest Ophthalmol Vis Sci 56:5238–5245. https://doi.org/10.1167/iovs.15-17059
Kim JM, Kim SH, Park KH et al (2011) Investigation of the association between Helicobacter pylori infection and normal tension glaucoma. Invest Ophthalmol Vis Sci 52:665–668. https://doi.org/10.1167/iovs.10-6096
Chen H, Cho KS, Vu THK et al (2018) Commensal microflora-induced T cell responses mediate progressive neurodegeneration in glaucoma. Nat Commun 9:3209. https://doi.org/10.1038/s41467-018-05681-9
Polla D, Astafurov K, Hawy E et al (2017) A pilot study to evaluate the oral microbiome and dental health in primary open-angle glaucoma. J Glaucoma 26:320–327. https://doi.org/10.1097/IJG.0000000000000465
Astafurov K, Elhawy E, Ren L et al (2014) Oral microbiome link to neurodegeneration in glaucoma. PLoS ONE 9:e104416. https://doi.org/10.1371/journal.pone.0104416
Honda R, Toshida H, Suto C et al (2011) Effect of long-term treatment with eyedrops for glaucoma on conjunctival bacterial flora. Infect Drug Resist 4:191–196. https://doi.org/10.2147/IDR.S24250
Klindworth A, Pruesse E, Schweer T et al (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41:e1. https://doi.org/10.1093/nar/gks808
Edgar RC, Haas BJ, Clemente JC et al (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Bolyen E, Rideout JR, Dillon MR et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Lopez-Garcia A, Pineda-Quiroga C, Atxaerandio R et al (2018) Comparison of Mothur and QIIME for the analysis of Rumen microbiota composition based on 16S rRNA amplicon sequences. Front Microbiol 9:3010. https://doi.org/10.3389/fmicb.2018.03010
Cole JR, Wang Q, Fish JA et al (2014) Ribosomal Database Project: data and tools for high throughput rRNA analysis. Nucleic Acids Res 42:D633-642. https://doi.org/10.1093/nar/gkt1244
Sun J, Nishiyama T, Shimizu K et al (2013) TCC: an R package for comparing tag count data with robust normalization strategies. BMC Bioinformatics 14:219. https://doi.org/10.1186/1471-2105-14-219
Benjamini YHY (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Series B 57:289–300
Hochberg Y, Benjamini Y (1990) More powerful procedures for multiple significance testing. Stat Med 9:811–818
Ozkan J, Coroneo M, Willcox M et al (2018) Identification and visualization of a distinct microbiome in ocular surface conjunctival tissue. Invest Ophthalmol Vis Sci 59:4268–4276. https://doi.org/10.1167/iovs.18-24651
Ozkan J, Willcox MD (2019) The ocular microbiome: molecular characterisation of a unique and low microbial environment. Curr Eye Res 1-10. https://doi.org/10.1080/02713683.2019.1570526
Lee J, Iwasaki T, Ohtani S et al (2020) Benzalkonium chloride resistance in staphylococcus epidermidis on the ocular surface of glaucoma patients under long-term administration of eye drops. Transl Vis Sci Technol 9:9. https://doi.org/10.1167/tvst.9.8.9
Asbell PA, DeCory HH (2018) Antibiotic resistance among bacterial conjunctival pathogens collected in the Antibiotic Resistance Monitoring in Ocular Microorganisms (ARMOR) surveillance study. PLoS ONE 13:e0205814. https://doi.org/10.1371/journal.pone.0205814
Ku CA, Forcina B, LaSala PR et al (2015) Granulicatella adiacens, an unusual causative agent in chronic dacryocystitis. J Ophthalmic Inflamm Infect 5:12. https://doi.org/10.1186/s12348-015-0043-2
Zhang H, Zhao F, Hutchinson DS et al (2017) Conjunctival microbiome changes associated with soft contact lens and orthokeratology lens wearing. Invest Ophthalmol Vis Sci 58:128–136. https://doi.org/10.1167/iovs.16-20231
Asao K, Hashida N, Ando S et al (2019) Conjunctival dysbiosis in mucosa-associated lymphoid tissue lymphoma. Sci Rep 9:8424. https://doi.org/10.1038/s41598-019-44861-5
Hayes RA, Bennett HY, O’Hagan S (2017) Rothia dentocariosa endophthalmitis following intravitreal injection-a case report. J Ophthalmic Inflamm Infect 7:24. https://doi.org/10.1186/s12348-017-0142-3
Garcia-Lopez M, Meier-Kolthoff JP, Tindall BJ et al (2019) Analysis of 1,000 type-strain genomes improves taxonomic classification of bacteroidetes. Front Microbiol 10:2083. https://doi.org/10.3389/fmicb.2019.02083
Gong H, Zhang S, Li Q et al (2020) Gut microbiota compositional profile and serum metabolic phenotype in patients with primary open-angle glaucoma. Exp Eye Res 191:107921. https://doi.org/10.1016/j.exer.2020.107921
Brooke JS, Di Bonaventura G, Berg G et al (2017) Editorial: a multidisciplinary look at stenotrophomonas maltophilia: an emerging multi-drug-resistant global opportunistic pathogen. Front Microbiol 8:1511. https://doi.org/10.3389/fmicb.2017.01511
Wen X, Miao L, Deng Y et al (2017) The influence of age and sex on ocular surface microbiota in healthy adults. Invest Ophthalmol Vis Sci 58:6030–6037. https://doi.org/10.1167/iovs.17-22957
Acknowledgments
This study was supported by a VHS Medical Center Research Grant (grant number: VHSMC 19022), Republic of Korea. The authors thank the Theragen Bio for providing technical support for genome sequencing and metagenome analysis
Funding
This study was supported by a VHS Medical Center Research Grant (grant number: VHSMC 19022), Republic of Korea.
Author information
Authors and Affiliations
Contributions
S-HL, JHS, and J-WL: study design and conception, data interpretation, and writing the manuscript; YL: data analysis and interpretation, writing the manuscript; JHS: study design and conception, data analysis and interpretation, and writing and revising the manuscript. All authors finally approved the manuscript and agreed to be accountable.
Corresponding author
Ethics declarations
Ethics approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. The protocol was approved by the Institutional Review Board (IRB) of each center: Veterans Health Service Medical Center, Korea (IRB No. 2018–12-015), Pusan National University Hospital, (IRB No. H-1904–001-077), Daegu Veterans Health Service Medical Center (IRB No. 2019–001), and Pusan National University Yangsan Hospital (IRB No.05–2019-022).
Consent to participate
Written informed consent was obtained from all subjects prior to their participation in the study.
Consent for publication
Not applicable.
Competing interests
The authors have no conflicts of interest to declare that are relevant to the content of this article.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
417_2021_5218_MOESM1_ESM.xlsx
Supplementary Table S1. Differential gene expression analysis of the eyelid microbiome in open-angle glaucoma patients treated with prostaglandin analogs compared to naïve-open-angle glaucoma patients (XLSX 113 KB)
417_2021_5218_MOESM2_ESM.xlsx
Supplementary Table S2. Differential gene expression analysis of the buccal microbiome in open-angle glaucoma patients treated with prostaglandin analogs compared to naïve-open-angle glaucoma patients (XLSX 119 KB)
Rights and permissions
About this article
Cite this article
Lim, SH., Shin, J.H., Lee, JW. et al. Differences in the eyelid and buccal microbiome of glaucoma patients receiving long-term administration of prostaglandin analog drops. Graefes Arch Clin Exp Ophthalmol 259, 3055–3065 (2021). https://doi.org/10.1007/s00417-021-05218-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00417-021-05218-9