Abstract
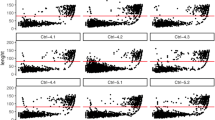
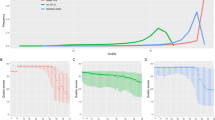
Mitochondrial DNA (mtDNA) is a small but significant part of the human genome, whose applicability potential has gradually increased with the advent of massively parallel sequencing (MPS) technology. Knowledge of the particular workflow, equipment, and reagents used, along with extensive usage of negative controls to monitor all preparation steps constitute the prerequisites for confident reporting of results. In this study, we performed an assessment of Illumina® Human mtDNA Genome assay on MiSeq FGx™ instrument. Through analysis of several types of negative controls, as well as mtDNA positive controls, we established thresholds for data analysis and interpretation, consisting of several components: minimum read depth (220 reads), minimum quality score (41), percentage of minor allele sufficient for analysis (3.0%), percentage of minor allele sufficient for interpretation (6.0%), and percentage of major allele sufficient for homoplasmic variant call (97.0%). Based on these criteria, we defined internal guidelines for analysis and interpretation of mtDNA results obtained by MPS. Our study shows that the whole mtDNA assay on MiSeq FGx™ produces repeatable and reproducible results, independent of the analyst, which are also concordant with Sanger-type sequencing results for mtDNA control region, as well as with MPS results produced by NextSeq®. Overall, established thresholds and interpretation guidelines were successfully applied for the sequencing of complete mitochondrial genomes from high-quality samples. The underlying principles and proposed methodology on the definition of internal laboratory guidelines for analysis and interpretation of MPS results may be applicable to similar MPS workflows, e.g. targeting good-quality samples in forensic genetics and molecular diagnostics.



Similar content being viewed by others
Data availability
The datasets generated and analysed during this study are available from the corresponding author on reasonable request.
References
Butler JM (2012) Mitochondrial DNA analysis. In: Butler JM (ed) Advanced topics in forensic DNA typing: methodology. Academic Press, Elsevier, Cambridge, pp 405–456
Parson W, Huber G, Moreno L, Madel MB, Brandhagen MD, Nagl S, Xavier C, Eduardoff M, Callaghan TC, Irwin JA (2015) Massively parallel sequencing of complete mitochondrial genomes from hair shaft samples. Forensic Sci Int Genet 15:8–15
Lyons EA, Scheible MK, Sturk-Andreaggi K, Irwin JA, Just RS (2013) A high-throughput Sanger strategy for human mitochondrial genome sequencing. BMC Genomics 14:881
Just RS, Scheible MK, Fast SA, Sturk-Andreaggi K, Röck AW, Bush JM, Higginbotham JL, Peck MA, Ring JD, Huber GE, Xavier C, Strobl C, Lyons EA, Diegoli TM, Bodner M, Fendt L, Kralj P, Nagl S, Niederwieser D, Zimmermann B, Parson W, Irwin JA (2015) Full mtGenome reference data: development and characterization of 588 forensic-quality haplotypes representing three U.S. populations. Forensic Sci Int Genet 14:141–155
Børsting C, Morling N (2015) Next generation sequencing and its applications in forensic genetics. Forensic Sci Inte Genet 18:78–89
Clarke AC, Prost S, JAL S, WTJ W, Kaplan ME, Matisoo-Smith EA, The Genographic Consortium (2014) From cheek swabs to consensus sequences: an A to Z protocol for high-throughput DNA sequencing of complete human mitochondrial genomes. BMC Genomics 15:68
ENFSI DNA Working Group (2010) Recommended minimum criteria for the validation of various aspects of the DNA profiling process, Issue no 001
Scientific Working Group on DNA Analysis Methods – SWGDAM (2016) Validation guidelines for DNA analysis methods
Parson W, Gusmão L, Hares DR, Irwin JA, Mayr WR, Morling N, Pokorak E, Prinz M, Salas A, Schneider PM, Parsons TJ (2014) DNA Commission of the International Society for Forensic Genetics: revised and extended guidelines for mitochondrial DNA typing. Forensic Sci Int Genet 13:134–142
Scientific Working Group on DNA Analysis Methods – SWGDAM (2019) Interpretation guidelines for mitochondrial DNA analysis by forensic DNA testing laboratories
Parson W, Strobl C, Huber G, Zimmermann B, Gomes SM, Souto L, Fendt L, Delport R, Langit R, Wootton S, Lagacé R, Irwin J (2013) Evaluation of next generation mtGenome sequencing using the Ion Torrent Personal Genome Machine (PGM). Forensic Sci Int Genet 7:543–549
Strobl C, Eduardoff M, Bus MM, Allen M, Parson W (2018) Evaluation of the precision ID whole MtDNA genome panel for forensic analyses. Forensic Sci Int Genet 35:21–25
Woerner AE, Ambers A, Wendt FR, King JL, Moura-Neto RS, Silva R, Budowle B (2018) Evaluation of the precision ID mtDNA whole genome panel on two massively parallel sequencing systems. Forensic Sci Int Genet 36:213–224
King JL, LaRue BL, Novroski NM, Stoljarova M, Seo SB, Zeng X, Warshauer DH, Davis CP, Parson W, Sajantila A, Budowle B (2014) High-quality and high-throughput massively parallel sequencing of the human mitochondrial genome using the Illumina MiSeq. Forensic Sci Int Genetics 12:128–135
McElhoe JA, Holland MM, Makova KD, Su MS, Paul IM, Baker CH, Faith SA, Young B (2014) Development and assessment of an optimized next-generation DNA sequencing approach for the mitogenome using the Illumina MiSeq. Forensic Sci Int Genet 13:20–29
Peck MA, Brandhagen MD, Marshall C, Diegoli TM, Irwin JA, Sturk-Andreaggi K (2016) Concordance and reproducibility of a next generation mtGenome sequencing method for high-quality samples using the Illumina MiSeq. Forensic Sci Int: Genetics 24:103–111
Peck MA, Sturk-Andreaggi K, Thomas JT, Oliver RS, Barritt-Ross S, Marshall C (2018) Developmental validation of a Nextera XT mitogenome Illumina MiSeq sequencing method for high-quality samples. Forensic Sci Int Genet 34:25–36
Qiagen (2014) EZ1® DNA Investigator® Handbook
Qiagen (2014) QIAamp® DNA Micro Handbook
National Institute of Standards and Technology – NIST (2018) Certificate of Analysis: Standard Reference Material® 2392 Mitochondrial DNA Sequencing (Human).
National Institute of Standards and Technology – NIST (2018) Certificate of Analysis: Standard Reference Material® 2392-I Mitochondrial DNA Sequencing (Human HL-60 DNA)
Illumina (2016) Protocol: Human mtDNA Genome for the Illumina Sequencing Platform, Document #15037958 v01
TaKaRa Bio Inc. (2015) PrimerSTAR® GXL DNA Polymerase Product Manual, Cat. #R050A, v201509Da
Illumina (2016) MiSeq® System Denature and Dilute Libraries Guide, Document #15039740 v01
Illumina (2019) NextSeq® System Denature and Dilute Libraries Guide, Document #15048776 v10
Barbarić L, Lipovac K, Sukser V, Rožić S, Korolija M, Zimmermann B, Parson W (2020) Maternal perspective of Croatian genetic diversity. Forensic Sci Int Genet 44:102190
Illumina (2016) mtDNA Variant Processor v1.0 BaseSpace App Guide, Document #1000000007931 v00
Anderson S, Bankier AT, Barrell BG, de Bruijn MH, Coulson AR, Drouin J, Eperon IC, Nierlich DP, Roe BA, Sanger F, Schreier PH, Smith AJ, Staden R, Young IG (1981) Sequence and organization of the human mitochondrial genome. Nature 290(5806):457–465
Andrews RM, Kubacka I, Chinnery PF, Lightowlers RN, Turnbull DM, Howell N (1999) Reanalysis and revision of the Cambridge reference sequence for human mitochondrial DNA. Nat Genet 23(2):147
Gilder JR, Doom TE, Inman K, Krane DE (2007) Run-specific limits of detection and quantitation for STR-based DNA testing. J Forensic Sci 52(1):97–101
Riman S, Kiesler KM, Borsuk LA, Vallone PM (2017) Characterization of NIST human mitochondrial DNA SRM-2392 and SRM-2392-I standard reference materials by next generation sequencing. Forensic Sci Int Genet 29:181–192
Robinson JT, Thorvaldsdóttir H, Winckler W, Guttman M, Lander ES, Getz G, Mesirov JP (2011) Integrative genomics viewer. Nat Biotechnol 29(1):24–26
Robinson JT, Thorvaldsdóttir H, Wenger AM, Zehir A, Mesirov JP (2017) Variant review with the integrative genomics viewer (IGV). Cancer Res 77(21):31–34
Illumina (2015) MiSeq® System Specification Sheet
Illumina (2019) Cluster Optimization: Overview Guide, Document #1000000071511 v00
Hussing C, Kampmann ML, Smidt Mogensen H, Børsting C, Morling N (2018) Quantification of massively parallel sequencing libraries – a comparative study of eight methods. Sci Rep 8:1110
Loman NJ, Misra RV, Dallman TJ, Constantinidou C, Gharbia SE, Wain J, Pallen MJ (2012) Performance comparison of benchtop high-throughput sequencing platforms. Nat Biotechnol 30(5):434–439
Quail MA, Smith M, Coupland P, Otto TD, Harris SR, Connor TR, Bertoni A, Swerdlow HP, Gu Y (2012) A tale of three next generation sequencing platforms: comparison of Ion Torrent, Pacific Biosciences and Illumina MiSeq sequencers. BMC Genomics 13:341
Schirmer M, Ijaz UZ, D’Amore R, Hall N, Sloan WT, Quince C (2015) Insight into biases and sequencing errors for amplicon sequencing with the Illumina MiSeq platform. Nucleic Acids Res 43(6):e37
Ring JD, Sturk-Andreaggi K, Peck MA, Marshall C (2017) A performance evaluation of Nextera XT and KAPA HyperPlus for rapid Illumina library preparation of long-range mitogenome amplicons. Forensic Sci Int Genet 29:174–180
Brandhagen MD, Just RS, Irwin JA (2020) Validation of NGS for mitochondrial DNA casework at the FBI laboratory. Forensic Sci Int Genet 44:102151
Acknowledgements
The authors thank all participants in the study for their valuable contributions in the form of samples and detailed informed consents. The authors are also thankful to Oliver Vugrek, PhD, Head of Laboratory for Advanced Genomics, Division of Molecular Medicine at “Ruđer Bošković” Institute, and their laboratory staff for collaboration in concordance study. Theauthors are grateful to Sara Rožić and Ivana Račić, PhD, who made valuable contributions in the experimental part of this study.
Funding
This work was supported by the Ministry of the Interior of the Republic of Croatia.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
This study involved samples collected from human participants. All procedures performed in the study were in accordance with the institutional and national ethical standards.
Consent to participate
Informed consent was obtained from all individual participants included in this study.
Consent for publication
Not applicable.
Code availability
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(XLSX 1107 kb)
Rights and permissions
About this article
Cite this article
Sukser, V., Rokić, F., Barbarić, L. et al. Assessment of Illumina® Human mtDNA Genome assay: workflow evaluation with development of analysis and interpretation guidelines. Int J Legal Med 135, 1161–1178 (2021). https://doi.org/10.1007/s00414-021-02508-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-021-02508-z