Abstract
One of the subjects within the meiotic field that has been actively investigated in the recent years is the temporal and functional relationships between meiotic recombination, cohesin loading and synaptonemal complex (SC) assembly. Although the study of meiotic mutants has shed some light, many questions remain to be answered. Here, we have studied this topic in the orthopteran Paratettix meridionalis, a species with telocentric chromosomes, which shows two unusual cytological features: pairing and synapsis of homologues during prophase I are restricted to the non-centromeric distal regions and extremely distal chiasma localization in metaphase I bivalents. In order to determine whether there is a relationship between both phenomena, we have used: (1) a spreading technique for following the ultrastructure of SC assembly and (2) immunofluorescence for SMC3 and SMC1α cohesin subunits, which mark the development of the axial element (a SC component); the histone γ-H2AX, which mostly labels the sites of double-strand breaks; and the recombinase RAD51. Spermatocytes showed conspicuous polarization of both the maturation of cohesin axes and the initiation of meiotic recombination events. Consequently, it is proposed that maturation of cohesin axes, which begins in very distal regions, could drive the latter loading of recombinases to such regions. This restricted distribution of recombination events along homologues would finally be responsible for the incomplete pairing and synapsis observed in all autosomes of the complement and hence for chiasma localization.






Similar content being viewed by others
References
Albini S, Jones GH (1988) Synaptonemal complex spreading in Allium cepa and Allium fistulosum: II: pachytene observations: the SC karyotype and the correspondence of late recombination nodules and chiasmata. Genome 30:399–410
Ashley T, Plug AW, Xu J, Solari AJ, ReddyG Golub EI, Ward DC (1995) Dynamic changes in Rad51 distribution on chromatin during meiosis in male and female vertebrates. Chromosoma 104:19–28
Barlow AL, Benson FE, West SC, Hulten MA (1997) Distribution of the Rad51 recombinase in human and mouse spermatocytes. Embo J 16:5207–5215
Baudat F, de Massy B (2007) Regulating double-stranded DNA break repair towards crossover or non-crossover during mammalian meiosis. Chromosome Res 15:565–577
Baudat F, Manova K, Yuen JP, Jasin M, Keeney S (2000) Chromosome synapsis defects and sexually dimorphic meiotic progression in mice lacking Spo11. Mol Cell 6:989–998
Bishop DK (1994) RecA homologs Dmc1 and Rad51 interact to form multiple nuclear complexes prior to meiotic chromosome synapsis. Cell 79:1081–1092
Bishop DK, Zickler D (2004) Early decision; meiotic crossover interference prior to stable strand exchange and synapsis. Cell 117:9–15
Bishop DK, Park D, Xu L, Kleckner N (1992) DMC1: a meiosis-specific yeast homolog of E. coli recA required for recombination, synaptonemal complex formation, and cell cycle progression. Cell 69:439–456
Calvente A, Viera A, Page J, Parra MT, Gomez R, Suja JA, Rufas JS, Santos JL (2005) DNA double-strand breaks and homology search: inferences from a species with incomplete pairing and synapsis. J Cell Sci 118:2957–2963
Celeste A, Fernandez-Capetillo O, Kruhlak MJ, Pilch DR, Staudt DW, Lee A, Bonner RF, Bonner WM, Nussenzweig A (2003) Histone H2AX phosphorylation is dispensable for the initial recognition of DNA breaks. Nat Cell Biol 5:675–679
Chicheportiche A, Bernardino-Sgherri J, de Massy B, Dutrillaux B (2007) Characterization of Spo11-dependent and independent phospho-H2AX foci during meiotic prophase I in the male mouse. J Cell Sci 120:1733–1742
Darlington CD (1931) The Mechanism of Crossing-Over. Science 73:561–562
del Cerro AL, Santos JL (1995) Synapsis in grasshopper bivalents heterozygous for centric shifts. Genome 38:616–622
del Cerro AL, Jones GH, Santos JL (1997) Chiasma localization and incomplete synapsis in two species of Tetrigidae (Orthoptera). Chromosome Res 5:69–71
Fernandez-Capetillo O, Allis CD, Nussenzweig A (2004) Phosphorylation of histone H2B at DNA double-strand breaks. J Exp Med 199:1671–1677
Fontana PG, Vickery VR (1975) The B-chromosome system of Tettigidea lateralis (Say). II. New karyomorphs, patterns of pycnosity and giemsa-banding. Chromosoma 50:371–391
Fox DP (1973) The control of chiasma distribution in the locust, Schistocerca gregaria (Forskal). Chromosoma 43:289–328
Franklin AE, McElver J, Sunjevaric I, Rothstein R, Bowen B, Cande WZ (1999) Three-dimensional microscopy of the Rad51 recombination protein during meiotic prophase. Plant Cell 11:809–824
Grelon M, Vezon D, Gendrot G, Pelletier G (2001) AtSPO11-1 is necessary for efficient meiotic recombination in plants. Embo J 20:589–600
Henderson SA (1961) The chromosomes of the British Tetrigidae (Orthoptera). Chromosoma 12:553–572
Henderson KA, Keeney S (2004) Tying synaptonemal complex initiation to the formation and programmed repair of DNA double-strand breaks. Proc Natl Acad Sci USA 101:4519–4524
Henderson KA, Keeney S (2005) Synaptonemal complex formation: where does it start? Bioessays 27:995–998
Hirano T (2002) The ABCs of SMC proteins: two-armed ATPases for chromosome condensation, cohesion, and repair. Genes Dev 16:399–414
Janssens FA (1924) La chiasmatypie dans les insectes. La Cellule 34:135–359
Jones GH (1973) Light and electron microscope studies of chromosome pairing in relation to chiasma localisation in Stethophyma grossum (Orthoptera: Acrididae). Chromosoma 42:145–162
Jones GH (1987) Chiasmata. In: Moens PB (ed) Meiosis. Academic, London, pp 213–244
Jones GH, Wallace BMN (1980) Meiotic chromosome pairing in Stethophyma grossum spermatocytes studies by a surfacespreading and silver staining technique. Chromosoma 78:187–201
Keeney S, Giroux CN, Kleckner N (1997) Meiosis-specific DNA double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell 88:375–384
Kleckner N (1996) Meiosis: how could it work? Proc Natl Acad Sci USA 93:8167–8174
Kleckner N (2006) Chiasma formation: chromatin/axis interplay and the role(s) of the synaptonemal complex. Chromosoma 115:175–194
Li W, Chen C, Markmann-Mulisch U, Timofejeva L, Schmelzer E, Ma H, Reiss B (2004) The Arabidopsis AtRAD51 gene is dispensable for vegetative development but required for meiosis. Proc Natl Acad Sci USA 101:10596–10601
Lorenz A, Whitby MC (2006) Crossover promotion and prevention. Biochem Soc Trans 34:537–541
Madigan JP, Chotkowski HL, Glaser RL (2002) DNA double-strand break-induced phosphorylation of Drosophila histone variant H2Av helps prevent radiation-induced apoptosis. Nucleic Acids Res 30:3698–36705
Mahadevaiah SK, Turner JM, Baudat F, Rogakou EP, de Boer P, Blanco-Rodriguez J, Jasin M, Keeney S, Bonner WM, Burgoyne PS (2001) Recombinational DNA double-strand breaks in mice precede synapsis. Nat Genet 27:271–276
Mather K (1938) Crossing-over. Biol Rev 13:252–292
Mercier R, Vezon D, Bullier E, Motamayor JC, Sellier A, Lefevre F, Pelletier G, Horlow C (2001) SWITCH1 (SWI1): a novel protein required for the establishment of sister chromatid cohesion and for bivalent formation at meiosis. Genes Dev 15:1859–1871
Mercier R, Armstrong SJ, Horlow C, Jackson NP, Makaroff CA, Vezon D, Pelletier G, Jones GH, Franklin FC (2003) The meiotic protein SWI1 is required for axial element formation and recombination initiation in Arabidopsis. Development 130:3309–3318
Moens PB, Chen DJ, Shen Z, Kolas N, Tarsounas M, Heng HH, Spyropoulos B (1997) Rad51 immunocytology in rat and mouse spermatocytes and oocytes. Chromosoma 106:207–215
Moens PB, Kolas NK, Tarsounas M, Marcon E, Cohen PE, Spyropoulos B (2002) The time course and chromosomal localization of recombination-related proteins at meiosis in the mouse are compatible with models that can resolve the early DNA–DNA interactions without reciprocal recombination. J Cell Sci 115:1611–1622
Oakley HA, Jones GH (1982) Meiosis in Mesostoma ehrenbergii (Turbellaria, Rhabdocoela). I. Chromosome pairing, synaptonemal complexes and chiasma localisation in spermatogenesis. Chromosoma 85:311–322
Orellana J, Giráldez R (1981) Metaphase I bound arms and crossing-over frequency in rye. I. Open pollinated varieties. Chromosoma 84:439–449
Page J, Suja JA, Santos JL, Rufas JS (1998) Squash procedure for protein immunolocalization in meiotic cells. Chromosome Res 6:639–642
Page J, Berrios S, Rufas JS, Parra MT, Suja JA, Heyting C, Fernandez-Donoso R (2003) The pairing of X and Y chromosomes during meiotic prophase in the marsupial species Thylamys elegans is maintained by a dense plate developed from their axial elements. J Cell Sci 116:551–560
Paigen K, Szatkiewicz JP, Sawyer K, Leahy N, Parvanov ED, Ng SH, Graber JH, Broman KW, Petkov PM (2008) The recombinational anatomy of a mouse chromosome. PLoS Genet 4:e1000119
Parra MT, Page J, Yen TJ, He D, Valdeolmillos A, Rufas JS, Suja JA (2002) Expression and behaviour of CENP-E at kinetochores during mouse spermatogenesis. Chromosoma 111:53–61
Pawlowski WP, Golubovskaya IN, Cande WZ (2003) Altered nuclear distribution of recombination protein RAD51 in maize mutants suggests the involvement of RAD51 in meiotic homology recognition. Plant Cell 15:1807–1816
Peoples TL, Dean E, Gonzalez O, Lambourne L, Burgess SM (2002) Close, stable homolog juxtaposition during meiosis in budding yeast is dependent on meiotic recombination, occurs independently of synapsis, and is distinct from DSB-independent pairing contacts. Genes Dev 16:1682–1695
Peters AH, Plug AW, van Vugt MJ, de Boer P (1997) A drying-down technique for the spreading of mammalian meiocytes from the male and female germline. Chromosome Res 5:66–68
Peters AH, O'Carroll D, Scherthan H, Mechtler K, Sauer S, Schofer C, Weipoltshammer K, Pagani M, Lachner M, Kohlmaier A et al (2001) Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell 107:323–337
Redon C, Pilch D, Rogakou E, Sedelnikova O, Newrock K, Bonner W (2002) Histone H2A variants H2AX and H2AZ. Curr Opin Genet Dev 12:162–169
Rockmill B, Sym M, Scherthan H, Roeder GS (1995) Roles for two RecA homologs in promoting meiotic chromosome synapsis. Genes Dev 9:2684–2695
Roeder GS (1997) Meiotic chromosomes: it takes two to tango. Genes Dev 11:2600–2621
Romanienko PJ, Camerini-Otero RD (2000) The mouse Spo11 gene is required for meiotic chromosome synapsis. Mol Cell 6:975–987
Sanchez-Moran E, Santos JL, Jones GH, Franklin FC (2007) ASY1 mediates AtDMC1-dependent interhomolog recombination during meiosis in Arabidopsis. Genes Dev 21:2220–2233
Santos JL, Giráldez R (1978) The effect of C-heterochromatin in chiasma terminalization in Chorthippus biguttulus (Acrididae, Orthoptera). Chromosoma 70:59–66
Shinohara A, Ogawa H, Ogawa T (1992) Rad51 protein involved in repair and recombination in S. cerevisiae is a RecA-like protein. Cell 69:457–470
Storlazzi A, Tesse S, Ruprich-Robert G, Gargano S, Poggeler S, Kleckner N, Zickler D (2008) Coupling meiotic chromosome axis integrity to recombination. Genes Dev 22:796–809
Tesse S, Storlazzi A, Kleckner N, Gargano S, Zickler D (2003) Localization and roles of Ski8p protein in Sordaria meiosis and delineation of three mechanistically distinct steps of meiotic homolog juxtaposition. Proc Natl Acad Sci USA 100:12865–12870
Tsubouchi H, Roeder GS (2003) The importance of genetic recombination for fidelity of chromosome pairing in meiosis. Dev Cell 5:915–925
Valdeolmillos AM, Viera A, Page J, Prieto I, Santos JL, Parra MT, Heck MM, Martinez AC, Barbero JL, Suja JA, Rufas JS (2007) Sequential loading of cohesin subunits during the first meiotic prophase of grasshoppers. PLoS Genet 3:e28
van Heemst D, Heyting C (2000) Sister chromatid cohesion and recombination in meiosis. Chromosoma 109:10–26
Viera A, Parra MT, Page J, Santos JL, Rufas JS, Suja JA (2003) Dynamic relocation of telomere complexes in mouse meiotic chromosomes. Chromosome Res 11:797–807
Viera A, Calvente A, Page J, Parra MT, Gomez R, Suja JA, Rufas JS, Santos JL (2004a) X and B chromosomes display similar meiotic characteristics in male grasshoppers. Cytogenet Genome Res 106:302–308
Viera A, Santos JL, Page J, Parra MT, Calvente A, Cifuentes M, Gomez R, Lira R, Suja JA, Rufas JS (2004b) DNA double-strand breaks, recombination and synapsis: the timing of meiosis differs in grasshoppers and flies. EMBO Rep 5:385–391
Zickler D, Kleckner N (1999) Meiotic chromosomes: integrating structure and function. Annu Rev Genet 33:603–754
Acknowledgements
We would like to express our gratitude to Dr. Carlos García de la Verga for his valuable criticisms and corrections on the manuscript and to the Consejería de Medio Ambiente y Desarrollo Rural (Comunidad de Castilla-La Mancha; España) for producing the authorization for sampling of animals. Dr. Jorge Iñiguez and Dr. David Lluciá Pomares helped in the geographic location of populations, Dr. Raimundo Outerelo made the taxonomical identification of the specimens and Dr. José Luis Barbero generously gave the antibodies against the cohesins SMC3 and SMC1α. This work was supported by grants BFU2005-02431 and BFU2006–06655 from Ministerio de Educación y Ciencia, Spain.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Keeney
All authors contributed equally to this work.
Electronic supplementary materials
Below is the link to the electronic supplementary material.
Supplementary Figure 1
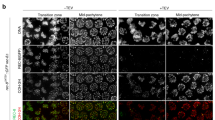
γ-H2AX staining in L. migratoria during prophase I. Double immunolabelling of SMC3 (red) and γ-H2AX (blue) onto spread (A–E and G) or squashed (F–F‴, H–H‴) spermatocytes of L. migratoria. When it is clearly discernible, the position of the X chromosome (X) is indicated (C–H‴). (A) Early leptotene: SMC3 axes are present in a dotted-like fashion, and there is no labelling of γ-H2AX. (B) Late leptotene: linear filaments of SMC3 are spread in the nucleus as well as a bunch of γ-H2AX labels. (C–E) Zygotene: SMC3 axes start progressive pairing and form thicker filaments. The massive γ-H2AX labelling observed in early zygotene (C) becomes restricted to the regions showing unpaired SMC3 axes as well as the single X chromosome in mid and late zygotene (D and E). (F–F‴) Late zygotene: γ-H2AX is only associated with the autosomal SMC3 axes which remain unpaired and to the single X chromosome axis. These regions are magnified in F‴. (G) Mid pachytene: γ-H2AX decorates the unpaired cohesin axis of the sex chromosome, and a reduced number of foci on the paired SMC axes of autosomes. (H-H‴) An identical pattern of γ-H2AX foci distribution on synapsed autosomes is observed in squashed preparations. However, the sex chromosome is not labelled, possibly as a result of steric interactions between the two primary antibodies. Bar represents 10 µm (GIF 178 kb)
Supplementary Figure 2
RAD51 loading in L. migratoria during prophase I. Double immunolabelling of SMC3 (red) and RAD51 (green) onto spread L. migratoria spermatocytes. The position of the single X chromosome is indicated (X) when it is discernible (C and D). (A) Late leptotene: RAD51 foci are observable on the trajectories of unpaired SMC3 axes. (B and C) Zygotene: the number of RAD51 foci is dramatically increased and are located on both paired and unpaired SMC3 axes. (D) Mid pachytene: a discrete number of RAD51 foci are associated to the paired cohesin axes. The presence of some RAD51 foci that do not locate on the cohesin axes from zygotene onwards is noteworthy. This situation is not observed in squashed spermatocytes, but it is a common feature of recombinases in spread preparations. Therefore, the previously reported dynamic pattern of RAD51 in grasshopper spermatocytes is also maintained with spreading techniques. Bar represents 10 µm (GIF 77 kb)
Rights and permissions
About this article
Cite this article
Viera, A., Santos, J.L. & Rufas, J.S. Relationship between incomplete synapsis and chiasma localization. Chromosoma 118, 377–389 (2009). https://doi.org/10.1007/s00412-009-0204-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-009-0204-x