Abstract
Purpose
Hair in the pilonidal sinus is not growing within the sinus cavity, as hair follicles are not present there. Not few pilonidal patients do not have intergluteal hair, which is said to be the causative agent of folliculitis and pilonidal genesis. So, what is the real source of the hair forming the typical pilonidal hair nest?
Methods
A trifold approach was used: First, axial hair strength testing of pilonidal hair and body hair harvested from head, lower back (glabella sacralis), and cranial third of intergluteal fold. Hair strength match was compared clinically. Second, comparative morphological examination by expert forensic biologist of hair from sinus and dorsal body hair. Third, statistical Bayesian classification of every single sinus hair based on its strength was done to determine the most probable region of origin.
Results
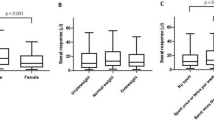
Using clinical hair strength comparison, in 13/20 patients, head hair is the stiffest hair, followed by intergluteal hair. Only in 6/20 patients, this is the case with hair from the glabella sacralis. According to comparative morphological comparison, a minimum of 5 of 13 hair nests with possible hair allocation examined contain hair from the occiput. In 5/18 nests, hair could not be determined to a specific location though. Statistical classification with correction for multiple testing shows that 2 nests have hair samples that are at least 100 times more probable to originate from head or lower back than from intergluteal fold.
Conclusion
We saw our null hypothesis that “hair in the sinus cavity is from the intergluteal region” rejected by each of three different approaches. There is strong evidence that occipital hair is present regularly in pilonidal sinus nests. We should start thinking of occipital hair as an important hair source for the development of the pilonidal hair nest.

Similar content being viewed by others
References
Akinci OF, Bozer M, Uzunköy A, Düzgün SA et al (1999) Incidence and aetiological factors in pilonidal sinus among Turkish soldiers. Eur J Surg 165:339–342
Allen-Mersh TG (1990) Pilonidal sinus: finding the right track for treatment. BrJSurg 77:123–132
Anderson AW (1847) Hair extracted from an ulcer. Boston Med Surg J 36:74
Ardelt M, Dennler U, Fahrner R, Hallof G, et al. (2017) Adolescence is a major factor of sinus pilonidal disease - gender specific based investigation of case development in Germany from 2007 until 2015. Chirurg 88(11):961–967
Bolandparvaz S, Moghadam Dizaj P, Salahi R, Paydar S et al (2012) Evaluation of the risk factors of pilonidal sinus: a single center experience. Turk J Gastroenterol 23:535–537
Bosche F, Luedi MM, van der Zypen D, Moersdorf P et al (2017) The hair in the sinus: sharp-ended rootless head hair fragments can be found in large amounts in pilonidal sinus nests. World J Surg 42(2):567–573
Buie LA (1944) Jeep disease (pilonidal disease of mechanized warfare). South Med J 37:103–109
Dahl HD, Henrich MH (1992) Light and scanning electron microscopy study of the pathogenesis of pilonidal sinus and anal fistula. Langenbecks Arch Chir 377:118–124
Davage ON (1954) The origin of sacrococcygeal pilonidal sinuses based on an analysis of four hundred sixty-three cases. Am J Pathol 30:1191–1205
Doll D, Bosche FD, Stauffer VK, Sinicina I, Hoffmann S, van der Zypen D, Luedi MM (2017) Strength of occipital hair as an explanation for pilonidal sinus disease caused by intruding hair. Dis Colon Rectum 60:979–986
Doll D, Luedi MM, Wieferich K, van der Zypen D et al (2015) Stop insulting the patient: neither incidence nor recurrence in pilonidal sinus disease is linked to personal hygiene. Pilonidal Sinus J 1:11–19
Doll D, Stauffer VK, Luedi MM (2016) Intra-anal pilonidal sinus disease: a unique diagnosis possibly pointing to the occiput. ANZ J Surg 86:622–625
Elliot D, Quyyumi S (1981) A “pennanidal” sinus. J R Soc Med 74:847–848
Evers T, Doll D, Matevossian E, Noe S, Neumann K, Li HL, Hüser N, Lüdde R, Hoffmann S, Krapohl BD (2011) Trends in incidence and long-term recurrence rate of pilonidal sinus disease and analysis of associated influencing factors. Zhonghua Wai Ke Za Zhi 49:799–803
Favre R, Delacroix P (1964) Apropos of 1,110 cases of pilonidal disease of coccy-perineal localization. Mem Acad Chir (Paris) 90:669–676
Gosselink MP, Jenkins L, Toh JWT (2017) Scanning electron microscope imaging of pilonidal disease. Tech Coloproctol 21(11):905–906
Karahan Ö, Eryilmaz MA, Torun V, Sevinç B et al (2010) Is the increase in the number of pilonidal sinus surgery normal? Turkish J Surg 26:207–211
Klass AA (1956) The so-called pilo-nidal sinus. Can Med Assoc J 75:737–742
Lord PH (1970) Unusual case of pilonidal sinus. Proc R Soc Med 63:967–968
Mohanna PN, Al-Sam SZ, Flemming AF (2001) Subungual pilonidal sinus of the hand in a dog groomer. Br J Plast Surg 54:176–178
Mount LA (1949) Congenital dermal sinuses as a cause of meningitis, intraspinal abscess and intracranial abscess. J Am Med Assoc 139:1263–1268
Oien CT (2009) Forensic hair comparison: background information for interpretation. FBI Forensic Case Rev 11
Patey D (1971) Pilonidal sinus: a postscript. Lancet 1:245
Sekmen Ü, Kara VM, Altintoprak F, Şenol Z (2010) Pilonidal sinus in the army: its incidence and risk factors. Turkish J Surg 26:95–98
Senapati A (2012) The management of pilonidal sinus disease. Contemporary Coloproctology Springer-Verlag London Ltd: 67–75
Stone HB (1924) Pilonidal sinus: coccygeal fistula. Ann Surg 79:410–414
Tait GB, Wilks JM, Sames CP (1948) Pilonidal sinus in a barber’s hand. Lancet 2:121
Vaiude P, Dhital M, Hancock K (2011) A true pilonidal sinus in the hand of a sheep shearer. J Surg Case Rep 2011:6
Warren JM (1854) Abscess, containing hair, on the nates. Am J Sci 28:113
Yildiz T, Elmas B, Yucak A, Turgut HT, Ilce Z (2017) Risk factors for pilonidal sinus disease in teenagers. Indian J Pediatr 84:134–138
Zimmerman K (1970) Pilonidal disease. Dis Colon Rectum 13:330–332
Author information
Authors and Affiliations
Contributions
Statistical analysis and calculations: AH, DD
Manuscript editing and interpretation of data: DD, FB, MML, FR, IS, AH, PM
Manuscript writing: DD, AH, DD, JG, FR, FB, MML
Graphic design: DD, AH, FB
Data acquisition: FB, AH, DD, JG, FR
Corresponding author
Ethics declarations
The ethics committee of the medical association of Niedersachsen, Berliner Allee 20, 30175 Hannover, Germany (Prof. Dr. med. Andreas Creutzig, chair), fully and unanimously approved the study based on § 15 of the Niedersachsen Medical Association’s professional code of conduct.
Conflict of interest
The authors declare that there is no conflict of interest.
Appendix. Statistical remarks
Appendix. Statistical remarks
Aggregating hair strength measurements
The strength of every hair was measured up to six times in sequence; we used a linear model with the hair as a factor variable and the measurement number as a numerical variable to aggregate multiple measurements into a single strength value for each hair (see the section “Statistical analysis”). The most important assumption of this model, the homoscedasticity assumption, was fulfilled by the data (see Fig. 2, left), but the distribution of the residuals was rather long-tailed and slightly skewed than normal (see Fig. 2, right). Despite this fact, we considered the overall model fit good enough to be used for aggregating multiple strength measurements for hair samples.
Classification of pilonidal sinus hair to regions of origins
We performed linear discriminant analysis (LDA) to statistically attribute pilonidal sinus hair of each patient to their most probable region of origin (protuberantia occipitalis externa [POE], glabella sacralis [GS], or intergluteal fold [IGF], respectively). This proceeded as follows:
-
1.
We fitted one LDA model per patient, using hair strengths from POE, GS, and IGF samples to train the model. The LDA model assumes hair strengths of each region of origin are normally distributed with equal variance, but different means.
-
2.
We classified each pilonidal sinus hair of a patient by calculating its posterior probabilities to originate from POE, GS, and IGF, respectively. This calculation was done based on the normal distributions of hair strength fitted in step 1, and based on the prior probabilities, we assigned to each region of origin; we assigned equal prior probability to each region of origin (POE, GS, and IGF; “objective” or “uninformed” prior.
-
3.
In order to quantify the uncertainty in the calculated posteriors, we performed non-parametric bootstrap. That means, we resampled hair samples for each patient 999 times (random sampling with replacement), and repeated steps 1 and 2 above for every simulated sample. We used the empirical distribution of the 999 bootstrap posteriors calculated like this to derive 95% confidence intervals for the estimated posteriors.
This analysis has several potentially delicate points:
-
Assumption of a model (normally distributed hair strength with equal variance in each region of origin) that may or may not be appropriate. By visual inspection of the distribution of hair strengths in different regions (POE, GS, and IGF), we convinced ourselves that the assumption of this model seems safe.
-
Selection bias. In order to reliably fit an LDA model, we would need (random) hair samples from the three regions (POE, GS, and IGF). However, it is not possible to ensure random sampling in practice: hair are chosen and harvested by human researchers. The sampling strategy that was applied for most patients actually contradicted this requirement: in the beginning of the study, hair were sampled based on visual strength, having forensic examination and visual inspection in mind. To estimate the validity of models trained on these non-random hair samples, new samples were collected from three patients (#1, #4, #15), this time making sure selection is as random as possible for a human researcher. We compared these randomly selected hair samples to the older samples selected based on visible strength.
Interestingly, the hair samples collected with the aim of selecting strong hairs from the different regions were on average not stronger than the random samples (Fig. 3). When comparing hair strength from random and non-random sampling with T tests, we got unadjusted p values of 0.018 (patient #1, IGF; mean hair force from randomly sampled hair is larger), 0.045 (patient #15, GS; mean hair force from randomly sampled hair is larger), and above 0.05 in other cases. After adjusting the p values for multiple testing with the Holm correction, the minimal (adjusted) p value was 0.160. We concluded that the problem of non-random selection of hair samples can be neglected in practice. It seems that it is difficult to determine the hair strength by eye, and that the samples of supposedly strong hair are indeed representative for the whole range of hair strengths of the patients, which is good for fitting the LDA models.
-
Small sample size. Most models are based on only 6 hair from each region (POE, GS, and IGF); the calculated posteriors are therefore not very precise. We performed the non-parametric bootstrap (see above) in order to quantify the uncertainty. As expected, confidence intervals are large for most hair samples, and most pilonidal sinus hair cannot be statistically attributed to a single region of origin with high probability (see also detailed results in Fig. 3).
Classification results for all pilonidal sinus hair samples are depicted in Fig. 4.
Posterior probabilities of all classified hair to originate from POE, GS, or IGF, respectively. Numbers in brackets denote the 95% confidence intervals calculated by bootstrap. Red cells refer to a high probability and green cells to a low probability. Patients not shown here did not have a sinus hair sample (patients #7, #12), or no IGF hair sample (patients #2, #14; the null hypothesis that sinus hair stems from IGF cannot be tested in this case). Reading example for patient #1: hair no. 1 is assigned a probability between 43 and 90% of originating from POE, 2 to 27% of originating from GS, and 4 to 36% of originating from IGF
Rights and permissions
About this article
Cite this article
Doll, D., Bosche, F., Hauser, A. et al. The presence of occipital hair in the pilonidal sinus cavity—a triple approach to proof. Int J Colorectal Dis 33, 567–576 (2018). https://doi.org/10.1007/s00384-018-2988-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00384-018-2988-8