Abstract
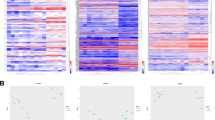
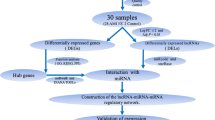
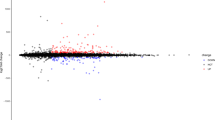
Myocardial infarction (MI) is the leading cause of fatality worldwide. Our study aimed to investigate the dysregulated long non-coding RNA (lncRNA) in MI and elucidate the mechanism of it in MI. The lncRNA and mRNA expression profiling of the whole left ventricular tissue of MI mice model (8 mice) and Sham group (8 mice) was obtained based on microarray analysis. Differentially expressed lnRNAs/mRNA (DELs/DEMs) were identified in MI. DELs/DEMs co-expression network construction, gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis were conducted to predict the biological functions of DEMs. Quantitative real-time polymerase chain reaction (qRT-PCR) was subjected to validate the abnormally expressed DELs in left ventricular tissues of MI mice model. Total of 168 DELs (37 up- and 131 down-regulated) and 126 DEMs (87 up- and 39 down-regulated) were identified in MI compared with Sham group. The co-expression network of candidate DELs and DEMs was constructed, which covered 219 nodes and 1775 edges. The qRT-PCR validation results indicated that ENSMUST00000124047 was significantly down-regulated in MI group and AK166279 was significantly up-regulated in MI group. ENSMUST00000121611 and NR_015515 had the up-regulated tendency in MI group compared with Sham group. The DEMs in MI were significantly enriched in 41 signaling pathways including complement and coagulation cascades, cytokine–cytokine receptor interaction and chemokine signaling pathway. The expression profiling of dysregulated DELs in MI was identified. Our results might provide useful information for exploring the pathogenesis mechanism of MI.




Similar content being viewed by others
References
Lopez AD, Mathers CD, Ezzati M, Jamison DT, Murray CJ (2006) Global and regional burden of disease and risk factors, 2001: systematic analysis of population health data. Lancet 367(9524):1747–1757
Davies MJ, Thomas AC (1985) Plaque fissuring—the cause of acute myocardial infarction, sudden ischaemic death, and crescendo angina. Br Heart J 53(4):363–373
Wolf D, Stachon P, Bode C, Zirlik A (2014) Inflammatory mechanisms in atherosclerosis. Hamostaseologie 34(1):63–71
White HD, Chew DP (2008) Acute myocardial infarction. Lancet 372(9638):570–584
Inami T, Okabe M, Matsushita M, Kobayashi N, Inokuchi K, Hata N, Seino Y, Shimizu W (2016) JAK2 mutation and acute coronary syndrome complicated with stent thrombosis. Heart Vessels 31(10):1714–1716
Szpakowicz A, Pepinski W, Waszkiewicz E, Maciorkowska D, Skawronska M, Niemcunowicz-Janica A, Dobrzycki S, Musial WJ, Kaminski KA (2016) The influence of renal function on the association of rs854560 polymorphism of paraoxonase 1 gene with long-term prognosis in patients after myocardial infarction. Heart Vessels 31(1):15–22
Kiliszek M, Szpakowicz A, Franaszczyk M, Pepinski W, Waszkiewicz E, Skawronska M, Ploski R, Niemcunowicz-Janica A, Budnik M, Poludniewska D, Musial WJ, Kaminski KA, Opolski G (2016) The 9p21 polymorphism is linked with atrial fibrillation during acute phase of ST-segment elevation myocardial infarction. Heart Vessels 31(10):1590–1594
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136(2):215–233
Huang W, Tian SS, Hang PZ, Sun C, Guo J, Du ZM (2016) Combination of microRNA-21 and microRNA-146a attenuates cardiac dysfunction and apoptosis during acute myocardial infarction in mice. Mol Ther Nucleic Acids 5:e296
Wang Y, Pan X, Fan Y, Hu X, Liu X, Xiang M, Wang J (2015) Dysregulated expression of microRNAs and mRNAs in myocardial infarction. Am J Transl Res 7(11):2291–2304
Tony H, Meng K, Wu B, Yu A, Zeng Q, Yu K, Zhong Y (2015) MicroRNA-208a dysregulates apoptosis genes expression and promotes cardiomyocyte apoptosis during ischemia and its silencing improves cardiac function after myocardial infarction. Mediat Inflamm 2015:479123
Dey BK, Mueller AC, Dutta A (2014) Long non-coding RNAs as emerging regulators of differentiation, development, and disease. Transcription 5(4):e944014
Wang K, Liu F, Liu CY, An T, Zhang J, Zhou LY, Wang M, Dong YH, Li N, Gao JN, Zhao YF, Li PF (2016) The long noncoding RNA NRF regulates programmed necrosis and myocardial injury during ischemia and reperfusion by targeting miR-873. Cell Death Differ 23(8):1394–1405
Wang K, Liu CY, Zhou LY, Wang JX, Wang M, Zhao B, Zhao WK, Xu SJ, Fan LH, Zhang XJ, Feng C, Wang CQ, Zhao YF, Li PF (2015) APF lncRNA regulates autophagy and myocardial infarction by targeting miR-188-3p. Nat Commun 6:6779
Liao J, He Q, Li M, Chen Y, Liu Y, Wang J (2016) LncRNA MIAT: myocardial infarction associated and more. Gene 578(2):158–161
Ishii N, Ozaki K, Sato H, Mizuno H, Saito S, Takahashi A, Miyamoto Y, Ikegawa S, Kamatani N, Hori M, Saito S, Nakamura Y, Tanaka T (2006) Identification of a novel non-coding RNA, MIAT, that confers risk of myocardial infarction. J Hum Genet 51(12):1087–1099
Qu X, Du Y, Shu Y, Gao M, Sun F, Luo S, Yang T, Zhan L, Yuan Y, Chu W, Pan Z, Wang Z, Yang B, Lu Y (2017) MIAT is a pro-fibrotic long non-coding rna governing cardiac fibrosis in post-infarct myocardium. Sci Rep 7:42657
Frade AF, Laugier L, Ferreira LR, Baron MA, Benvenuti LA, Teixeira PC, Navarro IC, Cabantous S, Ferreira FM, da Silva Candido D, Gaiotto FA, Bacal F, Pomerantzeff P, Santos RH, Kalil J, Cunha-Neto E, Chevillard C (2016) Myocardial infarction-associated transcript, a long noncoding RNA, is overexpressed during dilated cardiomyopathy due to chronic chagas disease. J Infect Dis 214(1):161–165
Zangrando J, Zhang L, Vausort M, Maskali F, Marie PY, Wagner DR, Devaux Y (2014) Identification of candidate long non-coding RNAs in response to myocardial infarction. BMC Genom 15:460
Li L, Weng Z, Yao C, Song Y, Ma T (2015) Aquaporin-1 deficiency protects against myocardial infarction by reducing both edema and apoptosis in mice. Sci Rep 5:13807
Smyth GK (2005) Limma: linear models for microarray data. Bioinformatics and computational biology solutions using R and bioconductor. Springer, New York, pp 397–420
Benesty J, Chen J, Huang Y, Cohen I (2009) Pearson correlation coefficient. Noise reduction in speech processing. Springer, New York, pp 1–4
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative CT method. Nat Protoc 3(6):1101–1108
Takahashi N, Goto T, Kusudo T, Moriyama T, Kawada T (2005) The structures and functions of peroxisome proliferator-activated receptors (PPARs). Nihon rinsho 63(4):557–564
Desvergne B, Wahli W (1999) Peroxisome proliferator-activated receptors: nuclear control of metabolism. Endocr Rev 20(5):649–688
Chen MC, Chang JP, Lin YS, Pan KL, Ho WC, Liu WH, Chang TH, Huang YK, Fang CY, Chen CJ (2016) Deciphering the gene expression profile of peroxisome proliferator-activated receptor signaling pathway in the left atria of patients with mitral regurgitation. J Transl Med 14(1):157
Colak D, Alaiya AA, Kaya N, Muiya NP, AlHarazi O, Shinwari Z, Andres E, Dzimiri N (2016) Integrated left ventricular global transcriptome and proteome profiling in human end-stage dilated cardiomyopathy. PLoS One 11(10):e0162669
Liu C, Guo Q, Lu M, Li Y (2015) An experimental study on amelioration of dyslipidemia-induced atherosclesis by clematichinenoside through regulating peroxisome proliferator-activated receptor-alpha mediated apolipoprotein A-I, A-II and C-III. Eur J Pharmacol 761:362–374
Peloso GM, Demissie S, Collins D, Mirel DB, Gabriel SB, Cupples LA, Robins SJ, Schaefer EJ, Brousseau ME (2010) Common genetic variation in multiple metabolic pathways influences susceptibility to low HDL-cholesterol and coronary heart disease. J Lipid Res 51(12):3524–3532
Song X, Tian SP, Ju HY, Zhang F, Li YN, Wu F, Yang L (2015) Predictive value of apolipoprotein for coronary atherosclerosis in asymptomatic non-diabetic population. Zhongguo yi xue ke xue yuan xue bao 37(1):55–60
Li H, Han Y, Qi R, Wang Y, Zhang X, Yu M, Tang Y, Wang M, Shu YN, Huang W, Liu X, Rodrigues B, Han M, Liu G (2015) Aggravated restenosis and atherogenesis in ApoCIII transgenic mice but lack of protection in ApoCIII knockouts: the effect of authentic triglyceride-rich lipoproteins with and without ApoCIII. Cardiovasc Res 107(4):579–589
Gerritsen G, Rensen PC, Kypreos KE, Zannis VI, Havekes LM, Willems van Dijk K (2005) ApoC-III deficiency prevents hyperlipidemia induced by apoE overexpression. J Lipid Res 46(7):1466–1473
Ali A, Gale RE, Shakoori AR (2016) Detection of FLT3/TKD and IDH1 mutations in Pakistani acute myeloid leukemia patients by denaturing HPLC. J Cell Biochem 118(5):1174–1181
Sironi S, Wagner M, Kuett A, Drolle H, Polzer H, Spiekermann K, Rieger C, Fiegl M (2015) Microenvironmental hypoxia regulates FLT3 expression and biology in AML. Sci Rep 5:17550
deLapeyriere O, Naquet P, Planche J, Marchetto S, Rottapel R, Gambarelli D, Rosnet O, Birnbaum D (1995) Expression of Flt3 tyrosine kinase receptor gene in mouse hematopoietic and nervous tissues. Differentiation 58(5):351–359
Pfister O, Lorenz V, Oikonomopoulos A, Xu L, Hauselmann SP, Mbah C, Kaufmann BA, Liao R, Wodnar-Filipowicz A, Kuster GM (2014) FLT3 activation improves post-myocardial infarction remodeling involving a cytoprotective effect on cardiomyocytes. J Am Coll Cardiol 63(10):1011–1019
Gu L, Liu W, Yan Y, Su L, Wu G, Liang B, Tan J, Huang G (2014) Influence of the beta-fibrinogen-455G/A polymorphism on development of ischemic stroke and coronary heart disease. Thromb Res 133(6):993–1005
Lovely RS, Yang Q, Massaro JM, Wang J, D’Agostino RB Sr, O’Donnell CJ, Shannon J, Farrell DH (2011) Assessment of genetic determinants of the association of gamma’ fibrinogen in relation to cardiovascular disease. Arterioscler Thromb Vasc Biol 31(10):2345–2352
Oikonomopoulou K, Ricklin D, Ward PA, Lambris JD (2012) Interactions between coagulation and complement—their role in inflammation. Semin immunopathol 34(1):151–165
Cao Y, Li R, Li Y, Zhang T, Wu N, Zhang J, Guo Z (2017) Identification of transcription factor-gene regulatory network in acute myocardial infarction. Heart Lung Circ 26(4):343–353
Rajasekaran NS, Firpo MA, Milash BA, Weiss RB, Benjamin IJ (2008) Global expression profiling identifies a novel biosignature for protein aggregation R120GCryAB cardiomyopathy in mice. Physiol Genom 35(2):165–172
Gigante B, Bellis A, Visconti R, Marino M, Morisco C, Trimarco V, Galasso G, Piscione F, De Luca N, Prince JA, de Faire U, Trimarco B (2007) Retrospective analysis of coagulation factor II receptor (F2R) sequence variation and coronary heart disease in hypertensive patients. Arterioscler Thromb Vasc Biol 27(5):1213–1219
Zhou H, Qiu Z, Gao S, Chen Q, Li S, Tan W, Liu X, Wang Z (2016) Integrated analysis of expression profile based on differentially expressed genes in middle cerebral artery occlusion animal models. Int J Mol Sci 17(5):E7776
Wang W, Liu H, Song M, Fang W, Yuan F (2016) Clinical effect of cardiac shock wave therapy on myocardial ischemia in patients with ischemic heart failure. J Cardiovasc Pharmacol Ther 21(4):381–387
Liu J, Wang H, Li J (2016) Inflammation and inflammatory cells in myocardial infarction and reperfusion injury: a double-edged sword. Clin Med Insights Cardiol 10:79–84
Swirski FK, Nahrendorf M (2013) Leukocyte behavior in atherosclerosis, myocardial infarction, and heart failure. Science 339(6116):161–166
Fan Y, Xiong X, Zhang Y, Yan D, Jian Z, Xu B, Zhao H (2016) MKEY, a peptide inhibitor of CXCL4-CCL5 heterodimer formation, protects against stroke in mice. J Am Heart Assoc 5(9):e003615
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
The work was supported by The National Natural Sciences Foundation of China (Project for Young Scientists, 30800999) and The National Natural Sciences Foundation of China (Project of International Cooperation and Exchanges, 812111369).
Conflict of interest
All of authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
380_2017_990_MOESM6_ESM.tif
Figure S1: GO terms and KEGG signaling pathways enrichment of DEMs in MI. (A): molecular function enrichment of GO terms from DEMs; (B) biological process enrichment of GO terms from DEMs; (C): cellular component enrichment of GO terms from DEMs; (D): KEGG signaling pathway enrichment from DEMs. (TIFF 743 kb)
Rights and permissions
About this article
Cite this article
Wu, T., Wu, Hd., Xu, Zx. et al. Abnormal expression of long non-coding RNAs in myocardial infarction. Heart Vessels 32, 1253–1261 (2017). https://doi.org/10.1007/s00380-017-0990-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00380-017-0990-7