Abstract
Background and Purpose
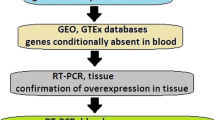
Extrachromosomal circular DNAs (eccDNAs) have been recognized for their significant involvement in numerous biological processes. Nonetheless, the existence and molecular characteristics of eccDNA in the peripheral blood of patients diagnosed with clear cell renal cell carcinoma (ccRCC) have not yet been reported. Our aim was to identify potentially marked plasma eccDNAs in ccRCC patients.
Methods and Materials
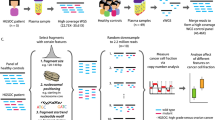
The detection of plasma eccDNA in ccRCC patients and healthy controls was performed using the Tn5-tagmentation and next-generation sequencing (NGS) method. Comparisons were made between ccRCC patients and healthy controls regarding the distribution of length, gene annotation, pattern of junctional nucleotide motif, and expression pattern of plasma eccDNA.
Results
We found 8,568 and 8,150 plasma eccDNAs in ccRCC patients and healthy controls, respectively. There were no statistical differences in the length distribution, gene annotation, and motif signature of plasma eccDNAs between the two groups. A total of 701 differentially expressed plasma eccDNAs were identified, and 25 plasma eccDNAs with potential diagnostic value for ccRCC have been successfully screened. These up-regulated plasma eccDNAs also be indicated to originate from the genomic region of the tumor-associated genes.
Conclusion
This work demonstrates the characterization of plasma eccDNAs in ccRCC and suggests that the up-regulated plasma eccDNAs could be considered as a promising non-invasive biomarker in ccRCC.




Similar content being viewed by others
Data availability
The data that supports the findings of this study are available in the supplementary materials and the raw data are available from the corresponding author upon reasonable request.
References
Dizman N, Philip EJ, Pal SK (2020) Genomic profiling in renal cell carcinoma. Nat Rev Nephrol 16(8):435–451. https://doi.org/10.1038/s41581-020-0301-x
D’Avella C, Abbosh P, Pal SK, Geynisman DM (2020) Mutations in renal cell carcinoma. Urol Oncol 38(10):763–773. https://doi.org/10.1016/j.urolonc.2018.10.027
Møller HD, Mohiyuddin M, Prada-Luengo I, Sailani MR, Halling JF, Plomgaard P et al (2018) Circular DNA elements of chromosomal origin are common in healthy human somatic tissue. Nat Commun 9(1):1069. https://doi.org/10.1038/s41467-018-03369-8
Dillon LW, Kumar P, Shibata Y, Wang YH, Willcox S, Griffith JD et al (2015) Production of Extrachromosomal MicroDNAs Is Linked to Mismatch Repair Pathways and Transcriptional Activity. Cell Rep 11(11):1749–1759. https://doi.org/10.1016/j.celrep.2015.05.020
Hull RM, King M, Pizza G, Krueger F, Vergara X, Houseley J (2019) Transcription-induced formation of extrachromosomal DNA during yeast ageing. PLoS Biol. https://doi.org/10.1371/journal.pbio.3000471
Hull RM, Houseley J (2020) The adaptive potential of circular DNA accumulation in ageing cells. Curr Genet 66(5):889–894. https://doi.org/10.1007/s00294-020-01069-9
Sin STK, Jiang P, Deng J, Ji L, Cheng SH, Dutta A et al (2020) Identification and characterization of extrachromosomal circular DNA in maternal plasma. Proc Natl Acad Sci U S A 117(3):1658–1665. https://doi.org/10.1073/pnas.1914949117
Shibata Y, Kumar P, Layer R, Willcox S, Gagan JR, Griffith JD et al (2012) Extrachromosomal microDNAs and chromosomal microdeletions in normal tissues. Science 336(6077):82–86. https://doi.org/10.1126/science.1213307
Kunisada T, Yamagishi H, Ogita Z, Kirakawa T, Mitsui Y (1985) Appearance of extrachromosomal circular DNAs during in vivo and in vitro ageing of mammalian cells. Mech Ageing Dev 29(1):89–99. https://doi.org/10.1016/0047-6374(85)90050-8
Zhu Y, Gujar AD, Wong CH, Tjong H, Ngan CY, Gong L et al (2021) Oncogenic extrachromosomal DNA functions as mobile enhancers to globally amplify chromosomal transcription. Cancer Cell 39(5):694-707.e7. https://doi.org/10.1016/j.ccell.2021.03.006
Neumann AA, Watson CM, Noble JR, Pickett HA, Tam PP, Reddel RR (2013) Alternative lengthening of telomeres in normal mammalian somatic cells. Genes Dev 27(1):18–23. https://doi.org/10.1101/gad.205062.112
Wang Y, Wang M, Djekidel MN, Chen H, Liu D, Alt FW et al (2021) eccDNAs are apoptotic products with high innate immunostimulatory activity. Nature 599(7884):308–314. https://doi.org/10.1038/s41586-021-04009-w
Paulsen T, Shibata Y, Kumar P, Dillon L, Dutta A (2019) Small extrachromosomal circular DNAs, microDNA, produce short regulatory RNAs that suppress gene expression independent of canonical promoters. Nucleic Acids Res 47(9):4586–4596. https://doi.org/10.1093/nar/gkz155
Koche RP, Rodriguez-Fos E, Helmsauer K, Burkert M, MacArthur IC, Maag J et al (2020) Extrachromosomal circular DNA drives oncogenic genome remodeling in neuroblastoma. Nat Genet 52(1):29–34. https://doi.org/10.1038/s41588-019-0547-z
Cen Y, Fang Y, Ren Y, Hong S, Lu W, Xu J (2022) Global characterization of extrachromosomal circular DNAs in advanced high grade serous ovarian cancer. Cell Death Dis 13(4):342. https://doi.org/10.1038/s41419-022-04807-8
Yang Y, Yang Y, Huang H, Song T, Mao S, Liu D et al (2023) PLCG2 can exist in eccDNA and contribute to the metastasis of non-small cell lung cancer by regulating mitochondrial respiration. Cell Death Dis 14(4):257. https://doi.org/10.1038/s41419-023-05755-7
Lin C, Chen Y, Zhang F, Liu B, Xie C, Song Y (2022) Encoding gene RAB3B exists in linear chromosomal and circular extrachromosomal DNA and contributes to cisplatin resistance of hypopharyngeal squamous cell carcinoma via inducing autophagy. Cell Death Dis 13(2):171. https://doi.org/10.1038/s41419-022-04627-w
Song K, Minami JK, Huang A, Dehkordi SR, Lomeli SH, Luebeck J et al (2022) Plasticity of extrachromosomal and intrachromosomal BRAF amplifications in overcoming targeted therapy dosage challenges. Cancer Discov 12(4):1046–1069. https://doi.org/10.1158/2159-8290.CD-20-0936
Sumiyoshi T, Yamasaki T, Takeda M, Mizuno K, Utsunomiya N, Sakamoto H et al (2021) Detection of von Hippel-Lindau gene mutation in circulating cell-free DNA for clear cell renal cell carcinoma. Cancer Sci 112(8):3363–3374. https://doi.org/10.1111/cas.14972
Geertsen L, Koldby KM, Thomassen M, Kruse T, Lund L (2022) Circulating tumor DNA in patients with renal cell carcinoma. a systematic review of the literature. Eur Urol Open Sci 37:27–35. https://doi.org/10.1016/j.euros.2021.12.006
Li M, Li L, Zheng J, Li Z, Li S, Wang K et al (2023) Liquid biopsy at the frontier in renal cell carcinoma: recent analysis of techniques and clinical application. Mol Cancer 22(1):37. https://doi.org/10.1186/s12943-023-01745-7
Kumar P, Dillon LW, Shibata Y, Jazaeri AA, Jones DR, Dutta A (2017) Normal and cancerous tissues release extrachromosomal circular DNA (eccDNA) into the circulation. Mol Cancer Res 15(9):1197–1205. https://doi.org/10.1158/1541-7786.MCR-17-0095
Zhu J, Zhang F, Du M, Zhang P, Fu S, Wang L (2017) Molecular characterization of cell-free eccDNAs in human plasma. Sci Rep 7(1):10968. https://doi.org/10.1038/s41598-017-11368-w
Wu X, Li P, Yimiti M, Ye Z, Fang X, Chen P et al (2022) Identification and Characterization of Extrachromosomal Circular DNA in Plasma of Lung Adenocarcinoma Patients. Int J Gen Med 15:4781–4791. https://doi.org/10.2147/IJGM.S363425
Penkov D, Zubkova E, Parfyonova Y (2023) Tn5 DNA Transposase in Multi-Omics Research. Methods Protoc 6(2):24. https://doi.org/10.3390/mps6020024
Pang J, Pan X, Lin L, Li L, Yuan S, Han P et al (2022) Characterization of Plasma Extrachromosomal Circular DNA in Gouty Arthritis. Front Genet 13:859513. https://doi.org/10.3389/fgene.2022.859513
Zhou H, He Y, Huang Y, Li R, Zhang H, Xia X et al (2023) Comprehensive analysis of prognostic value, immune implication and biological function of CPNE1 in clear cell renal cell carcinoma. Front Cell Dev Biol 11:1157269. https://doi.org/10.3389/fcell.2023.1157269
Yu L, Dong L, Li H, Liu Z, Luo Z, Duan G et al (2020) Ubiquitination-mediated degradation of SIRT1 by SMURF2 suppresses CRC cell proliferation and tumorigenesis. Oncogene 39(22):4450–4464. https://doi.org/10.1038/s41388-020-1298-0
Wu YH, Huang YF, Chang TH, Chen CC, Wu PY, Huang SC et al (2021) COL11A1 activates cancer-associated fibroblasts by modulating TGF-β3 through the NF-κB/IGFBP2 axis in ovarian cancer cells. Oncogene 40(26):4503–4519. https://doi.org/10.1038/s41388-021-01865-8
Strell C, Norberg KJ, Mezheyeuski A, Schnittert J, Kuninty PR, Moro CF et al (2017) Stroma-regulated HMGA2 is an independent prognostic marker in PDAC and AAC. Br J Cancer 117(1):65–77. https://doi.org/10.1038/bjc.2017.140
Zhu N, Chen X, Zhao J, Fang L, Yao Y, Zhou F et al (2022) Hypoxia-induced LINC00674 facilitates hepatocellular carcinoma progression by activating the NOX1/mTOR signaling pathway. J Cancer 13(11):3177–3188. https://doi.org/10.7150/jca.76458
He H, Tian W, Chen H, Jiang K (2016) MiR-944 functions as a novel oncogene and regulates the chemoresistance in breast cancer. Tumour Biol 37(2):1599–1607. https://doi.org/10.1007/s13277-015-3844-x
Jiang M, Zhong T, Zhang W, Xiao Z, Hu G, Zhou H et al (2017) Reduced expression of miR-205-5p promotes apoptosis and inhibits proliferation and invasion in lung cancer A549 cells by upregulation of ZEB2 and downregulation of erbB3. Mol Med Rep 15(5):3231–3238. https://doi.org/10.3892/mmr.2017.6398
Acknowledgements
We thank Cloud-Seq Biotech Ltd. Co. (Shanghai, China) for eccDNA sequencing service.
Funding
This work was supported by University Natural Science Research Project of Anhui Province (KJ2021ZD0102).
Author information
Authors and Affiliations
Contributions
Gang Feng and Guo-Rong Li designed the study. Rui-Xuan Zhang, Jing-Jing Yang, and Hou-Bao Huang collected the samples and data. Qing Li, Rui-Xuan Zhang, Jing-Jing Yang, and Hou-Bao Huang analyzed the data. Gang Feng, Qing Li and Guo-Rong Li drafted and revised the manuscript. All authors gave final approval of the final version and agreed to its submission.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval and consent to participate
Approval for this research was granted by the Scientific research IRB of Wannan Medical College Yijishan Hospital. Consent was obtained from 6 participants included after they were provided with written information.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Q., Zhang, RX., Yang, JJ. et al. Characterization of extrachromosomal circular DNAs in plasma of patients with clear cell renal cell carcinoma. World J Urol 42, 328 (2024). https://doi.org/10.1007/s00345-024-05031-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00345-024-05031-z