Abstract
Objectives
Our purpose was to differentiate between malignant from benign soft tissue neoplasms using a combination of MRI-based radiomics metrics and machine learning.
Methods
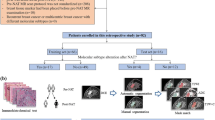
Our retrospective study identified 128 histologically diagnosed benign (n = 36) and malignant (n = 92) soft tissue lesions. 3D ROIs were manually drawn on 1 sequence of interest and co-registered to other sequences obtained during the same study. One thousand seven hundred eight radiomics features were extracted from each ROI. Univariate analyses with supportive ROC analyses were conducted to evaluate the discriminative power of predictive models constructed using Real Adaptive Boosting (Adaboost) and Random Forest (RF) machine learning approaches.
Results
Univariate analyses demonstrated that 36.89% of individual radiomics varied significantly between benign and malignant lesions at the p ≤ 0.05 level. Adaboost and RF performed similarly well, with AUCs of 0.77 (95% CI 0.68–0.85) and 0.72 (95% CI 0.63–0.81), respectively, after 10-fold cross-validation. Restricting the machine learning models to only sequences extracted from T2FS and STIR sequences maintained comparable performance, with AUCs of 0.73 (95% CI 0.64–0.82) and 0.75 (95% CI 0.65–0.84), respectively.
Conclusion
Machine learning decision classifiers constructed from MRI-based radiomics features show promising ability to preoperatively discriminate between benign and malignant soft tissue masses. Our approach maintains applicability even when the dataset is restricted to T2FS and STIR fluid-sensitive sequences, which may bolster practicality in clinical application scenarios by eliminating the need for complex co-registrations for multisequence analysis.
Key Points
• Predictive models constructed from MRI-based radiomics data and machine learning–augmented approaches yielded good discriminative power to correctly classify benign and malignant lesions on preoperative scans, with AUCs of 0.77 (95% CI 0.68–0.85) and 0.72 (95% CI 0.63–0.81) for Real Adaptive Boosting (Adaboost) and Random Forest (RF), respectively.
• Restricting the models to only use metrics extracted from T2 fat-saturated (T2FS) and Short-Tau Inversion Recovery (STIR) sequences yielded similar performance, with AUCs of 0.73 (95% CI 0.64–0.82) and 0.75 (95% CI 0.65–0.84) for Adaboost and RF, respectively.
• Radiomics-based machine learning decision classifiers constructed from multicentric data more closely mimic the real-world practice environment and warrant additional validation ahead of prospective implementation into clinical workflows.






Similar content being viewed by others
Abbreviations
- 3D:
-
3-dimensional 3D
- AUC:
-
Area under the curve
- Adaboost:
-
Real Adaptive Boosting
- FIESTA:
-
Fast Imaging Employing Steady-state Acquisition
- GLCM:
-
Gray Level Co-Occurrence Matrix
- GLSZM:
-
Gray Level Size Zone Matrix
- LTE:
-
Laws Texture Energy
- NPV:
-
Negative predictive value
- PD:
-
Proton density
- PPV:
-
Positive predictive value
- RF:
-
Random Forest
- ROC:
-
Receiver operating characteristic
- ROI:
-
Region of interest
- STIR:
-
Short-Tau Inversion Recovery
- STS:
-
Soft tissue sarcoma
- T2FS:
-
T2 fat-saturated
- VIBE:
-
Volumetric Interpolated Breath-hold Examination
- WHO:
-
World Health Organization
References
Zhang Y, Zhu Y, Shi X et al (2019) Soft tissue sarcomas: preoperative predictive histopathological grading based on radiomics of MRI. Acad Radiol 26(9):1262–1268
Zhao F, Ahlawat S, Farahani SJ et al (2014) Can MR imaging be used to predict tumor grade in soft-tissue sarcoma? Radiology 272(1):192–201
Vallieres M, Freeman CR, Skamene SR, El Naqa I (2015) A radiomics model from joint FDG-PET and MRI texture features for the prediction of lung metastases in soft-tissue sarcomas of the extremities. Phys Med Biol 60(14):5471–5496
Patel DB, Matcuk GR Jr (2018) Imaging of soft tissue sarcomas. Chin Clin Oncol 7(4):35
Fields BKK, Hwang D, Cen S et al (2020) Quantitative magnetic resonance imaging (q-MRI) for the assessment of soft-tissue sarcoma treatment response: a narrative case review of technique development. Clin Imaging 63:83–93
Baheti AD, O'Malley RB, Kim S et al (2016) Soft-tissue sarcomas: an update for radiologists based on the revised 2013 World Health Organization classification. AJR Am J Roentgenol 206(5):924–932
Fletcher CDM, Bridge JA, Hogendoorn PCW, Mertens F (eds) (2013) WHO classification of tumours of soft tissue and bone, 4th edn. International Agency for Research on Cancer (IARC), Lyon, FR
De La Hoz PM, Dick E, Bhumbra R, Pollock R, Sandhu R, Saifuddin A (2017) Surgical considerations when reporting MRI studies of soft tissue sarcoma of the limbs. Skeletal Radiol 46(12):1667–1678
Manaster BJ (2013) Soft-tissue masses: optimal imaging protocol and reporting. AJR Am J Roentgenol 201(3):505–514
Chhabra A, Soldatos T (2012) Soft-tissue lesions: when can we exclude sarcoma? AJR Am J Roentgenol 199(6):1345–1357
Wu JS, Hochman MG (2009) Soft-tissue tumors and tumorlike lesions: a systematic imaging approach. Radiology 253(2):297–316
Wang H, Nie P, Wang Y et al (2020) Radiomics nomogram for differentiating between benign and malignant soft-tissue masses of the extremities. J Magn Reson Imaging 51(1):155–163
Wang H, Zhang J, Bao S et al (2020) Preoperative MRI-based radiomic machine-learning nomogram may accurately distinguish between benign and malignant soft-tissue lesions: a two-center study. J Magn Reson Imaging 52(3):873–882
Fayad LM, Jacobs MA, Wang X, Carrino JA, Bluemke DA (2012) Musculoskeletal tumors: how to use anatomic, functional, and metabolic MR techniques. Radiology 265(2):340–356
Crombe A, Alberti N, Stoeckle E et al (2016) Soft tissue masses with myxoid stroma: can conventional magnetic resonance imaging differentiate benign from malignant tumors? Eur J Radiol 85(10):1875–1882
Arkun R, Argin M (2014) Pitfalls in MR imaging of musculoskeletal tumors. Semin Musculoskelet Radiol 18(1):63–78
Hirschmann A, van Praag VM, Haas RL, van de Sande MAJ, Bloem JL (2020) Can we use MRI to detect clinically silent recurrent soft-tissue sarcoma? Eur Radiol 30(9):4724–4733
Juntu J, Sijbers J, De Backer S, Rajan J, Van Dyck D (2010) Machine learning study of several classifiers trained with texture analysis features to differentiate benign from malignant soft-tissue tumors in T1-MRI images. J Magn Reson Imaging 31(3):680–689
Crombe A, Marcellin PJ, Buy X et al (2019) Soft-tissue sarcomas: assessment of MRI features correlating with histologic grade and patient outcome. Radiology 291(3):710–721
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278(2):563–577
Aerts HJ (2016) The potential of radiomic-based phenotyping in precision medicine: a review. JAMA Oncol 2(12):1636–1642
Hwang DH, Varghese BA, Chang M, et al (2017) Radiomics-based quantitative biomarker discovery: development of a robust image processing infrastructure. Proc SPIE 10160, 12th International Symposium on Medical Information Processing and Analysis, 1016017, January 26, 2017
Varghese BA, Cen SY, Hwang DH, Duddalwar VA (2019) Texture analysis of imaging: what radiologists need to know. AJR Am J Roentgenol 212(3):520–528
Varghese BA, Hwang D, Cen SY et al (2019) Reliability of CT-based texture features: phantom study. J Appl Clin Med Phys 20(8):155–163
Shafiq-Ul-Hassan M, Latifi K, Zhang G, Ullah G, Gillies R, Moros E (2018) Voxel size and gray level normalization of CT radiomic features in lung cancer. Sci Rep 8(1):10545
Whitney HM, Li H, Ji Y, Liu P, Giger ML (2020) Harmonization of radiomic features of breast lesions across international DCE-MRI datasets. J Med Imaging (Bellingham) 7(1):012707
Limkin EJ, Sun R, Dercle L et al (2017) Promises and challenges for the implementation of computational medical imaging (radiomics) in oncology. Ann Oncol 28(6):1191–1206
Peeken JC, Spraker MB, Knebel C et al (2019) Tumor grading of soft tissue sarcomas using MRI-based radiomics. EBioMedicine 48:332–340
Crombe A, Perier C, Kind M et al (2019) T2-based MRI delta-radiomics improve response prediction in soft-tissue sarcomas treated by neoadjuvant chemotherapy. J Magn Reson Imaging 50(2):497–510
Crombe A, Le Loarer F, Sitbon M et al (2020) Can radiomics improve the prediction of metastatic relapse of myxoid/round cell liposarcomas? Eur Radiol 30(5):2413–2424
Tagliafico AS, Bignotti B, Rossi F, Valdora F, Martinoli C (2019) Local recurrence of soft tissue sarcoma: a radiomic analysis. Radiol Oncol 53(3):300–306
Spraker MB, Wootton LS, Hippe DS et al (2019) MRI radiomic features are independently associated with overall survival in soft tissue sarcoma. Adv Radiat Oncol 4(2):413–421
Corino VDA, Montin E, Messina A et al (2018) Radiomic analysis of soft tissues sarcomas can distinguish intermediate from high-grade lesions. J Magn Reson Imaging 47(3):829–840
Wang H, Chen H, Duan S, Hao D, Liu J (2020) Radiomics and machine learning with multiparametric preoperative MRI may accurately predict the histopathological grades of soft tissue sarcomas. J Magn Reson Imaging 51(3):791–797
Li L, Wang K, Ma X et al (2019) Radiomic analysis of multiparametric magnetic resonance imaging for differentiating skull base chordoma and chondrosarcoma. Eur J Radiol 118:81–87
Malinauskaite I, Hofmeister J, Burgermeister S et al (2020) Radiomics and machine learning differentiate soft-tissue lipoma and liposarcoma better than musculoskeletal radiologists. Sarcoma 2020:7163453
Xie H, Hu J, Zhang X, Ma S, Liu Y, Wang X (2019) Preliminary utilization of radiomics in differentiating uterine sarcoma from atypical leiomyoma: comparison on diagnostic efficacy of MRI features and radiomic features. Eur J Radiol 115:39–45
Xie H, Zhang X, Ma S, Liu Y, Wang X (2019) Preoperative differentiation of uterine sarcoma from leiomyoma: comparison of three models based on different segmentation volumes using radiomics. Mol Imaging Biol 21(6):1157–1164
Gulati M, Hu JS, Desai B, Hwang DH, Grant EG, Duddalwar VA (2015) Contrast-enhanced sonography for monitoring neoadjuvant chemotherapy in soft tissue sarcomas. J Ultrasound Med 34(8):1489–1499
Friston K, Ashburner J, Kiebel S, Nichols T, Penny W (eds) (2007) Statistical parametric mapping: the analysis of functional brain images, 1st edn. Academic Press, London
Fan TW, Malhi H, Varghese B et al (2019) Computed tomography-based texture analysis of bladder cancer: differentiating urothelial carcinoma from micropapillary carcinoma. Abdom Radiol (NY) 44(1):201–208
Varghese B, Chen F, Hwang D et al (2019) Objective risk stratification of prostate cancer using machine learning and radiomics applied to multiparametric magnetic resonance images. Sci Rep 9(1):1570
Huhdanpaa H, Hwang D, Cen S et al (2015) CT prediction of the Fuhrman grade of clear cell renal cell carcinoma (RCC): towards the development of computer-assisted diagnostic method. Abdom Imaging 40(8):3168–3174
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B Methodol 57(1):289–300
Hastie T, Tibshirani R, Friedman J (2009) The elements of statistical learning: data mining, inference, and prediction, 2nd edn. Springer, New York, NY, p 363
Loh W-Y (2009) Improving the precision of classification trees. Ann Appl Stat 3(4):1710–1737
Laws KI (1980) Rapid texture identification. Proc SPIE 0238, Image Processing for Missile Guidance, December 23, 1980
Miller P, Astley S (1992) Classification of breast-tissue by texture analysis. Image Vision Comput 10(5):277–282
Chu Y, Li L, Goldgof DB, Qui Y, Clark RA (2003) Classification of masses on mammograms using support vector machine. Proc SPIE 5032, Medical Imaging 2003: Image Processing, May 15, 2003
Cox G, Hoare F, de Jager G (1992) Experiments in lung cancer nodule detection using texture analysis and neural network classifiers. Third South African Workshop on Pattern Recognition 31:136–142
Dilger SK, Judisch A, Uthoff J, Hammond E, Newell JD, Sieren JC (2015) Improved pulmonary nodule classification utilizing lung parenchyma texture features. Proc SPIE 9414, Medical Imaging 2015: Computer-Aided Diagnosis, 94142T, March 20, 2015
Barata C, Marques JS, Mendonça T (2013) Bag-of-features classification model for the diagnose of melanoma in dermoscopy images using color and texture descriptors. In: Kamel M, Campilho A (eds) Image analysis and recognition. Lecture Notes in Computer Science, vol 7950. ICIAR 2013:547–555
Tixier F, Le Rest CC, Hatt M et al (2011) Intratumor heterogeneity characterized by textural features on baseline 18F-FDG PET images predicts response to concomitant radiochemotherapy in esophageal cancer. J Nucl Med 52(3):369–378
Cook GJ, Yip C, Siddique M et al (2013) Are pretreatment 18F-FDG PET tumor textural features in non-small cell lung cancer associated with response and survival after chemoradiotherapy? J Nucl Med 54(1):19–26
Parmar C, Leijenaar RT, Grossmann P et al (2015) Radiomic feature clusters and prognostic signatures specific for Lung and Head & Neck cancer. Sci Rep 5:11044
Leijenaar RT, Carvalho S, Hoebers FJ et al (2015) External validation of a prognostic CT-based radiomic signature in oropharyngeal squamous cell carcinoma. Acta Oncol 54(9):1423–1429
Tokuda O, Harada Y, Matsunaga N (2009) MRI of soft-tissue tumors: fast STIR sequence as substitute for T1-weighted fat-suppressed contrast-enhanced spin-echo sequence. AJR Am J Roentgenol 193(6):1607–1614
Couronne R, Probst P, Boulesteix AL (2018) Random forest versus logistic regression: a large-scale benchmark experiment. BMC Bioinformatics 19(1):270
Traverso A, Kazmierski M, Zhovannik I et al (2020) Machine learning helps identifying volume-confounding effects in radiomics. Phys Med 71:24–30
Acknowledgements
The authors would like to thank Rosaura Diaz for her assistance with obtaining IRB approval. The authors would also like to thank Robert Fields CPA, MBA, for his assistance with restructuring the data output for interpretation and reporting. We thank the Radiological Society of North America’s Research & Education Foundation for their support and funding of our work.
Funding
This study was funded by Radiological Society of North America Research Medical Student Grant RMS#1909.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is G.R. Matcuk Jr.
Conflict of interest
GRM is a consultant for Canon Medical Systems, USA. VD is a consultant for Radmetrix and serves on the advisory board for DeepTek. The remaining authors declare that they have no other disclosures.
Statistics and biometry
SYC and XL have significant statistical expertise.
Informed consent
Written informed consent was waived by the Institutional Review Board.
Ethical approval
Institutional Review Board approval was obtained.
Study subjects overlap
8 study subjects have been previously reported in Reference 5, 39.
Methodology
•Retrospective
•Diagnostic or prognostic study
•Performed at one institution
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(XLSX 314 kb)
Rights and permissions
About this article
Cite this article
Fields, B.K.K., Demirjian, N.L., Hwang, D.H. et al. Whole-tumor 3D volumetric MRI-based radiomics approach for distinguishing between benign and malignant soft tissue tumors. Eur Radiol 31, 8522–8535 (2021). https://doi.org/10.1007/s00330-021-07914-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-021-07914-w