Abstract
Key message
ABI5 functions in ABA-mediated anthocyanin accumulation in plant response to low phosphate.
Abstract
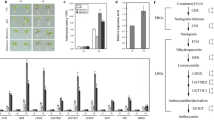
Low phosphate (LP)-induced anthocyanin biosynthesis and accumulation play an important role in plant adaptive response to phosphate starvation conditions. However, whether and how the stress phytohormone abscisic acid (ABA) participates in LP-induced anthocyanin accumulation remain elusive. Here, we report that ABA is required for LP-induced anthocyanin accumulation in Arabidopsis thaliana. Disrupting ABA DEFICIENT2 (ABA2), a key ABA-biosynthetic gene, or BETA-GLUCOSIDASE1 (BG1), a major gene implicated in converting conjugated ABA to active ABA, significantly impairs LP-induced anthocyanin accumulation, as LP-induced expression of the anthocyanin-biosynthetic genes Chalcone Synthase (CHS) is dampened in the aba2 and bg1 mutant. In addition, LP-induced anthocyanin accumulation is defective in the mutants of ABA signaling pathway, including ABA receptors, ABA Insensitive2, and the transcription factors ABA Insensitive5 (ABI5), suggesting a role of ABI5 in ABA-mediated upregulation of anthocyanin-biosynthetic genes in plant response to LP. Indeed, LP-induced expression of CHS is repressed in the abi5-7 mutant but further promoted in the ABI5-overexpressing plants compared to the wild-type. Moreover, ABI5 can bind to and transcriptionally activate CHS, and the defectiveness of LP-induced anthocyanin accumulation in abi5-7 can be restored by overexpressing CHS. Collectively, our findings illustrates that ABI5 functions in ABA-mediated LP-induced anthocyanin accumulation in Arabidopsis.






Similar content being viewed by others
Data availability
All the data generated or analyzed during this study are included in this published article.
References
Besseau S, Hoffmann L, Geoffroy P, Lapierre C, Pollet B, Legrand M (2007) Flavonoid accumulation in Arabidopsis repressed in lignin synthesis affects auxin transport and plant growth. Plant Cell 19:148–162
Carles C, Bies-Etheve N, Aspart L, Léon-Kloosterziel KM, Koornneef M, Echeverria M, Delseny M (2002) Regulation of Arabidopsis thaliana Em genes: role of ABI5. Plant J: Cell Mol Biol 30(3):373–383. https://doi.org/10.1046/j.1365-313x.2002.01295.x
Chen YF, Li LQ, Xu Q, Kong YH, Wang H, Wu WH (2009) The WRKY6 transcription factor modulates PHOSPHATE1 expression in response to low Pi stress in Arabidopsis. Plant Cell 21:3554–3566
Chen J, Wang Y, Wang F, Yang J, Gao M, Li C et al (2015) The rice CK2 kinase regulates trafficking of phosphate transporters in response to phosphate levels. Plant Cell 27:711–723
Chen Q, Hu T, Li X, Song CP, Zhu JK, Chen L, Zhao Y (2022) Phosphorylation of SWEET sucrose transporters regulates plant root:shoot ratio under drought. Nat Plants 8:68–77
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacteriummediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Gonzalez E, Solano R, Rubio V, Leyva A, Paz-Ares J (2005) PHOSPHATE TRANSPORTER TRAFFIC FACILITATOR1 is a plant-specific SEC12-related protein that enables the endoplasmic reticulum exit of a highaffinity phosphate transporter in Arabidopsis. Plant Cell 17:3500–3512
Gou JY, Felippes FF, Liu CJ, Weigel D, Wang JW (2011) Negative regulation of anthocyanin biosynthesis in Arabidopsis by a miR156-targeted SPL transcription factor. Plant Cell 23:1512–1522
Guo JX, Song RF, Lu KK, Zhang Y, Chen HH, Zuo JX, Li TT, Li XF, Liu WC (2022a) Cycc1;1 negatively modulates ABA signaling by interacting with and inhibiting ABI5 during seed germination. Plant Physiol 190:2812–2827
Guo M, Zhang Y, Jia X, Wang X, Zhang Y, Liu J et al (2022b) Alternative splicing of regulator of leaf inclination 1 modulates phosphate starvation signaling and growth in plants. Plant Cell 34:3319–3338
Gould KS (2004) Nature’s Swiss army knife: the diverse protective roles of anthocyanins in leaves. J Biomed Biotechnol 2004:314–320
He Y, Zhang X, Li L, Sun Z, Li J, Chen X, Hong G (2021) SPX4 interacts with both PHR1 and PAP1 to regulate critical steps in phosphorus-status-dependent anthocyanin biosynthesis. New Phytol 230:205–217
Huang KL, Ma GJ, Zhang ML, Xiong H, Wu H, Zhao CZ et al (2018) The ARF7 and ARF19 transcription factors positively regulate PHOSPHATE STARVATION RESPONSE1 in Arabidopsis roots. Plant Physiol 178:413–427
Karimi SM, Freund M, Wager BM, Knoblauch M, Fromm J, M Mueller H, Ache P, Krischke M, Mueller MJ, Müller T, Dittrich M, Geilfus CM, Alfarhan AH, Hedrich R, Deeken R (2021) Under salt stress guard cells rewire ion transport and abscisic acid signaling. New Phytol 231:1040–1055
Li C, Shi L, Li X, Wang Y, Bi Y, Li W, Ma H, Chen B, Zhu L, Fu Y (2022) ECAP is a key negative regulator mediating different pathways to modulate salt stress-induced anthocyanin biosynthesis in Arabidopsis. New Phytol 233:2216–2231
Li H, He K, Zhang Z, Hu Y (2023) Molecular mechanism of phosphorous signaling inducing anthocyanin accumulation in Arabidopsis. Plant Physiol Biochem 196:121–129
Liu WC, Song RF, Zheng SQ, Li T-T, Zhang BL, Gao X, Lu YT (2022a) Coordination of plant growth and abiotic stress responses by tryptophan synthase β subunit1 through modulating tryptophan and ABA homeostasis in Arabidopsis. Mol Plant 15:1–18
Liu WC, Song RF, Qiu YM, Zheng SQ, Li TT, Wu Y, Song CP, Lu YT, Yuan HM (2022b) Sulfenylation of ENOLASE2 facilitates H2O2-conferred freezing tolerance in Arabidopsis. Dev Cell 57:1883–1898
Lu KK, Song RF, Guo JX, Zhang Y, Zuo JX, Chen HH, Liao CY, Hu XY, Ren F, Lu YT, Liu WC (2023) CycC1; 1-WRKY75 complex-mediated transcriptional regulation of SOS1 controls salt stress tolerance in Arabidopsis. Plant Cell 35:2570–2591
Malapeira J, Mas P (2014) ChIP-Seq analysis of histone modifications at the core of the Arabidopsis circadian clock. Methods Mol Biol 1158:57–69
Nagatoshi Y, Ikazaki K, Kobayashi Y, Mizuno N, Sugita R, Takebayashi Y, Kojima M, Sakakibara H, Kobayashi NI, Tanoi K, Fujii K, Baba J, Ogiso-Tanaka E, Ishimoto M, Yasui Y, Oya T, Fujita Y (2023) Phosphate starvation response precedes abscisic acid response under progressive mild drought in plants. Nat Commun 14:5047
Nakashima K, Fujita Y, Kanamori N, Katagiri T, Umezawa T, Kidokoro S, Maruyama K, Yoshida T, Ishiyama K, Kobayashi M, Shinozaki K, Yamaguchi-Shinozaki K (2009) Three Arabidopsis SnRK2 protein kinases, SRK2D/SnRK2.2, SRK2E/SnRK2.6/OST1 and SRK2I/SnRK2.3, involved in ABA signaling are essential for the control of seed development and dormancy. Plant & Cell Physiol 50(7):1345–1363. https://doi.org/10.1093/pcp/pcp083
Ni J, Premathilake AT, Gao Y, Yu W, Tao R, Teng Y, Bai S (2021) Ethylene-activated PpERF105 induces the expression of the repressor-type R2R3-MYB gene PpMYB140 to inhibit anthocyanin biosynthesis in red pear fruit. Plant J 105:167–181
Nilsson L, Muller R, Nielsen TH (2007) Increased expression of the MYB-related transcription factor, PHR1, leads to enhanced phosphate uptake in Arabidopsis thaliana. Plant Cell Environ 30:1499–1512
Peer WA, Bandyopadhyay A, Blakeslee JJ, Makam SN, Chen RJ, Masson PH, Murphy AS (2004) Variation in expression and protein localization of the PIN family of auxin efflux facilitator proteins in flavonoid mutants with altered auxin transport in Arabidopsis thaliana. Plant Cell 16:1898–1911
Raghavendra AS, Gonugunta VK, Christmann A, Grill E (2010) ABA perception and signalling. Trends Plant Sci 15:395–401
Ried MK, Wild R, Zhu JS, Pipercevic J, Sturm K, Broger L et al (2021) Inositol pyrophosphates promote the interaction of SPX domains with the coiled-coil motif of PHR transcription factors to regulate plant phosphate homeostasis. Nat Commun 12:384
Rubio V, Linhares F, Solano R, Martin AC, Iglesias J, Leyva A et al (2001) A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev 15:2122–2133
Shi HT, Liu GY, Wei YX, Chan ZL (2018) The zinc-finger transcription factor ZAT6 is essential for hydrogen peroxide induction of anthocyanin synthesis in Arabidopsis. Plant Mol Bio 97:165–176
Shin H, Shin HS, Dewbre GR, Harrison MJ (2004) Phosphate transport in Arabidopsis: Pht1;1 and Pht1;4 play a major role in phosphate acquisition from both low- and high-phosphate environments. Plant J 39:629–642
Song RF, Li TT, Liu WC (2021) Jasmonic acid impairs arabidopsis seedling salt stress tolerance through MYC2-mediated repression of CAT2 expression. Front Plant Sci 12:730228
Stracke R, De Vos RC, Bartelniewoehner L, Ishihara H, Sagasser M, Martens S, Weisshaar B (2009) Metabolomic and genetic analyses of flavonol synthesis in Arabidopsis thaliana support the in vivo involvement of leucoanthocyanidin dioxygenase. Planta 229:427–445
Su T, Xu Q, Zhang FC, Chen Y, Li LQ, Wu WH et al (2015) WRKY42 modulates phosphate homeostasis through regulating phosphate translocation and acquisition in Arabidopsis. Plant Physiol 167:1579–1717
Wan J, Zhang P, Wang R, Sun L, Wang W, Zhou H, Xu J (2018) UV-B radiation induces root bending through the flavonoid-mediated auxin pathway in arabidopsis. Frontiers Plant Sci 9:618
Wang Y, Li L, Ye T, Lu Y, Chen X, Wu Y (2013) The inhibitory effect of ABA on floral transition is mediated by ABI5 in Arabidopsis. J Exp Bot 64:675–684
Wang ZY, Ruan WY, Shi J, Zhang L, Xiang D, Yang C et al (2014) Rice SPX1 and SPX2 inhibit phosphate starvation responses through interacting with PHR2 in a phosphate-dependent manner. Proc Natl Acad Sci USA 111:14953–14958
Wang K, He J, Zhao Y, Wu T, Zhou X, Ding Y, Kong L, Wang X, Wang Y, Li J et al (2018) EAR1 negatively regulates ABA signaling by enhancing 2C protein phosphatase activity. Plant Cell 30:815–834
Wang X, Wang HF, Chen Y, Sun MM, Wang Y, Chen YF (2020) The transcription factor NIGT1.2 modulates both phosphate uptake and nitrate influx during phosphate starvation in Arabidopsis and maize. Plant Cell 32:3519–3534
Wang LF, Li TT, Zhang Y, Guo JX, Lu KK, Liu WC (2021) CAND2/PMTR1 is required for melatonin-conferred osmotic stress tolerance in Arabidopsis. Int J Mol Sci 22:4014
Wang LF, Lu KK, Li TT, Zhang Y, Guo JX, Song RF, Liu WC (2022) Maize PHYTOMELATONIN RECEPTOR1 functions in plant tolerance to osmotic and drought stress. J Exp Bot 73(7):5961–5973. https://doi.org/10.1093/jxb/erab553
Winkel-Shirley B (2001) Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol 126:485–493
Yan H, Pei X, Zhang H, Li X, Zhang X, Zhao M, Chiang VL, Sederoff RR, Zhao X (2021) MYB-Mediated regulation of anthocyanin biosynthesis. Int J Mol Sci 22:3103
Yoshida T, Fujita Y, Maruyama K, Mogami J, Todaka D, Shinozaki K, Yamaguchi-Shinozaki K (2015) Four Arabidopsis AREB/ABF transcription factors function predominantly in gene expression downstream of SnRK2 kinases in abscisic acid signalling in response to osmotic stress. Plant Cell Environ 38:35–49
Yu B, Zheng W, Xing L, Zhu JK, Persson S, Zhao Y (2022) Root twisting drives halotropism via stress-induced microtubule reorientation. Dev Cell 57:2412–2425
Zhang Y, Ji TT, Li TT, Tian YY, Wang LF, Liu WC (2020) Jasmonic acid promotes leaf senescence through MYC2-mediated repression of CATALASE2 expression in Arabidopsis. Plant Sci 299:110604
Zhang Y, Song RF, Yuan HM, Li TT, Wang LF, Lu KK, Guo JX, Liu WC (2021) Overexpressing the N-terminus of CATALASE2 enhances plant jasmonic acid biosynthesis and resistance to necrotrophic pathogen Botrytis cinerea B05.10. Mol Plant Pathol 22:1226–1238
Zhang Y, Wang LF, Han SY, Ren F, Liu WC (2022a) Sorting Nexin1 negatively modulates phosphate uptake by facilitating phosphate transporter1;1 degradation in Arabidopsis. Plant J 111:72–84
Zhang Y, Li TT, Wang LF, Guo JX, Lu KK, Song RF, Liu WC (2022b) Abscisic acid facilitates phosphate acquisition through the transcription factor aba insensitive5 in Arabidopsis. Plant J 111:269–281
Zhao HY, Nie KL, Zhou HP, Yan XJ, Zhan QD, Zheng Y et al (2020) ABI5 modulates seed germination via feedback regulation of the expression of the PYR/PYL/RCAR ABA receptor genes. New Phytol 228:596–608
Acknowledgements
We thank the anonymous reviewers for their thoughtful reviews and constructive suggestions which have improved this paper. The authors are grateful to Prof. Xia Li (Huazhong Agricultural University, China) for the abi2-1C mutant seed; Prof. Yingfang Zhu (Henan University, China) for the pyr1pyl12458 (hexpyl) mutant seed. This work was supported by National Natural Science Foundation of China (#32322010), Hainan Province Science and Technology Special Fund (ZDYF2022XDNY251), and National Natural Science Foundation of China (#32260400).
Funding
National Natural Science Foundation of China,32322010,Wen-Cheng Liu, 32260400, Hong-Mei Yuan, Hainan Province Science and Technology Special Fund, ZDYF2022XDNY251, Hong-Mei Yuan.
Author information
Authors and Affiliations
Contributions
H-MY and W-CL conceived and designed the research. R-FS and X-YH conducted the study. H-MY and W-CL analyzed the data and wrote the article.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest with this manuscript.
Additional information
Communicated by Sheng Ying.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Song, RF., Hu, XY., Liu, WC. et al. ABA functions in low phosphate-induced anthocyanin accumulation through the transcription factor ABI5 in Arabidopsis. Plant Cell Rep 43, 55 (2024). https://doi.org/10.1007/s00299-024-03146-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00299-024-03146-6