Abstract
Objective
This work aimed to develop a radiomics nomogram to predict 3-year overall survival of esophageal cancer patients after chemoradiotherapy.
Methods
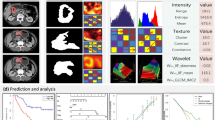
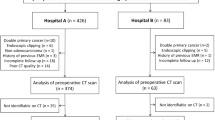
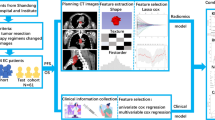
A total of 109 esophageal cancer patients, diagnosed from November 2012 to February 2015, were enrolled in this retrospective study. They were randomly divided into training set (77 cases) and verification set (32 cases). Image standardization was performed prior to feature extraction. And then, about 1670 radiomics features were extracted from the pretreatment diagnostic computed tomography image. A radiomics signature was constructed with the lasso algorithm; then, a radiomics score was calculated to reflect survival probability using the radiomics signature for each patient. A radiomics nomogram was developed by incorporating the radiomics score and clinical factors. A clinical model was constructed using clinical factors only. The performance of the nomogram was assessed with respect to its calibration and discrimination. Kaplan–Meier survival analysis was performed.
Results
Sixteen radiomics features were selected to build the radiomics signature. The radiomics nomogram showed better calibration and classification capacity than the clinical model with AUC 0.96 vs. 0.72 for the training cohort, and 0.87 vs. 0.67 for the validation cohort. The model showed good discrimination with a Harrell’s Concordance Index of 0.76 in the training cohort and 0.81 in the validation cohort. Decision curve analysis demonstrated the clinical usefulness of the radiomics nomogram. A significant difference (p value < 0.05; log-rank test) was observed between the survival curves of the nomogram-predicted survival and non-survival groups.
Conclusions
The present study proposed a radiomics-based nomogram involving the radiomics signature and clinical factors. It can be potentially applied in the individual preoperative prediction of 3-year survival in esophageal cancer patients.







Similar content being viewed by others
Data availability
The data that support the findings of this study are available from Shanxi Cancer Hospital. Restrictions apply to the availability of these data, which were used under license for this study.
References
Enzinger PC, Mayer RJ (2003) Esophageal cancer. N Engl J Med 349(23):2241–2252
Van Hagen P et al (2012) Preoperative chemoradiotherapy for esophageal or junctional cancer. N Engl J Med 366(22):2074–2084
Stewart BW, Kleihues P (eds) (2003) World cancer report. IARC Press, Lyon
Burmeister BH, Smithers BM, Gebski V, Fitzgerald L, Simes RJ, Devitt P et al (2005) Surgery alone versus chemoradiotherapy followed by surgery for resectable cancer of the oesophagus: a randomised controlled phase III trial. Lancet Oncol 6:659–668. https://doi.org/10.1016/S1470-2045(05)70288-6
Tanaka K et al (2016) Negative influence of programmed death-1-ligands on the survival of esophageal cancer patients treated with chemotherapy. Cancer Sci 107(6):726–733
Walsh TN, Noonan N, Hollywood D, Kelly A, Keeling N, Hennessy TPJ (1996) A comparison of multimodal therapy and surgery for esophageal adenocarcinoma. N Engl J Med 335:462–467. https://doi.org/10.1056/NEJM199608153350702
Huang FL, Yu SJ (2018) Esophageal cancer: risk factors, genetic association, and treatment. Asian J Surg 41(3):210–215. https://doi.org/10.1016/j.asjsur.2016.10.005
Goense L, Merrell KW, Arnett AL, Hallemeier CL, Meijer GJ, Ruurda JP, Hofstetter WL, van Hillegersberg R, Lin SH (2018) Validation of a nomogram predicting survival after trimodality therapy for esophageal cancer. Ann Thorac Surg 106(5):1541–1547. https://doi.org/10.1016/j.athoracsur.2018.05.055
Jin X, Zheng X, Chen D, Jin J, Zhu G, Deng X, Han C, Gong C, Zhou Y, Liu C, Xie C (2019) Prediction of response after chemoradiation for esophageal cancer using a combination of dosimetry and CT radiomics. Eur Radiol 29(11):6080–6088. https://doi.org/10.1007/s00330-019-06193-w
Larue RTHM, Klaassen R, Jochems A, Leijenaar RTH, Hulshof MCCM, van Berge Henegouwen MI, Schreurs WMJ, Sosef MN, van Elmpt W, van Laarhoven HWM, Lambin P (2018) Pre-treatment CT radiomics to predict 3-year overall survival following chemoradiotherapy of esophageal cancer. Acta Oncol 57(11):1475–1481. https://doi.org/10.1080/0284186X.2018.1486039 (Epub 2018 Aug 1)
Nga WTB, Eloumou SAFB, Engbang JPN, Bell EMD, Mayeh AMM, Atenguena E, Biwole ME, Ayissi GBN, Kenfack G, Noah DN, Luma HN, Sone AM, Ndom P, Ndam ECN (2019) Facteurs pronostiques du cancer de l’œsophage au Cameroun: étude multicentrique [Prognosis and survival of esophageal cancer in Cameroon: a prognostic study]. Pan Afr Med J 31(33):73. https://doi.org/10.11604/pamj.2019.33.73.16112
Wang F, Ning S, Yu B, Wang Y (2021) USP14: structure, function, and target inhibition. Front Pharmacol 12: https://doi.org/10.3389/fphar.2021.801328
Lv Z, Yu Z, Xie S, Alamri A (2022) Deep Learning-based Smart Predictive Evaluation for Interactive Multimedia-enabled Smart Healthcare. ACM Trans Multimedia Comput Commun Appl 18(15):1–20. https://doi.org/10.1145/3468506
Liu, S., Yang, B., Wang, Y., Tian, J., Yin, L., Zheng, W. (2022). 2D/3D Multimode Medical Image Registration Based on Normalized Cross-Correlation. Appl Sci, 12(6). doi: 10.3390/app12062828
Xu Q, Zeng Y, Tang W, Peng W, Xia T, Li Z, Guo J (2020) Multi-task joint learning model for segmenting and classifying tongue images using a deep neural network. IEEE J Biomed Health Inform 24(9):2481–2489. https://doi.org/10.1109/JBHI.2020.2986376
Katsila T, Liontos M, Patrinos GP, Bamias A, Kardamakis D (2018) The new age of-omics in urothelial cancer—re-wording its diagnosis and treatment. EBioMedicine 28:43–50
Lambin P, Rios-Velazquez E, Leijenaar R, Carvalho S, van Stiphout RG, Granton P et al (2012) Radiomics: extracting more information from medical images using advanced feature analysis. Eur J Cancer 48(4):441–446
O’Connor JPB, Aboagye EO, Adams JE et al (2017) Imaging biomarker roadmap for cancer studies. Nat Rev Clin Oncol 14:169–186
Aerts HJWL, Velazquez ER, Leijenaar RTH, Parmar C, Grossmann P, Cavalho S et al (2014) Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nat Commun 5:4006
Huang YQ, Liang CH, He L, Tian J, Liang CS, Chen X et al (2016) Development and validation of a radiomics nomogram for preoperative prediction of lymph node metastasis in colorectal cancer. Sci Found China 34(4):2157–2164
Lee G, Lee HY, Park H, Schiebler ML, van Beek EJ, Ohno Y et al (2017) Radiomics and its emerging role in lung cancer research, imaging biomarkers and clinical management: state of the art. Eur J Radiol 86:297–307
Wu S, Zheng J, Li Y, Yu H, Shi S, Xie W et al (2017) A radiomics nomogram for the preoperative prediction of lymph node metastasis in bladder cancer. Clin Cancer Res 23(22):6904–6911
Wu Y, Xu L, Yang P, Lin N, Huang X, Pan W, Li H, Lin P, Li B, Bunpetch V, Luo C, Jiang Y, Yang D, Huang M, Niu T, Ye Z (2018) Survival prediction in high-grade osteosarcoma using radiomics of diagnostic computed tomography. EBioMedicine 34:27–34. https://doi.org/10.1016/j.ebiom.2018.07.006 (Epub 2018 Jul 17)
Ganeshan B, Skogen K, Pressney I et al (2012) Tumour heterogeneity in oesophageal cancer assessed by CT texture analysis: preliminary evidence of an association with tumour metabolism, stage, and survival. Clin Radiol 67:157–164
Yip C, Davnall F, Kozarski R et al (2015) Assessment of changes in tumor heterogeneity following neoadjuvant chemotherapy in primary esophageal cancer. Dis Esophagus 28:172–179
Yip C, Landau D, Kozarski R et al (2014) Primary esophageal cancer: heterogeneity as potential prognostic biomarker in patients treated with definitive chemotherapy and radiation therapy. Radiology 270:141–148
Balagurunathan Y, Gu Y, Wang H et al (2014) Reproducibility and prognosis of quantitative features extracted from CT images. Transl Oncol 7(1):72–87
Shafiq-Ul-Hassan M, Zhang GG, Latifi K, Ullah G, Hunt DC, Balagurunathan Y, Abdalah MA, Schabath MB, Goldgof DG, Mackin D, Court LE, Gillies RJ, Moros EG (2017) Intrinsic dependencies of CT radiomic features on voxel size and number of gray levels. Med Phys 44(3):1050–1062. https://doi.org/10.1002/mp.12123
Zhao B, Tan Y, Tsai WY, Qi J, Xie C, Lu L, Schwartz LH (2016) Reproducibility of radiomics for deciphering tumor phenotype with imaging. Sci Rep 6:23428. https://doi.org/10.1038/srep23428
Lin L, Ehmke RC, Schwartz LH et al (2016) Assessing agreement between radiomic features computed for multiple CT imaging settings. PLoS ONE 11(12):e0166550
Encinas de la Iglesia J, Corral de la Calle MA, Fernández Pérez GC, Ruano Pérez R, Álvarez DA (2016) Esophageal cancer: anatomic particularities, staging, and imaging techniques. Radiologia 58(5):352–365. https://doi.org/10.1016/j.rx.2016.06.004 (English, Spanish. Epub 2016 Jul 25)
Larue RTHM, van Timmeren JE, de Jong EEC et al (2017) Influence of grey level discretization on radiomic feature stability for different CT scanners, tube currents and slice thicknesses: a comprehensive phantom study. Acta Oncol 56:1544–1553
Fedorov A, Beichel R, Kalpathy-Cramer J, Finet J, Fillion-Robin JC, Pujol S, Bauer C, Jennings D, Fennessy F, Sonka M, Buatti J, Aylward S, Miller JV, Pieper S, Kikinis R (2012) 3D slicer as an image computing platform for the Quantitative Imaging Network. Magn Reson Imaging 30(9):1323–1341. https://doi.org/10.1016/j.mri.2012.05.001 (Epub 2012 Jul 6)
Zwanenburg A, Vallières M, Abdalah MA et al (2020) The image biomarker standardization initiative: standardized quantitative radiomics for high-throughput image-based phenotyping. Radiology 295(2):328–338. https://doi.org/10.1148/radiol.2020191145 (Epub 2020 Mar 10)
Sun C, Wee WG (1983) Neighboring grey level dependence matrix for texture classification. Comput Graph Image Process 23:341–352
Haralick RM, Shanmugam K, Dinstein I (1973) Textural features for image classification. IEEE Trans Syst Man Cybern SMC-3:610–621
Galloway MM (1975) Texture analysis using gray level run lengths. Comput Graph Image Process 4:172–179
Thibault G, Fertil B, Navarro C (2009) Texture indexes and gray level size zone matrix application to cell nuclei classification. In: 10th international conference on pattern recognition and information processing, Minsk, Belarus, pp 140–145
Amadasun M, King R (1989) Textural features corresponding to textural properties. IEEE Trans Syst Man Cybern 19:1264–1274
Lambin P, Leijenaar RTH, Deist TM et al (2017) Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol 14:749–762
Tibshirani R (2011) Regression shrinkage and selection via the lasso: a retrospective. J Roy Stat Soc Ser B (Stat Methodol) 73(3):267–288
Bozdogan H (1987) Model selection and Akaike’s information criterion (AIC): the general theory and its analytical extensions. Psychometrika 52(3):345–370
Kutner MH, Nachtsheim C, Neter J (2004) Applied linear regression models. McGraw-Hill, Irwin
Kramer AA, Zimmerman JE (2007) Assessing the calibration of mortality benchmarks in critical care: the Hosmer–Lemeshow test revisited. Crit Care Med 35(9):2052–2056
Vickers AJ, Elkin EB (2006) decision curve analysis: a novel method for evaluating prediction models. Med Decis Mak 26(6):565–574
Yan J, Yao Y, Yan S, Gao R, Lu W, He W (2020) Chiral protein supraparticles for tumor suppression and synergistic immunotherapy: an enabling strategy for bioactive supramolecular chirality construction. Nano Lett 20(8):5844–5852. https://doi.org/10.1021/acs.nanolett.0c01757
Cao Z, Wang Y, Zheng W, Yin L, Tang Y, Miao W, Liu S, Yang B (2022) The algorithm of stereo vision and shape from shading based on endoscope imaging. Biomed Signal Process Control 76:103658. https://doi.org/10.1016/j.bspc.2022.103658
Zhang M, Chen Y, Lin J (2021) A privacy-preserving optimization of neighborhood-based recommendation for medical-aided diagnosis and treatment. IEEE Internet Things J 8(13):10830–10842. https://doi.org/10.1109/JIOT.2021.3051060
Si X, Gao L, Song Y, Khayatnezhad M, Minaeifar AA (2020) Understanding population differentiation using geographical, morphological and genetic characterization. Indian J Genet 80(4):459–467
Peng X, Khayatnezhad M, Ghezeljehmeidan L (2021) Rapd profiling in detecting genetic variation in stellaria l. (caryophyllaceae). Genetika-Belgrade 53(1):349–362
Ma S, Khayatnezhad M, Minaeifar A (2021) Genetic diversity and relationships among Hypericum L. species by ISSR Markers: A high value medicinal plant from Northern of Iran. Caryologia 74(1):97–107
Parmar C, Rios Velazquez E, Leijenaar R et al (2014) Robust radiomics feature quantification using semiautomatic volumetric segmentation. PLoS ONE 9:e102107
Author information
Authors and Affiliations
Contributions
JW and XY conceived and designed the work. JZ, HL and PQ wrote the manuscript. All the authors listed have approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest regarding the publication of this paper.
Research involving human participants and/or animals
No.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, J., Yu, X., Zeng, J. et al. Radiomics model for preoperative prediction of 3-year survival-based CT image biomarkers in esophageal cancer. Eur Arch Otorhinolaryngol 279, 5433–5443 (2022). https://doi.org/10.1007/s00405-022-07510-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00405-022-07510-8