Abstract
Purpose
This study aimed to identify differentially expressed genes (DEGs) and pathways in benign prostatic hyperplasia (BPH) by comprehensive bioinformatics analysis.
Methods
Data of the gene expression microarray (GSE6099) were downloaded from GEO database. DEGs were obtained by GEO2R. Functional and enrichment analyses of selected genes were performed using DAVID database. Protein–protein interaction network was constructed through STRING. Anterior gradient 2 (ARG2) and lumican (LUM) staining in paraffin-embedded specimens from BPH and normal prostate (NP) were detected by immunohistochemistry (IHC). Differences between groups were analyzed by the Student’s t test.
Results
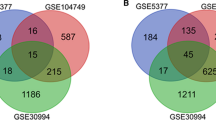
A total of 24 epithelial DEGs and 39 stromal DEGs were determined. The GO analysis results showed that epithelial DEGs between BPH and NP were enriched in biological processes of glucose metabolic process, glucose homeostasis and negative regulation of Rho protein signal transduction. For DEGs in stroma, enriched biological processes included response to ischemia, antigen processing and presentation, cartilage development, T cell costimulation and energy reserve metabolic process. ARG2, as one of the epithelial DEGs, was mainly located in epithelial cells of prostate. In addition, LUM is primarily expressed in the stroma. We further confirmed that compared with NP, the BPH have the lower ARG2 protein level (p = 0.029) and higher LUM protein level (p = 0.003) using IHC.
Conclusions
Our study indicated that there are possible differentially expressed genes in epithelial and stromal cells, such as ARG2 and LUM, which may provide a novel insight for the pathogenesis of BPH.


Similar content being viewed by others
Data availability
Yes.
Code availability
Yes.
References
Chughtai B, Forde JC, Thomas DD, Laor L, Hossack T, Woo HH, Te AE, Kaplan SA (2016) Benign prostatic hyperplasia. Nat Rev Dis Primers 2:16031. https://doi.org/10.1038/nrdp.2016.31
Sarma AV, Wei JT (2012) Clinical practice. Benign prostatic hyperplasia and lower urinary tract symptoms. N Engl J Med 367(3):248–257. https://doi.org/10.1056/NEJMcp1106637
Xiao H, Jiang Y, He W, Xu D, Chen P, Liu D, Liu J, Wang X, DiSanto ME, Zhang X (2020) Identification and functional activity of matrix-remodeling associated 5 (MXRA5) in benign hyperplastic prostate. Aging (Albany NY) 12(9):8605–8621. https://doi.org/10.18632/aging.103175
Bushman W (2009) Etiology, epidemiology, and natural history of benign prostatic hyperplasia. Urol Clin North Am 36(4):403–415. https://doi.org/10.1016/j.ucl.2009.07.003 (v)
Liu D, Shoag JE, Poliak D, Goueli RS, Ravikumar V, Redmond D, Vosoughi A, Fontugne J, Pan H, Lee D, Thomas D, Salari K, Wang Z, Romanel A, Te A, Lee R, Chughtai B, Olumi AF, Mosquera JM, Demichelis F, Elemento O, Rubin MA, Sboner A, Barbieri CE (2020) Integrative multiplatform molecular profiling of benign prostatic hyperplasia identifies distinct subtypes. Nat Commun 11(1):1987. https://doi.org/10.1038/s41467-020-15913-6
Ke ZB, Cai H, Wu YP, Lin YZ, Li XD, Huang JB, Sun XL, Zheng QS, Xue XY, Wei Y, Xu N (2019) Identification of key genes and pathways in benign prostatic hyperplasia. J Cell Physiol 234(11):19942–19950. https://doi.org/10.1002/jcp.28592
Zhou H, Xiang Q, Hu C, Zhang J, Zhang Q, Zhang R (2020) Identification of MMP1 as a potential gene conferring erlotinib resistance in non-small cell lung cancer based on bioinformatics analyses. Hereditas 157(1):32. https://doi.org/10.1186/s41065-020-00145-x
Zhao B, Xu Y, Zhao Y, Shen S, Sun Q (2020) Identification of potential key genes associated with the pathogenesis, metastasis, and prognosis of triple-negative breast cancer on the basis of integrated bioinformatics analysis. Front Oncol 10:856. https://doi.org/10.3389/fonc.2020.00856
Tomlins SA, Mehra R, Rhodes DR, Cao X, Wang L, Dhanasekaran SM, Kalyana-Sundaram S, Wei JT, Rubin MA, Pienta KJ, Shah RB, Chinnaiyan AM (2007) Integrative molecular concept modeling of prostate cancer progression. Nat Genet 39(1):41–51. https://doi.org/10.1038/ng1935
Zhang B, Kwon OJ, Henry G, Malewska A, Wei X, Zhang L, Brinkley W, Zhang Y, Castro PD, Titus M, Chen R, Sayeeduddin M, Raj GV, Mauck R, Roehrborn C, Creighton CJ, Strand DW, Ittmann MM, Xin L (2016) Non-cell-autonomous regulation of prostate epithelial homeostasis by androgen receptor. Mol Cell 63(6):976–989. https://doi.org/10.1016/j.molcel.2016.07.025
De Nunzio C, Presicce F, Tubaro A (2016) Inflammatory mediators in the development and progression of benign prostatic hyperplasia. Nat Rev Urol 13(10):613–626. https://doi.org/10.1038/nrurol.2016.168
Tubaro A, De Nunzio C, Puccini F, Presicce F (2015) The evolving picture of lower urinary tract symptom management. Eur Urol 67(2):271–272. https://doi.org/10.1016/j.eururo.2014.10.016
Wu Y, Ding Y, Cao QF, Qian SB, Wang C, Duan HQ, Gu J, Shen HB (2020) The relationship between glucose homeostasis status and prostate size in aging Chinese males with benign prostatic hyperplasia. World J Urol. https://doi.org/10.1007/s00345-020-03084-4
Yang T, Zhou Y, Wang H, Chen S, Shen M, Hu Y, Wang T, Liu J, Jiang Z, Wang Z, Zhu X, Qian S, Yin X, Lu Q (2020) Insulin exacerbated high glucose-induced epithelial-mesenchymal transition in prostatic epithelial cells BPH-1 and prostate cancer cells PC-3 via MEK/ERK signaling pathway. Exp Cell Res 394(1):112145. https://doi.org/10.1016/j.yexcr.2020.112145
Zi H, Wang XJ, Zhao MJ, Huang Q, Wang XH, Zeng XT (2019) Fasting blood glucose level and hypertension risk in aging benign prostatic hyperplasia patients. Aging (Albany NY) 11(13):4438–4445. https://doi.org/10.18632/aging.102061
Thurmond P, Yang JH, Li Y, Lerner LB, Azadzoi KM (2015) Structural modifications of the prostate in hypoxia, oxidative stress, and chronic ischemia. Korean J Urol 56(3):187–196. https://doi.org/10.4111/kju.2015.56.3.187
Jin Cho W, Pyo JS (2020) Immunohistochemical analysis of the impact of ischemic change in benign prostatic hyperplasia. Pathol Res Pract 216(1):152694. https://doi.org/10.1016/j.prp.2019.152694
Alavi M, Mah V, Maresh EL, Bagryanova L, Horvath S, Chia D, Goodglick L, Liu AY (2015) High expression of AGR2 in lung cancer is predictive of poor survival. BMC Cancer 15:655. https://doi.org/10.1186/s12885-015-1658-2
Broustas CG, Hopkins KM, Panigrahi SK, Wang L, Virk RK, Lieberman HB (2019) RAD9A promotes metastatic phenotypes through transcriptional regulation of anterior gradient 2 (AGR2). Carcinogenesis 40(1):164–172. https://doi.org/10.1093/carcin/bgy131
Zhang Y, Ali TZ, Zhou H, D’Souza DR, Lu Y, Jaffe J, Liu Z, Passaniti A, Hamburger AW (2010) ErbB3 binding protein 1 represses metastasis-promoting gene anterior gradient protein 2 in prostate cancer. Cancer Res 70(1):240–248. https://doi.org/10.1158/0008-5472.CAN-09-2904
Tohti M, Li J, Tang C, Wen G, Abdujilil A, Yizim P, Ma C (2017) Serum AGR2 as a useful biomarker for pituitary adenomas. Clin Neurol Neurosurg 154:19–22. https://doi.org/10.1016/j.clineuro.2017.01.004
Bu H, Bormann S, Schafer G, Horninger W, Massoner P, Neeb A, Lakshmanan VK, Maddalo D, Nestl A, Sultmann H, Cato AC, Klocker H (2011) The anterior gradient 2 (AGR2) gene is overexpressed in prostate cancer and may be useful as a urine sediment marker for prostate cancer detection. Prostate 71(6):575–587. https://doi.org/10.1002/pros.21273
Shi X, Zhao Y, Fan C (2019) Zbed3 promotes proliferation and invasion of lung cancer partly through regulating the function of Axin-Gsk3beta complex. J Cell Mol Med 23(2):1014–1021. https://doi.org/10.1111/jcmm.14001
Wang T, Wang CJ, Tian S, Song HB (2019) Overexpressed IGFBP5 promotes cell proliferation and inhibits apoptosis of nucleus pulposus derived from rats with disc degeneration through inactivating the ERK/MAPK axis. J Cell Biochem 120(11):18782–18792. https://doi.org/10.1002/jcb.29191
Liu Y, Wang F, Liu Y, Yao Y, Lv X, Dong B, Li J, Ren S, Yao Y, Xu Y (2016) RNF135, RING finger protein, promotes the proliferation of human glioblastoma cells in vivo and in vitro via the ERK pathway. Sci Rep 6:20642. https://doi.org/10.1038/srep20642
Coulson-Thomas VJ, Coulson-Thomas YM, Gesteira TF, Andrade de Paula CA, Carneiro CR, Ortiz V, Toma L, Kao WW, Nader HB (2013) Lumican expression, localization and antitumor activity in prostate cancer. Exp Cell Res 319(7):967–981. https://doi.org/10.1016/j.yexcr.2013.01.023
Xiao D, Liang T, Zhuang Z, He R, Ren J, Jiang S, Zhu L, Wang K, Shi D (2020) Lumican promotes joint fibrosis through TGF-β signaling. FEBS Open Bio. https://doi.org/10.1002/2211-5463.12974
Papoutsidakis A, Giatagana EM, Berdiaki A, Spyridaki I, Spandidos DA, Tsatsakis A, Tzanakakis GN, Nikitovic D (2020) Lumican mediates HTB94 chondrosarcoma cell growth via an IGFIR/Erk1/2 axis. Int J Oncol 57(3):791–803. https://doi.org/10.3892/ijo.2020.5094
Middleton LW, Shen Z, Varma S, Pollack AS, Gong X, Zhu S, Zhu C, Foley JW, Vennam S, Sweeney RT, Tu K, Biscocho J, Eminaga O, Nolley R, Tibshirani R, Brooks JD, West RB, Pollack JR (2019) Genomic analysis of benign prostatic hyperplasia implicates cellular re-landscaping in disease pathogenesis. JCI Insight. https://doi.org/10.1172/jci.insight.129749
Li L, Ameri AH, Wang S, Jansson KH, Casey OM, Yang Q, Beshiri ML, Fang L, Lake RG, Agarwal S, Alilin AN, Xu W, Yin J, Kelly K (2019) EGR1 regulates angiogenic and osteoclastogenic factors in prostate cancer and promotes metastasis. Oncogene 38(35):6241–6255. https://doi.org/10.1038/s41388-019-0873-8
Acknowledgements
The authors are very grateful to the urology department of Beijing Tongren Hospital for their selfless help in this work.
Funding
This work was supported by the National Natural Science Foundation of China (Grant No. 81772698 to Hao Ping) and the Open Research Fund from Beijing Advanced Innovation Center for Big Data-Based Precision Medicine, Beijing Tongren Hospital, Beihang University and Capital Medical University (Grant No. BHTR-KFJJ-202005 to Hao Ping).
Author information
Authors and Affiliations
Contributions
HP: protocol/project development. PX, MDW and DL: data collection or management. WY, ZD, DG, PX and YXH: data analysis. PX and HP: manuscript writing/editing.
Corresponding author
Ethics declarations
Conflict of interest
All the authors declare that there are no conflict of interests.
Ethical standards
This study has been approved by the appropriate ethics committee.
Consent to participate
Yes.
Consent for publication
Yes.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Xiang, P., Liu, D., Guan, D. et al. Identification of key genes in benign prostatic hyperplasia using bioinformatics analysis. World J Urol 39, 3509–3516 (2021). https://doi.org/10.1007/s00345-021-03625-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00345-021-03625-5