Abstract
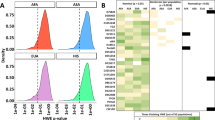
The recently developed probabilistic genotyping software package MaSTR™ (SoftGenetics LLC) was used to develop statistical weight estimates for a variety of two-person STR mixture profiles with differentially degraded sources of DNA. A total of 864 analyses, on 144 two-person profiles, were performed. Mixture ratios ranged from 1:1 to 1:10, including pristine sources of DNA and various combinations of artificially degraded DNA (average size fragments of 150 or 250 bps). Quantities of DNA template were varied (0.1 to 0.5 ngs of total input) and MaSTR™ analysis was performed with eight chains of 10,000 or 40,000 iterations, with or without a conditioning profile to generate likelihood ratio (LR) values. Overall, the software performed as expected. The resulting log(LR) values for pristine mixture profiles were typically greater than 1030. Lower-quality mixture data associated with sources of DNA at ~ 0.05 ngs for each contributor resulted in peak imbalance and allelic dropout which reduced the weight in support of a contributor. This was exacerbated by higher levels of degradation, with some instances resulting in log(LR) values in support of an exclusion. These studies provide additional support for the use of probabilistic genotyping software solutions in forensic investigations, addressing concerns raised by the President’s Council of Advisors on Science and Technology (PCAST).





Similar content being viewed by others
Data availability
Data files associated with this study cannot be made available as consent to do so was not obtained.
Code availability
The code for MaSTR™ is proprietary.
References
Coble MD, Bright JA (2019) Probabilistic genotyping software: an overview. Forensic Sci Int Genet 38:219–224. https://doi.org/10.1016/j.fsigen.2018.11.009
Benschop CCG, Hoogenboom J, Hovers P, Slagter M, Kruise D, Parag R et al (2019) DNAxs/DNAStatistX: development and validation of a software suite for the data management and probabilistic interpretation of DNA profiles. Forensic Sci Int Genet 42:81–89. https://doi.org/10.1016/j.fsigen.2019.06.015
Buckleton JS, Bright JA, Gittelson S, Moretti TR, Onorato A, Bieber FR et al (2019) The probabilistic genotyping software STRmix: utility and evidence for its validity. J Forensic Sci 64:393–405. https://doi.org/10.1111/1556-4029.13898
Bright JA, Cheng K, Kerr Z, McGovern C, Kelly H, Moretti TR et al (2019) STRmix collaborative exercise on DNA mixture interpretation. Forensic Sci Int Genet 40:1–8. https://doi.org/10.1016/j.fsigen.2019.01.006
Moretti TR, Just RS, Kehl SC, Willis LE, Buckleton JS, Bright JA et al (2017) Internal validation of STRmix™ for the interpretation of single source and mixed DNA profiles. Forensic Sci Int Genet 29:126–144. https://doi.org/10.1016/j.fsigen.2017.04.004
Haned H, Gill P, Lohmueller K, Inman K, Rudin N (2016) Validation of probabilistic genotyping software for use in forensic casework: definitions and illustrations. Sci Justice 56:104–108. https://doi.org/10.1016/j.scijus.2015.11.007
Bleka Ø, Storvik G, Gill P (2016) EuroForMix: an open source software based on a continuous model to evaluate STR DNA profiles from a mixture of contributors with artifacts. Forensic Sci Int Genet 21:35–44. https://doi.org/10.1016/j.fsigen.2015.11.008
Bright JA, Taylor D, McGovern C, Cooper S, Russell L, Abarno D et al (2016) Developmental validation of STRmix, expert software for the interpretation of forensic DNA profiles. Forensic Sci Int Genet 23:226–239. https://doi.org/10.1016/j.fsigen.2016.05.007
Noël S, Noël J, Granger D, Lefebvre JF, Séguin D (2019) STRmix put to the test: 300,000 non-contributor profiles compared to four-contributor DNA mixtures and the impact of replicates. Forensic Sci Int Genet 41:24–31. https://doi.org/10.1016/j.fsigen.2019.03.017
Bille T, Weitz S, Buckleton JS, Bright JA (2019) Interpreting a major component from a mixed DNA profile with an unknown number of minor contributors. Forensic Sci Int Genet 40:150–159. https://doi.org/10.1016/j.fsigen.2019.02.017
Swaminathan H, Qureshi MO, Grgicak CM, Duffy K, Lun DS (2018) Four model variants within a continuous forensic DNA mixture interpretation framework: effects on evidential inference and reporting. PLoS ONE 13:1–23. https://doi.org/10.1371/journal.pone.0207599
Kalafut K, Schuerman C, Sutton J, Faris T, Armogida L, Bright JA et al (2018) Implementation and validation of an improved allele specific stutter filtering method for electropherogram interpretation. Forensic Sci Int Genet 35:50–56. https://doi.org/10.1016/j.fsigen.2018.03.016
Taylor D, Buckleton J, Bright JA (2016) Factors affecting peak height variability for short tandem repeat data. Forensic Sci Int Genet 21:126–133. https://doi.org/10.1016/j.fsigen.2015.12.009
Coble MD, Bright JA, Buckleton JS, Curran JM (2015) Uncertainty in the number of contributors in the proposed new CODIS set. Forensic Sci Int Genet 19:207–211. https://doi.org/10.1016/j.fsigen.2015.07.005
Tvedebrink T (2014) On the exact distribution of the numbers of alleles in DNA mixtures. Int J Legal Med 128:427–437. https://doi.org/10.1007/s00414-013-0951-3
Bright JA, Curran JM, Buckleton JS (2014) The effects of the uncertainly in the number of contributors to mixed DNA profiles on profile interpretation. Forensic Sci Int Genet 12:208–214. https://doi.org/10.1016/j.fsigen.2014.06.009
Bright JA, Taylor D, Curran JM, Buckleton JS (2013) Developing allelic and stutter peak height models for a continuous method of DNA interpretation. Forensic Sci Int Genet 7:296–304. https://doi.org/10.1016/j.fsigen.2012.11.013
Ballantyne J, Hanson EK, Perlin MW (2013) DNA mixture genotyping by probabilistic computer interpretation of binomially-sampled laser capture cell populations: combining quantitative data for greater identification information. Sci Justice 53:103–114. https://doi.org/10.1016/j.scijus.2012.04.004
Brookes C, Bright JA, Harbison S, Buckleton J (2012) Characterizing stutter in forensic STR multiplexes. Forensic Sci Int Genet 6:58–63. https://doi.org/10.1016/j.fsigen.2011.02.001
Paoletti DR, Doom TE, Krane CM, Raymer ML, Krane DE (2005) Empirical analysis of the STR profiles resulting from conceptual mixtures. J Forensic Sci 50:1361–1366
Coble MD, Buckleton J, Butler JM, Egeland T, Fimmers R, Gill P et al (2016) DNA commission of the International Society for Forensic Genetics: recommendations on the validation of software programs performing biostatistical calculations for forensic genetics applications. Forensic Sci Int Genet 25:191–197. https://doi.org/10.1016/j.fsigen.2016.09.002
Duke KR, Myers SP (2020) Systematic evaluation of STRmix™ performance on degraded DNA profile data. Forensic Sci Int Genet 44:102174. https://doi.org/10.1016/j.fsigen.2019.102174
Bright JA, Taylor D, Curran JM, Buckleton JS (2013) Degradation of forensic DNA profiles. Australian J Forensic Sci 45:445–449. https://doi.org/10.1080/00450618.2013.772235
Holdren JP, Lander ES (2016) Forensic science in criminal courts: ensuring scientific validity of feature-comparison methods. President’s Council of Advisors on Science and Technology. https://obamawhitehouse.archives.gov/sites/default/files/microsites/ostp/PCAST/pcast_forensic_science_report_final.pdf
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M et al (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55:611–622. https://doi.org/10.1373/clinchem.2008.112797
Courts C, Pfaffl MW, Sauer E, Parson W (2019) Pleading for adherence to the MIQE-guidelines when reporting quantitative PCR data in forensic genetic research. Forensic Sci Int Genet 42:e21–e24. https://doi.org/10.1016/j.fsigen.2019.06.021
Holland MM, Parson W (2011) GeneMarker® HID: a reliable software tool for the analysis of forensic STR data. J Forensic Sci 56:29–35. https://doi.org/10.1111/j.1556-4029.2010.01565.x
Hastings WK (1970) Monte Carlo sampling methods using Markov chains and their applications. Biometrika 57:97–109. https://doi.org/10.1093/biomet/57.1.97
Hill CR, Duewer DL, Kline MC, Coble MD, Butler JM (2013) U.S. population data for 29 autosomal STR loci. Forensic Sci Int Genet 7:e82–e83. https://doi.org/10.1016/j.fsigen.2012.12.004
Shapiro SS, Wilk MB (1965) An analysis of variance test for normality (complete samples). Biometrika 52:591–611. https://doi.org/10.2307/2333709
Kruskal WH, Wallis WA (1952) Use of ranks in one-criterion variance analysis. J Am Stat Assoc 47:583–621. https://doi.org/10.2307/2280779
Holm S (1979) A simple sequentially rejective multiple test procedure. Scandinavian J. Stat. 6:65–70. http://www.jstor.org/stable/4615733. Accessed 31 Dec 2021
RStudio Team (2015) RStudio: integrated development for R. RStudio Inc, Boston, MA
Taylor D, Bright JA, Buckleton J, Curran J (2014) An illustration of the effect of various sources of uncertainly on DNA likelihood ratio calculations. Forensic Sci Int Genet 11:56–63. https://doi.org/10.1016/j.fsigen.2014.02.003
Acknowledgements
The authors wish to thank John Fosnacht, Teresa Snyder-Leiby, Dan Erb, and Sarah Copeland from SoftGenetics LLC for their support with software development and generating reports of MaSTR™ analyses. The authors also wish to thank four anonymous donors for the samples associated with this study.
Funding
Financial support provided by the Eberly College of Science, Department of Biochemistry & Molecular Biology, Forensic Science Program at Penn State University.
Author information
Authors and Affiliations
Contributions
MMH: experimental design, data analysis, wrote the manuscript; TMT, AJB, and SAG-S: experimental design, laboratory and data analysis, review of the manuscript; JAM: statistical analysis, development of figures, review of the manuscript.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Samples were collected with consent according to an Institutional Biosafety Committee approved protocol #IBC-48221.
Consent for publication
Not applicable.
Conflict of interest
MMH serves as the Acting Laboratory Director of Mitotyping Technologies, a SoftGenetics company, LLC. MMH receives no monetary compensation for his role as Laboratory Director and has no financial interests in SoftGenetics or Mitotyping Technologies.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Holland, M.M., Tiedge, T.M., Bender, A.J. et al. MaSTR™: an effective probabilistic genotyping tool for interpretation of STR mixtures associated with differentially degraded DNA. Int J Legal Med 136, 433–446 (2022). https://doi.org/10.1007/s00414-021-02771-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-021-02771-0