Abstract
Objectives
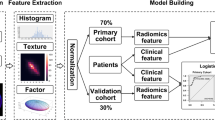
To develop a model using radiomic features extracted from MR images to distinguish radiation necrosis from tumour progression in brain metastases after Gamma Knife radiosurgery.
Methods
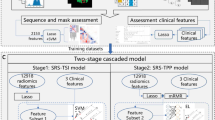
We retrospectively identified 87 patients with pathologically confirmed necrosis (24 lesions) or progression (73 lesions) and calculated 285 radiomic features from four MR sequences (T1, T1 post-contrast, T2, and fluid-attenuated inversion recovery) obtained at two follow-up time points per lesion per patient. Reproducibility of each feature between the two time points was calculated within each group to identify a subset of features with distinct reproducible values between two groups. Changes in radiomic features from one time point to the next (delta radiomics) were used to build a model to classify necrosis and progression lesions.
Results
A combination of five radiomic features from both T1 post-contrast and T2 MR images were found to be useful in distinguishing necrosis from progression lesions. Delta radiomic features with a RUSBoost ensemble classifier had an overall predictive accuracy of 73.2% and an area under the curve value of 0.73 in leave-one-out cross-validation.
Conclusions
Delta radiomic features extracted from MR images have potential for distinguishing radiation necrosis from tumour progression after radiosurgery for brain metastases.
Key points
• Some radiomic features showed better reproducibility for progressive lesions than necrotic ones
• Delta radiomic features can help to distinguish radiation necrosis from tumour progression
• Delta radiomic features had better predictive value than did traditional radiomic features




Similar content being viewed by others
Change history
14 March 2018
The original version of this article, published on 24 November 2017, unfortunately contained a mistake.
Abbreviations
- AUC:
-
Area under the curve
- CCC:
-
Concordance correlation coefficient
- COM:
-
Co-occurrence matrix
- FLAIR:
-
Fluid-attenuated inversion recovery
- HOG:
-
Histogram of oriented gradients
- IBEX:
-
Imaging Biomarker Explorer
- MRI:
-
Magnetic resonance imaging
- NGTDM:
-
Neighbourhood grey-tone difference matrix
- RLM:
-
Run length matrix
- ROC:
-
Receiver operating characteristic
- T1c:
-
T1 weighted post-contrast
References
Nayak L, Lee EQ, Wen PY (2012) Epidemiology of brain metastases. Current Oncology Reports 14:48–54
Kondziolka D, Martin JJ, Flickinger JC et al (2005) Long-term survivors after gamma knife radiosurgery for brain metastases. Cancer 104:2784–2791
Elaimy AL, Mackay AR, Lamoreaux WT et al (2011) Clinical outcomes of stereotactic radiosurgery in the treatment of patients with metastatic brain tumors. World Neurosurg 75:673–683
Minniti G, Clarke E, Lanzetta G et al (2011) Stereotactic radiosurgery for brain metastases: analysis of outcome and risk of brain radionecrosis. Radiation Oncology 6:48
Shaw E, Scott C, Souhami L et al (2000) Single dose radiosurgical treatment of recurrent previously irradiated primary brain tumors and brain metastases: Final report of RTOG protocol 90-05. International Journal of Radiation Oncology Biology Physics 47:291–298
Chuang MT, Liu YS, Tsai YS, Chen YC, Wang CK (2016) Differentiating Radiation-Induced Necrosis from Recurrent Brain Tumor Using MR Perfusion and Spectroscopy: A Meta-Analysis. PLoS ONE 11
Lai G, Mahadevan A, Hackney D et al (2015) Diagnostic Accuracy of PET, SPECT, and Arterial Spin-Labeling in Differentiating Tumor Recurrence from Necrosis in Cerebral Metastasis after Stereotactic Radiosurgery. American Journal of Neuroradiology 36:2250–2255
Aerts HJWL, Velazquez ER, Leijenaar RTH et al (2014) Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nature Communications 5
Fehr D, Veeraraghavan H, Wibmer A et al (2015) Automatic classification of prostate cancer Gleason scores from multiparametric magnetic resonance images. Proceedings of the National Academy of Sciences 112:E6265–E6273
Gevaert O, Mitchell LA, Achrol AS et al (2014) Glioblastoma multiforme: exploratory radiogenomic analysis by using quantitative image features. Radiology 273:168–174
Gillies RJ, Kinahan PE, Hricak H (2015) Radiomics: images are more than pictures, they are data. Radiology 278:563–577
Itakura H, Achrol AS, Mitchell LA et al (2015) Magnetic resonance image features identify glioblastoma phenotypic subtypes with distinct molecular pathway activities. Science translational medicine 7:303ra138–303ra138
Yamamoto S, Han W, Kim Y et al (2015) Breast cancer: radiogenomic biomarker reveals associations among dynamic contrast-enhanced MR imaging, long noncoding RNA, and metastasis. Radiology 275:384–392
Mattes D, Haynor DR, Vesselle H, Lewellen TK, Eubank W (2003) PET-CT image registration in the chest using free-form deformations. IEEE Transactions On Medical Imaging 22:120–128
Zhang L, Fried DV, Fave XJ, Hunter LA, Yang J, Court LE (2015) IBEX: An open infrastructure software platform to facilitate collaborative work in radiomics. Medical Physics 42:1341–1353
Yang J, Zhang L, Fave XJ et al (2016) Uncertainty analysis of quantitative imaging features extracted from contrast-enhanced CT in lung tumors. Comput Med Imaging Graph 48:1–8
Burger W, Burge MJ (2013) Edge-Preserving Smoothing FiltersPrinciples of Digital Image Processing. Springer, pp 119-167
Cunliffe AR, Armato Iii SG, Fei XM, Tuohy RE, Al-Hallaq HA (2013) Lung texture in serial thoracic CT scans: Registration-based methods to compare anatomically matched regionsa. Medical Physics 40:061906
Fried DV, Tucker SL, Zhou S et al (2014) Prognostic value and reproducibility of pretreatment CT texture features in stage III non-small cell lung cancer. Int J Radiat Oncol Biol Phys 90:834–842
Haralick RM, Shanmugam K, Dinstein I (1973) Textural features for image classification. IEEE T. Syst. Man Cyb. 3:610–621
Tang X (1998) Texture information in run-length matrices. IEEE Transactions on Image Processing 7:1602–1609
Legland D, Kiêu K, Devaux M-F (2007) Computation of Minkowski measures on 2D and 3D binary images. Image Anal Stereol 26:83–92
Basu S (2012) Developing predictive models for lung tumor analysis.
Amadasun M, King R (1989) Textural Features Corresponding to Textural Properties. Ieee Transactions on Systems Man and Cybernetics 19:1264–1274
Dalal N, Triggs B (2005) Histograms of oriented gradients for human detection IEEE Computer Society Conference on Computer Vision and Pattern Recognition (CVPR) pp 886-893
Tiwari P, Prasanna P, Wolansky L et al (2016) Computer-Extracted Texture Features to Distinguish Cerebral Radionecrosis from Recurrent Brain Tumors on Multiparametric MRI: A Feasibility Study. AJNR Am J Neuroradiol. https://doi.org/10.3174/ajnr.A4931
Galizia MS, Töre HG, Chalian H, McCarthy R, Salem R, Yaghmai V (2012) MDCT necrosis quantification in the assessment of hepatocellular carcinoma response to yttrium 90 radioembolization therapy: comparison of two-dimensional and volumetric techniques. Academic Radiology 19:48–54
Lin LIK (1989) A Concordance Correlation Coefficient to Evaluate Reproducibility. Biometrics 45:255–268
Duda RO, Hart PE, Stork DG (2001) Pattern Classification. John Wiley and Sons, Inc.
Seiffert C, Khoshgoftaar TM, Van Hulse J, Napolitano A (2010) RUSBoost: A Hybrid Approach to Alleviating Class Imbalance. Ieee Transactions on Systems Man and Cybernetics Part a-Systems and Humans 40:185-197
Barajas RF, Chang JS, Segal MR et al (2009) Differentiation of Recurrent Glioblastoma Multiforme from Radiation Necrosis after External Beam Radiation Therapy with Dynamic Susceptibility-weighted Contrast-enhanced Perfusion MR Imaging. Radiology 253:486–496
Kickingereder P, Burth S, Wick A et al (2016) Radiomic Profiling of Glioblastoma: Identifying an Imaging Predictor of Patient Survival with Improved Performance over Established Clinical and Radiologic Risk Models. Radiology 280:880–889
Li H, Zhu Y, Burnside ES et al (2016) MR Imaging Radiomics Signatures for Predicting the Risk of Breast Cancer Recurrence as Given by Research Versions of MammaPrint, Oncotype DX, and PAM50 Gene Assays. Radiology:152110
Larroza A, Moratal D, Paredes-Sánchez A et al (2015) Support vector machine classification of brain metastasis and radiation necrosis based on texture analysis in MRI. Journal of Magnetic Resonance Imaging 42:1362–1368
Shinohara RT, Sweeney EM, Goldsmith J et al (2014) Statistical normalization techniques for magnetic resonance imaging. NeuroImage: Clinical 6:9–19
Acknowledgements
The authors would like to thank Christine Wogan for reviewing this manuscript.
This work was presented in part at the 2016 American Association of Physicists in Medicine Annual Meeting.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Jing Li.
Conflict of interest
The authors of this manuscript declare no relationships with any companies, whose products or services may be related to the subject matter of the article.
Funding
This study has received funding by National Institutes of Health through Cancer Center Support Grant P30CA016672.
Statistics and biometry
No complex statistical methods were necessary for this paper.
Informed consent
Written informed consent was waived by the Institutional Review Board.
Ethical approval
Institutional Review Board approval was obtained.
Methodology
• retrospective
• experimental
• performed at one institution
Additional information
The original version of this article was revised: The presentation of Table 2 was incorrect.
Rights and permissions
About this article
Cite this article
Zhang, Z., Yang, J., Ho, A. et al. A predictive model for distinguishing radiation necrosis from tumour progression after gamma knife radiosurgery based on radiomic features from MR images. Eur Radiol 28, 2255–2263 (2018). https://doi.org/10.1007/s00330-017-5154-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-017-5154-8