Abstract
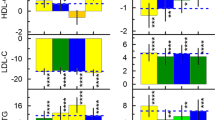
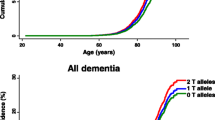
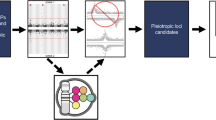
Cardiovascular (CV)- and lifestyle-associated risk factors (RFs) are increasingly recognized as important for Alzheimer’s disease (AD) pathogenesis. Beyond the ε4 allele of apolipoprotein E (APOE), comparatively little is known about whether CV-associated genes also increase risk for AD. Using large genome-wide association studies and validated tools to quantify genetic overlap, we systematically identified single nucleotide polymorphisms (SNPs) jointly associated with AD and one or more CV-associated RFs, namely body mass index (BMI), type 2 diabetes (T2D), coronary artery disease (CAD), waist hip ratio (WHR), total cholesterol (TC), triglycerides (TG), low-density (LDL) and high-density lipoprotein (HDL). In fold enrichment plots, we observed robust genetic enrichment in AD as a function of plasma lipids (TG, TC, LDL, and HDL); we found minimal AD genetic enrichment conditional on BMI, T2D, CAD, and WHR. Beyond APOE, at conjunction FDR < 0.05 we identified 90 SNPs on 19 different chromosomes that were jointly associated with AD and CV-associated outcomes. In meta-analyses across three independent cohorts, we found four novel loci within MBLAC1 (chromosome 7, meta-p = 1.44 × 10−9), MINK1 (chromosome 17, meta-p = 1.98 × 10−7) and two chromosome 11 SNPs within the MTCH2/SPI1 region (closest gene = DDB2, meta-p = 7.01 × 10−7 and closest gene = MYBPC3, meta-p = 5.62 × 10−8). In a large ‘AD-by-proxy’ cohort from the UK Biobank, we replicated three of the four novel AD/CV pleiotropic SNPs, namely variants within MINK1, MBLAC1, and DDB2. Expression of MBLAC1, SPI1, MINK1 and DDB2 was differentially altered within postmortem AD brains. Beyond APOE, we show that the polygenic component of AD is enriched for lipid-associated RFs. We pinpoint a subset of cardiovascular-associated genes that strongly increase the risk for AD. Our collective findings support a disease model in which cardiovascular biology is integral to the development of clinical AD in a subset of individuals.






Similar content being viewed by others
References
Allen M, Carrasquillo MM, Funk C et al (2016) Human whole genome genotype and transcriptome data for Alzheimer’s and other neurodegenerative diseases. Sci Data 3:160089. https://doi.org/10.1038/sdata.2016.89
Andreassen OA, Djurovic S, Thompson WK et al (2013) Improved detection of common variants associated with schizophrenia by leveraging pleiotropy with cardiovascular-disease risk factors. Am J Hum Genet 92:197–209. https://doi.org/10.1016/j.ajhg.2013.01.001
Andreassen OA, Thompson WK, Dale AM (2014) Boosting the power of schizophrenia genetics by leveraging new statistical tools. Schizophr Bull 40(1):13–17. https://doi.org/10.1093/schbul/sbt168
Andreassen OA, Thompson WK, Schork AJ et al (2013) Improved detection of common variants associated with schizophrenia and bipolar disorder using pleiotropy-informed conditional false discovery rate. PLoS Genet 9(4):e1003455. https://doi.org/10.1371/journal.pgen.1003455
Andreassen OA, Zuber V, Thompson WK, Schork AJ, Bettella F, Djurovic S et al (2014) Shared common variants in prostate cancer and blood lipids. Int J Epidemiol 43:1205–1214. https://doi.org/10.1093/ije/dyu090
Attems J, Jellinger KA (2014) The overlap between vascular disease and Alzheimer’s disease—lessons from pathology. BMC Med 12:206. https://doi.org/10.1186/s12916-014-0206-2
Barnes DE, Yaffe K (2011) The projected impact of risk factor reduction on Alzheimer’s disease prevalence. Lancet Neurol 10(9):819–828. https://doi.org/10.1016/S1474-4422(11)70072-2
Broce I, Karch CM, Wen N et al (2018) Immune-related genetic enrichment in frontotemporal dementia: an analysis of genome-wide association studies. PLoS Med 15(1):e1002487. https://doi.org/10.1371/journal.pmed.1002487
Carmona S, Hardy J, Guerreiro R (2018) The genetic landscape of Alzheimer disease, Chapter 26. In: Geschwind DH, Paulson HL, Klein C (eds) Handbook of clinical neurology, vol 148. Neurogenetics, Part II. Elsevier, Amsterdam, pp 395–408. https://doi.org/10.1016/b978-0-444-64076-5.00026-0
Dammer EB, Lee AK, Duong DM, Gearing M, Lah JJ, Levey AI et al (2014) Quantitative phosphoproteomics of Alzheimer’s disease reveals cross-talk between kinases and small heat shock proteins. J Proteom 15:508–519. https://doi.org/10.1002/pmic.201400189
Desikan RS, Fan CC, Wang Y et al (2017) Genetic assessment of age-associated Alzheimer disease risk: development and validation of a polygenic hazard score. PLoS Med 14(3):e1002258. https://doi.org/10.1371/journal.pmed.1002258
Desikan RS, Schork AJ, Wang Y et al (2015) Genetic overlap between Alzheimer’s disease and Parkinson’s disease at the MAPT locus. Mol Psychiatry 20(12):1588–1595. https://doi.org/10.1038/mp.2015.6
Desikan RS, Schork AJ, Wang Y et al (2015) Polygenic overlap between C-reactive protein, plasma lipids, and Alzheimer disease. Circulation 131(23):2061–2069. https://doi.org/10.1161/CIRCULATIONAHA.115.015489
Di Paolo G, Kim T-W (2011) Linking lipids to Alzheimer’s disease: cholesterol and beyond. Nat Rev Neurosci 12(5):284–296. https://doi.org/10.1038/nrn3012
Emdin CA, Khera AV, Natarajan P et al (2017) Genetic association of waist-to-hip ratio with cardiometabolic traits, type 2 diabetes, and coronary heart disease. JAMA 317(6):626–634. https://doi.org/10.1001/jama.2016.21042
Fagerberg L, Hallström BM, Oksvold P et al (2013) Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol Cell Proteom 13:397–406. https://doi.org/10.1074/mcp.m113.035600
Frikke-Schmidt R (2008) Association of loss-of-function mutations in the ABCA1 gene with high-density lipoprotein cholesterol levels and risk of ischemic heart disease. JAMA 299:2524. https://doi.org/10.1001/jama.299.21.2524
Pulit SL, Stoneman C, Morris AP (2018) Meta-analysis of genome-wide association studies for body fat distribution in 694,649 individuals of European ancestry. Biorxiv. https://doi.org/10.1101/304030
Hibar DP, Stein JL, Renteria ME et al (2015) Common genetic variants influence human subcortical brain structures. Nature 520(7546):224. https://doi.org/10.1038/nature14101
Huang K-L, Marcora E, Pimenova AA et al (2017) A common haplotype lowers PU.1 expression in myeloid cells and delays onset of Alzheimer’s disease. Nat Neurosci 20(8):1052–1061. https://doi.org/10.1038/nn.4587
Jun G, Ibrahim-Verbaas CA, Vronskaya M et al (2015) A novel Alzheimer disease locus located near the gene encoding tau protein. Mol Psychiatry 21:108–117. https://doi.org/10.1038/mp.2015.23
Karch CM, Ezerskiy LA, Bertelsen S, Consortium (ADGC) ADG, Goate AM (2016) Alzheimers disease risk polymorphisms regulate gene expression in the ZCWPW1 and the CELF1 loci. PLoS One 11(2):e0148717. https://doi.org/10.1371/journal.pone.0148717
Laird NM, Mosteller F (1990) Some statistical methods for combining experimental results. Int J Technol Assess Health Care 6:5–30. https://doi.org/10.1017/s0266462300008916
Lambert JC, Ibrahim-Verbaas CA, Harold D et al (2013) Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet 45(12):1452–1458. https://doi.org/10.1038/ng.2802
Larsson SC, Traylor M, Malik R et al (2017) Modifiable pathways in Alzheimer’s disease: Mendelian randomization analysis. BMJ 359:j5375
Livingston G, Sommerlad A, Orgeta V et al (2017) Dementia prevention, intervention, and care. Lancet 390(10113):2673–2734. https://doi.org/10.1016/S0140-6736(17)31363-6
Luchsinger JA (2001) Diabetes mellitus and risk of Alzheimers disease and dementia with stroke in a multiethnic cohort. Am J Epidemiol 154:635–641. https://doi.org/10.1093/aje/154.7.635
Mahajan A, Wessel J, Willems SM et al (2018) Refining the accuracy of validated target identification through coding variant fine-mapping in type 2 diabetes. Nat Genet 50(4):559–571. https://doi.org/10.1038/s41588-018-0084-1
Mahley RW (2016) Central nervous system lipoproteins: ApoE and regulation of cholesterol metabolism. Arterioscler Thromb Vasc Biol 36(7):1305–1315. https://doi.org/10.1161/ATVBAHA.116.307023
National Academies of Sciences, Engineering, and Medicine, Health and Medicine Division, Board on Health Sciences Policy, Committee on Preventing Dementia and Cognitive Impairment (2017) Preventing cognitive decline and dementia: a way forward. In: Downey A, Stroud C, Landis S, Leshner AI (eds) National Academies Press, Washington DC. http://www.ncbi.nlm.nih.gov/books/NBK436397/. Accessed 17 Apr 2018
Nelson CP, Goel A, Butterworth AS et al (2017) Association analyses based on false discovery rate implicate new loci for coronary artery disease. Nat Genet 49(9):1385–1391. https://doi.org/10.1038/ng.3913
Norton S, Matthews FE, Barnes DE, Yaffe K, Brayne C (2014) Potential for primary prevention of Alzheimer’s disease: an analysis of population-based data. Lancet Neurol 13(8):788–794. https://doi.org/10.1016/S1474-4422(14)70136-X
Østergaard SD, Mukherjee S, Sharp SJ et al (2015) Associations between potentially modifiable risk factors and Alzheimer disease: a Mendelian randomization study. PLoS Med 12(6):e1001841. https://doi.org/10.1371/journal.pmed.1001841
Profenno LA, Porsteinsson AP, Faraone SV (2010) Meta-analysis of Alzheimer’s disease risk with obesity, diabetes, and related disorders. Biol Psychiatry 67(6):505–512. https://doi.org/10.1016/j.biopsych.2009.02.013
Que X, Hung M-Y, Yeang C et al (2018) Oxidized phospholipids are proinflammatory and proatherogenic in hypercholesterolaemic mice. Nature 558(7709):301–306. https://doi.org/10.1038/s41586-018-0198-8
Ramasamy A, Trabzuni D, Guelfi S et al (2014) Genetic variability in the regulation of gene expression in ten regions of the human brain. Nat Neurosci 17(10):1418–1428. https://doi.org/10.1038/nn.3801
Reitz C (2013) Dyslipidemia and the risk of Alzheimer’s disease. Curr Atheroscler Rep 15(3):307. https://doi.org/10.1007/s11883-012-0307-3
Rietveld CA, Esko T, Davies G (2014) Common genetic variants associated with cognitive performance identified using the proxy-phenotype method. PNAS 111:13790–13794. https://doi.org/10.1073/pnas.1404623111
Sims R, van der Lee SJ, Naj AC et al (2017) Rare coding variants in PLCG2, ABI3, and TREM2 implicate microglial-mediated innate immunity in Alzheimer’s disease. Nat Genet 49(9):1373–1384. https://doi.org/10.1038/ng.3916
Sparks DL (2007) Cholesterol metabolism and brain amyloidosis: evidence for a role of copper in the clearance of Abeta through the liver. Curr Alzheimer Res 4(2):165–169
Staffaroni AM, Elahi FM, McDermott D et al (2017) Neuroimaging in dementia. Semin Neurol 37(5):510–537. https://doi.org/10.1055/s-0037-1608808
Stearns FW (2010) One hundred years of pleiotropy: a retrospective. J Genet 186(3):767–773. https://doi.org/10.1534/genetics.110.122549
Steele NZR, Carr JS, Bonham LW et al (2017) Fine-mapping of the human leukocyte antigen locus as a risk factor for Alzheimer disease: a case–control study. PLoS Med 14(3):e1002272. https://doi.org/10.1371/journal.pmed.1002272
Surakka I, Horikoshi M, Mägi R (2015) The impact of low-frequency and rare variants on lipid levels. Nat Genet 47:589–597. https://doi.org/10.1038/ng.3300
Westra H-J, Peters MJ, Esko T et al (2013) Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat Genet 45(10):1238–1243. https://doi.org/10.1038/ng.2756
Willer CJ, Schmidt EM, Sengupta S et al (2013) Discovery and refinement of loci associated with lipid levels. Nat Genet 45(11):1274–1283. https://doi.org/10.1038/ng.2797
Yengo L, Sidorenko J, Kemper KE et al (2018) Meta-analysis of genome-wide association studies for height and body mass index in ~700000 individuals of European ancestry. Hum Mol Genet. https://doi.org/10.1093/hmg/ddy271
Yokoyama JS, Wang Y, Schork AJ et al (2016) Association between genetic traits for immune-mediated diseases and Alzheimer disease. JAMA Neurol 73(6):691–697. https://doi.org/10.1001/jamaneurol.2016.0150
Yu C-E, Seltman H, Peskind ER (2007) Comprehensive analysis of APOE and selected proximate markers for late-onset Alzheimers disease: patterns of linkage disequilibrium and disease/marker association. J Genom 89:655–665. https://doi.org/10.1016/j.ygeno.2007.02.002
Zhang J, Liu Q (2015) Cholesterol metabolism and homeostasis in the brain. Protein Cell 6:254–264. https://doi.org/10.1007/s13238-014-0131-3
Xue-Shan Z, Juan P, Qi W, Zhong R, Li-Hong P, Zhi-Han T, Zhi-Sheng J, Gui-Xue W, Lu-Shan L (2016) Imbalanced cholesterol metabolism in Alzheimers disease. Clin Chim Acta 456:107–114. https://doi.org/10.1016/j.cca.2016.02.024
Jansen I, Savage J, Watanabe K, Bryois J, Williams D, Steinberg S, Sealock J, Karlsson I, Hagg S, Athanasiu LS (2018) Genetic meta-analysis identifies 9 novel loci and functional pathways for Alzheimers disease risk. Biorxiv. https://doi.org/10.1101/258533
Karlsson IK, Ploner A, Song C, Gatz M, Pedersen NL, Hägg S (2017) Genetic susceptibility to cardiovascular disease and risk of dementia. Transl Psychiatry. https://doi.org/10.1038/tp.2017.110
Kuźma E, Hannon E, Zhou A, Lourida I, Bethel A, Levine DA, Lunnon K, Thompson-Coon J, Hyppönen E, Llewellyn DJ (2018) Which risk factors causally influence dementia? A systematic review of Mendelian randomization studies. J Alzheimers Dis 64:181–193. https://doi.org/10.3233/jad-180013
Bulik-Sullivan B, Finucane HK, Anttila V, Gusev A, Day FR, Loh P-R, Duncan L, Perry JRB, Patterson N, Robinson EB, Daly MJ, Price AL, Neale BM (2015) An atlas of genetic correlations across human diseases and traits. Nat Genet 47:1236–1241. https://doi.org/10.1038/ng.3406
Grace C, Clarke R, Goel A, Farrall M, Watkins H, Hopewell JC (2018) Lack of genetic support for shared aetiology of coronary artery disease and late-onset Alzheimer’s disease. Sci Rep 1:1. https://doi.org/10.1038/s41598-018-25460-2
Mukherjee S, Walter S, Kauwe JS, Saykin AJ, Bennett DA, Larson EB, Crane PK, Glymour MM (2015) Genetically predicted body mass index and Alzheimers disease–related phenotypes in three large samples: Mendelian randomization analyses. Alzheimers Dementia 11:1439–1451. https://doi.org/10.1016/j.jalz.2015.05.015
Acknowledgements
We thank the Shiley-Marcos Alzheimer’s Disease Research Center at UCSD and the Memory and Aging Center at UCSF for continued support and the International Genomics of Alzheimer’s Project (IGAP) for providing summary result data for these analyses. This work was supported by Grants from the National Institutes of Health (NIH-AG046374, K01AG049152), Alzheimer’s Disease Genetics Consortium (U01 AG032984), National Alzheimer’s Coordinating Center Junior Investigator Award (RSD), RSNA Resident/Fellow Award (RSD), Foundation ASNR Alzheimer’s Imaging Grant (RSD), the Research Council of Norway (#213837, #225989, #223273, #237250/EU JPND), the South East Norway Health Authority (2013-123), Norwegian Health Association and the KG Jebsen Foundation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
JBB served on advisory boards for Elan, Bristol-Myers Squibb, Avanir, Novartis, Genentech, and Eli Lilly and holds stock options in CorTechs Labs, Inc. and Human Longevity, Inc. AMD is a founder of and holds equity in CorTechs Labs, Inc., and serves on its Scientific Advisory Board. He is also a member of the Scientific Advisory Board of Human Longevity, Inc. (HLI), and receives research funding from General Electric Healthcare (GEHC). The terms of these arrangements have been reviewed and approved by the University of California, San Diego, in accordance with its conflict of interest policies.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Broce, I.J., Tan, C.H., Fan, C.C. et al. Dissecting the genetic relationship between cardiovascular risk factors and Alzheimer’s disease. Acta Neuropathol 137, 209–226 (2019). https://doi.org/10.1007/s00401-018-1928-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-018-1928-6