Abstract
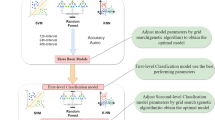
Determining postmortem interval (PMI) is one of the most challenging and essential endeavors in forensic science. Developments in PMI estimation can take advantage of machine learning techniques. Currently, applying an algorithm to obtain information on multiple organs and conducting joint analysis to accurately estimate PMI are still in the early stages. This study aimed to establish a multi-organ stacking model that estimates PMI by analyzing differential compounds of four organs in rats. In a total of 140 rats, skeletal muscle, liver, lung, and kidney tissue samples were collected at each time point after death. Ultra-performance liquid chromatography coupled with high-resolution mass spectrometry was used to determine the compound profiles of the samples. The original data were preprocessed using multivariate statistical analysis to determine discriminant compounds. In addition, three interrelated and increasingly complex patterns (single organ optimal model, single organ stacking model, multi-organ stacking model) were established to estimate PMI. The accuracy and generalized area under the receiver operating characteristic curve of the multi-organ stacking model were the highest at 93% and 0.96, respectively. Only 1 of the 14 external validation samples was misclassified by the multi-organ stacking model. The results demonstrate that the application of the multi-organ combination to the stacking algorithm is a potential forensic tool for the accurate estimation of PMI.







Similar content being viewed by others
Data and code availability
Part of the relevant code is within the Supporting Information files. All relevant data and code are available from the corresponding author upon request.
References
Peyron PA, Lehmann S, Delaby C, Baccino E, Hirtz C (2019) Biochemical markers of time since death in cerebrospinal fluid: a first step towards “Forensomics.” Crit Rev Clin Lab Sci 56(4):274–286. https://doi.org/10.1080/10408363.2019.1619158
De-Giorgio F, Ciasca G, D’Amico R, Trombatore P, D’Angelo A, Rinaldi P, Milano F, Locci E, De Spirito M, d’Aloja E, Colosimo C, Pascali VL (2020) An evaluation of the objectivity and reproducibility of shear wave elastography in estimating the post-mortem interval: a tissue biomechanical perspective. Int J Legal Med 134(5):1939–1948. https://doi.org/10.1007/s00414-020-02370-5
De-Giorgio F, Nardini M, Foti F, Minelli E, Papi M, d’Aloja E, Pascali VL, De Spirito M, Ciasca G (2019) A novel method for post-mortem interval estimation based on tissue nano-mechanics. Int J Legal Med 133(4):1133–1139. https://doi.org/10.1007/s00414-019-02034-z
Mansour H, Krebs O, Pinnschmidt HO, Griem N, Hammann-Ehrt I, Püschel K (2019) Factors affecting dental DNA in various real post-mortem conditions. Int J Legal Med 133(6):1751–1759. https://doi.org/10.1007/s00414-019-02151-9
Scrivano S, Sanavio M, Tozzo P, Caenazzo L (2019) Analysis of RNA in the estimation of post-mortem interval: a review of current evidence. Int J Legal Med 133(6):1629–1640. https://doi.org/10.1007/s00414-019-02125-x
Paltian JJ, da Fonseca CAR, Pinz MP, Luchese C, Antunes Wilhelm E (2019) Post-mortem interval estimative through determination of catalase and Δ-aminolevulinate dehydratase activities in hepatic, renal, skeletal muscle and cerebral tissues of Swiss mice. Biomark Biochem Indicat Exposure Response Suscept Chem 24(5):478–483. https://doi.org/10.1080/1354750x.2019.1619837
Choi KM, Zissler A, Kim E, Ehrenfellner B, Cho E, Lee SI, Steinbacher P, Yun KN, Shin JH, Kim JY, Stoiber W, Chung H, Monticelli FC, Kim JY, Pittner S (2019) Postmortem proteomics to discover biomarkers for forensic PMI estimation. Int J Legal Med 133(3):899–908. https://doi.org/10.1007/s00414-019-02011-6
Mickleburgh HL, Schwalbe EC, Bonicelli A, Mizukami H, Sellitto F, Starace S, Wescott DJ, Carter DO, Procopio N (2021) Human bone proteomes before and after decomposition: investigating the effects of biological variation and taphonomic alteration on bone protein profiles and the implications for forensic proteomics. J Proteome Res 20(5):2533–2546. https://doi.org/10.1021/acs.jproteome.0c00992
Pesko BK, Weidt S, McLaughlin M, Wescott DJ, Torrance H, Burgess K, Burchmore R (2020) Postmortomics: the potential of untargeted metabolomics to highlight markers for time since death. Omics: J Integ Biol 24(11):649–659. https://doi.org/10.1089/omi.2020.0084
Locci E, Stocchero M, Gottardo R, De-Giorgio F, Demontis R, Nioi M, Chighine A, Tagliaro F, d’Aloja E (2021) Comparative use of aqueous humour (1)H NMR metabolomics and potassium concentration for PMI estimation in an animal model. Int J Legal Med 135(3):845–852. https://doi.org/10.1007/s00414-020-02468-w
Baptista A, Pedrosa M, Curate F, Ferreira MT, Marques MPM (2022) Estimation of the post-mortem interval in human bones by infrared spectroscopy. Int J Legal Med 136(1):309–317. https://doi.org/10.1007/s00414-021-02641-9
Madea B (2016) Methods for determining time of death. Forensic Sci Med Pathol 12(4):451–485. https://doi.org/10.1007/s12024-016-9776-y
Wang Y, Wang Y, Wang M, Xu W, Zhang Y, Wang J (2021) Forensic entomology in China and its challenges. Insects 12(3):230. https://doi.org/10.3390/insects12030230
Dash HR, Das S (2020) Thanatomicrobiome and epinecrotic community signatures for estimation of post-mortem time interval in human cadaver. Appl Microbiol Biotechnol 104(22):9497–9512. https://doi.org/10.1007/s00253-020-10922-3
Díez López C, Vidaki A, Kayser M (2022) Integrating the human microbiome in the forensic toolkit: Current bottlenecks and future solutions. Forensic Sci Int Genet 56:102627. https://doi.org/10.1016/j.fsigen.2021.102627
Donaldson AE, Lamont IL (2015) Metabolomics of post-mortem blood: identifying potential markers of post-mortem interval. Metabolomics 11(1):237–245. https://doi.org/10.1007/s11306-014-0691-5
Castillo-Peinado LS, Luque de Castro MD (2016) Present and foreseeable future of metabolomics in forensic analysis. Anal Chim Acta 925:1–15. https://doi.org/10.1016/j.aca.2016.04.040
Mora-Ortiz M, Trichard M, Oregioni A, Claus SP (2019) Thanatometabolomics: introducing NMR-based metabolomics to identify metabolic biomarkers of the time of death. Metabolomics 15(3):37. https://doi.org/10.1007/s11306-019-1498-1
Kang Y-R, Park YS, Park YC, Yoon SM, JongAhn H, Kim G, Kwon SW (2012) UPLC/Q-TOF MS based metabolomics approach to post-mortem-interval discrimination: mass spectrometry based metabolomics approach. J Pharm Investig 42(1):41–46. https://doi.org/10.1007/s40005-012-0006-7
Locci E, Stocchero M, Noto A, Chighine A, Natali L, Napoli PE, Caria R, De-Giorgio F, Nioi M, d’Aloja E (2019) A (1)H NMR metabolomic approach for the estimation of the time since death using aqueous humour: an animal model. Metabolomics 15(5):76. https://doi.org/10.1007/s11306-019-1533-2
Zhu W, Zhai X, Zheng Z, Sun K, Yang M, Mo Y (2021) New contributions to the relationship between sequential changes of ATP-related metabolites and post-mortem interval in rats. Leg Med (Tokyo) 48:101809. https://doi.org/10.1016/j.legalmed.2020.101809
Zelentsova EA, Yanshole LV, Melnikov AD, Kudryavtsev IS, Novoselov VP, Tsentalovich YP (2020) Post-mortem changes in metabolomic profiles of human serum, aqueous humor and vitreous humor. Metabolomics 16(7):80. https://doi.org/10.1007/s11306-020-01700-3
Go A, Shim G, Park J, Hwang J, Nam M, Jeong H, Chung H (2019) Analysis of hypoxanthine and lactic acid levels in vitreous humor for the estimation of post-mortem interval (PMI) using LC-MS/MS. Forensic Sci Int 299:135–141. https://doi.org/10.1016/j.forsciint.2019.03.024
Locci E, Bazzano G, Chighine A, Locco F, Ferraro E, Demontis R, d’Aloja E (2020) Forensic NMR metabolomics: one more arrow in the quiver. Metabolomics 16(11):118. https://doi.org/10.1007/s11306-020-01743-6
Porto LF, Lima LNC, Franco A, Pianto D, Machado CEP, Vidal FB (2020) Estimating sex and age from a face: a forensic approach using machine learning based on photo-anthropometric indexes of the Brazilian population. Int J Legal Med 134(6):2239–2259. https://doi.org/10.1007/s00414-020-02346-5
Chaves D, Fidalgo E, Alegre E, Alaiz-Rodríguez R, Jáñez-Martino F, Azzopardi G (2020) Assessment and estimation of face detection performance based on deep learning for forensic applications. Sensors (Basel, Switzerland) 20(16):4491. https://doi.org/10.3390/s20164491
Khanagar SB, Vishwanathaiah S, Naik S, A Al-Kheraif A, Devang Divakar D, Sarode SC, Bhandi S, Patil S (2021) Application and performance of artificial intelligence technology in forensic odontology - a systematic review. Leg Med (Tokyo) 48:101826. https://doi.org/10.1016/j.legalmed.2020.101826
Ezzat A, Wu M, Li XL, Kwoh CK (2017) Drug-target interaction prediction using ensemble learning and dimensionality reduction. Methods (San Diego, Calif) 129:81–88. https://doi.org/10.1016/j.ymeth.2017.05.016
Kwon H, Park J, Lee Y (2019) Stacking ensemble technique for classifying breast cancer. Healthcare Inform Res 25(4):283–288. https://doi.org/10.4258/hir.2019.25.4.283
van der Laan MJ, Polley EC, Hubbard AE (2007) Super learner. Statistical applications in genetics and molecular biology 6:Article25. https://doi.org/10.2202/1544-6115.1309
Zhang X, Liang B, Zhang J, Hao X, Xu X, Chang HM, Leung PCK, Tan J (2021) Raman spectroscopy of follicular fluid and plasma with machine-learning algorithms for polycystic ovary syndrome screening. Mol Cell Endocrinol 523:111139. https://doi.org/10.1016/j.mce.2020.111139
Yy A, Long WB, Ying HA, Yan WA, Lh C, Sn A (2020) Classification of Parkinson’s disease based on multi-modal features and stacking ensemble learning. J Neurosci Methods 350:109019. https://doi.org/10.1016/j.jneumeth.2020.109019
Kim C, You SC, Reps JM, Cheong JY, Park RW (2020) Machine-learning model to predict the cause of death using a stacking ensemble method for observational data. J Am Med Inform Assoc 28(6):1098–1107. https://doi.org/10.1093/jamia/ocaa277
Sato T, Zaitsu K, Tsuboi K, Nomura M, Kusano M, Shima N, Abe S, Ishii A, Tsuchihashi H, Suzuki K (2015) A preliminary study on postmortem interval estimation of suffocated rats by GC-MS/MS-based plasma metabolic profiling. Anal Bioanal Chem 407(13):3659–3665. https://doi.org/10.1007/s00216-015-8584-7
Wu Z, Lu X, Chen F, Dai X, Ye Y, Yan Y, Liao L (2018) Estimation of early postmortem interval in rats by GC-MS-based metabolomics. Leg Med (Tokyo) 31:42–48. https://doi.org/10.1016/j.legalmed.2017.12.014
Dai X, Fan F, Ye Y, Lu X, Chen F, Wu Z, Liao L (2019) An experimental study on investigating the postmortem interval in dichlorvos poisoned rats by GC/MS-based metabolomics. Leg Med (Tokyo) 36:28–36. https://doi.org/10.1016/j.legalmed.2018.10.002
Zhao Y, Wu C, Zhu Y, Zhou C, Xiong Z, Samy Eweys A, Zhou H, Dong Y, Xiao X (2021) Metabolomics strategy for revealing the components in fermented barley extracts with Lactobacillus plantarum dy-1. Food Res Int (Ottawa, Ont) 139:109808. https://doi.org/10.1016/j.foodres.2020.109808
Du T, Lin Z, Xie Y, Ye X, Tu C, Jin K, Xie J, Shen Y (2018) Metabolic profiling of femoral muscle from rats at different periods of time after death. PLoS ONE 13(9):e0203920. https://doi.org/10.1371/journal.pone.0203920
Aiello D, Lucà F, Siciliano C, Frati P, Fineschi V, Rongo R, Napoli A (2021) Analytical strategy for MS-based thanatochemistry to estimate postmortem interval. J Proteome Res 20(5):2607–2617. https://doi.org/10.1021/acs.jproteome.0c01038
Cao J, Li J, Gu Z, Niu JJ, An GS, Jin QQ, Wang YY, Huang P, Sun JH (2022) Combined metabolomics and machine learning algorithms to explore metabolic biomarkers for diagnosis of acute myocardial ischemia. Int J Legal Med. https://doi.org/10.1007/s00414-022-02816-y
Liu Z, Li W, Geng L, Sun L, Wang Q, Yu Y, Yan P, Liang C, Ren J, Song M, Zhao Q, Lei J, Cai Y, Li J, Yan K, Wu Z, Chu Q, Li J, Wang S, Li C, Han JJ, Hernandez-Benitez R, Shyh-Chang N, Belmonte JCI, Zhang W, Qu J, Liu GH (2022) Cross-species metabolomic analysis identifies uridine as a potent regeneration promoting factor. Cell Discover 8(1):6. https://doi.org/10.1038/s41421-021-00361-3
Acknowledgements
We thank the National Natural Science Foundation of China. Thanks for the equipment support provided by the School of Forensic Medicine, Hebei Medical University, and the assistance of Professor Chun-ling Ma and Professor Di Wen.
Funding
This work was supported by grants from the National Key R&D Program of China (grant no. 2018YFC0807204) and the National Natural Science Foundation of China (81971795, 81901924).
Author information
Authors and Affiliations
Contributions
Jun-hong Sun, Ying-yuan Wang, and Jie Cao conceived the idea; Xiao-jun Lu, Jian Li, and Xue Wei conducted the data analysis and drafted the manuscript; Na Li, Li-hong Dang, and Guo-shuai An established the animal model; Qiu-xiang Du and Qian-qian Jin conceived the study and performed some interpretation of the results. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This article does not contain any studies with human participants performed by any of the authors and there is no patient consent. All experiments of rats were performed according to the ‘Guiding Principles in the Use and Care of Animals’ (NIH publication no. 85–23, revised 1996) and were approved by the Institutional Animal Care and Use Committee of Shanxi Medical University, China (rat batch number: SCXK [Jin] [2009-0001].
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
If there is any conflict, please consider Ph.D. Jun-hong Sun as the primary corresponding author.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lu, Xj., Li, J., Wei, X. et al. A novel method for determining postmortem interval based on the metabolomics of multiple organs combined with ensemble learning techniques. Int J Legal Med 137, 237–249 (2023). https://doi.org/10.1007/s00414-022-02844-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-022-02844-8