Abstract
Key message
VAMP726/VAMP725 and SYP131 can form a part of a SNARE complex to mediate vesicle secretion at the pollen tube apex.
Abstract
Secretory vesicle fusion with the plasma membrane of the pollen tube tip is a key step in pollen tube growth. Membrane fusion was mediated by SNAREs. However, little is known about the composition and function of the SNARE complex during pollen tube tip growth. In this study, we constructed a double mutant vamp725 vamp726 via CRISPR‒Cas9. Fluorescence labeling combined with microscopic observation, luciferase complementation imaging, co-immunoprecipitation and GST pull-down were applied in the study. We show that double mutation of the R-SNAREs VAMP726 and VAMP725 significantly inhibits pollen tube growth in Arabidopsis and slows vesicle exocytosis at the apex of the pollen tube. GFP-VAMP726 and VAMP725-GFP localize mainly to secretory vesicles and the plasma membrane at the apex of the pollen tube. In addition, fluorescence recovery after photobleaching (FRAP) experiments showed that mCherry-VAMP726 colocalizes with Qa-SNARE SYP131 in the central region of the pollen tube apical plasma membrane. Furthermore, we found that VAMP726 and VAMP725 can interact with the SYP131. Based on these results, we suggest that VAMP726/VAMP725 and SYP131 can form a part of a SNARE complex to mediate vesicle secretion at the pollen tube apex, and vesicle secretion may mainly occur at the central region of the pollen tube apical plasma membrane.







Similar content being viewed by others
Data availability
All relevant data can be found within the manuscript and its supporting materials.
References
Alexander MP (1969) Differential staining of aborted and nonaborted pollen. Stain Technol 44:117–122. https://doi.org/10.3109/10520296909063335
Baena G, Xia L, Waghmare S, Karnik R (2022) SNARE SYP132 mediates divergent traffic of plasma membrane H+-ATPase AHA1 and antimicrobial PR1 during bacterial pathogenesis. Plant Physiol 189:1639–1661. https://doi.org/10.1093/plphys/kiac149
Betz WJ, Mao F, Smith CB (1996) Imaging exocytosis and endocytosis. Curr Opin Neurobiol 6:365–371. https://doi.org/10.1016/s0959-4388(96)80121-8
Bhalla A, Chicka MC, Tucker WC, Chapman ER (2006) Ca2+-synaptotagmin directly regulates t-SNARE function during reconstituted membrane fusion. Nat Struct Mol Biol 13:323–330. https://doi.org/10.1038/nsmb1076
Bock JB, Matern HT, Peden AA, Scheller RH (2001) A genomic perspective on membrane compartment organization. Nature 409:839–841. https://doi.org/10.1038/35057024
Bolanos-Villegas P, Guo CL, Jauh GY (2015) Arabidopsis Qc-SNARE genes BET11 and BET12 are required for fertility and pollen tube elongation. Bot Stud. https://doi.org/10.1186/s40529-015-0102-x
Bove J, Vaillancourt B, Kroeger J, Hepler PK, Wiseman PW, Geitmann A (2008) Magnitude and direction of vesicle dynamics in growing pollen tubes using spatiotemporal image correlation spectroscopy and fluorescence recovery after photobleaching. Plant Physiol 147:1646–1658. https://doi.org/10.1104/pp.108.120212
Brunger AT, Choi UB, Lai Y, Leitz J, Zhou QJ (2018) Molecular mechanisms of fast neurotransmitter release. Annu Rev Biophys 47:469–497. https://doi.org/10.1146/annurev-biophys-070816-034117
Campanoni P, Blatt MR (2007) Membrane trafficking and polar growth in root hairs and pollen tubes. J Exp Bot 58:65–74. https://doi.org/10.1093/jxb/erl059
Chang F, Gu Y, Ma H, Yang ZB (2013) AtPRK2 promotes ROP1 activation via RopGEFs in the control of polarized pollen tube growth. Mol Plant 6:1187–1201. https://doi.org/10.1093/mp/sss103
Chen YA, Scheller RH (2001) SNARE-mediated membrane fusion. Nat Rev Mol Cell Biol 2:98–106. https://doi.org/10.1038/35052017
Chen HM, Zou Y, Shang YL, Lin HQ, Wang YJ, Cai R et al (2008) Firefly luciferase complementation imaging assay for protein-protein interactions in plants. Plant Physiol 146:368–376. https://doi.org/10.1104/pp.107.111740
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. https://doi.org/10.1046/j.1365-313x.1998.00343.x
Cui X, Wang S, Huang Y, Ding X, Wang Z, Zheng L et al (2022) Arabidopsis SYP121 acts as an ROP2 effector in the regulation of root hair tip growth. Mol Plant 15:1008–1023. https://doi.org/10.1016/j.molp.2022.04.008
Derksen J, Rutten T, Lichtscheidl IK, Dewin AHN, Pierson ES, Rongen G (1995) Quantitative-analysis of the distribution of organelles in tobacco pollen tubes: implications for exocytosis and endocytosis. Protoplasma 188:267–276. https://doi.org/10.1007/Bf01280379
Ebine K, Okatani Y, Uemura T, Goh T, Shoda K, Niihama M, Morita MT, Spitzer C, Otegui MS, Nakano A, Ueda T (2008) A SNARE complex unique to seed plants is required for protein storage vacuole biogenesis and seed development of Arabidopsis thaliana. Plant Cell 20:3006–3021. https://doi.org/10.1105/tpc.107.057711
El Kasmi F, Krause C, Hiller U, Stierhof YD, Mayer U, Conner L, Kong L, Reichardt I, Sanderfoot AA, Jurgens G (2013) SNARE complexes of different composition jointly mediate membrane fusion in Arabidopsis cytokinesis. Mol Biol Cell 24:1593–1601. https://doi.org/10.1091/mbc.E13-02-0074
Enami K, Ichikawa M, Uemura T, Kutsuna N, Hasezawa S, Nakagawa T et al (2009) Differential expression control and polarized distribution of plasma membrane-resident SYP1 SNAREs in Arabidopsis thaliana. Plant Cell Physiol 50:280–289. https://doi.org/10.1093/pcp/pcn197
Fasshauer D, Sutton RB, Brunger AT, Jahn R (1998) Conserved structural features of the synaptic fusion complex: SNARE proteins reclassified as Q- and R-SNAREs. Proc Natl Acad Sci U S A 95:15781–15786. https://doi.org/10.1073/pnas.95.26.15781
Fu Y, Wu G, Yang ZB (2001) Rop GTPase-dependent dynamics of tip-localized F-actin controls tip growth in pollen tubes. J Cell Biol 152:1019–1032. https://doi.org/10.1083/jcb.152.5.1019
Fujiwara M, Uemura T, Ebine K, Nishimori Y, Ueda T, Nakano A et al (2014) Interactomics of Qa-SNARE in Arabidopsis thaliana. Plant Cell Physiol 55:781–789. https://doi.org/10.1093/pcp/pcu038
Grebnev G, Cvitkovic M, Fritz C, Cai G, Smith AS, Kost B (2020) Quantitative structural organization of bulk apical membrane traffic in pollen tubes. Plant Physiol 183:1559–1585. https://doi.org/10.1104/pp.20.00380
Grefen C, Chen ZH, Honsbein A, Donald N, Hills A, Blatt MR (2010) A novel motif essential for SNARE interaction with the K+ channel KC1 and channel gating in Arabidopsis. Plant Cell 22:3076–3092. https://doi.org/10.1105/tpc.110.077768
Grefen C, Karnik R, Larson E, Lefoulon C, Wang YZ, Waghmare S, Zhang B, Hills A, Blatt MR (2015) A vesicle-trafficking protein commandeers Kv channel voltage sensors for voltage-dependent secretion. Nat Plants. https://doi.org/10.1038/nplants.2015.108
Gu X, Fonseka K, Agneessens J, Casson SA, Smertenko A, Guo G, Topping JF, Hussey PJ, Lindsey K (2021) The Arabidopsis R-SNARE VAMP714 is essential for polarisation of PIN proteins and auxin responses. New Phytol 230:550–566. https://doi.org/10.1111/nph.17205
Guo F, McCubbin AG (2012) The pollen-specific R-SNARE/longin PiVAMP726 mediates fusion of endo- and exocytic compartments in pollen tube tip growth. J Exp Bot 63:3083–3095. https://doi.org/10.1093/jxb/ers023
Guo JZ, Yang ZB (2020) Exocytosis and endocytosis: coordinating and fine-tuning the polar tip growth domain in pollen tubes. J Exp Bot 71:2428–2438. https://doi.org/10.1093/jxb/eraa134
Hachez C, Laloux T, Reinhardt H, Cavez D, Degand H, Grefen C, De Rycke R, Inze D, Blatt MR, Russinova E, Chaumont F (2014) Arabidopsis SNAREs SYP61 and SYP121 coordinate the trafficking of plasma membrane aquaporin PIP2;7 to modulate the cell membrane water permeability. Plant Cell 26:3132–3147. https://doi.org/10.1105/tpc.114.127159
He Y, Gao J, Luo M, GaoC LY, Wong HY et al (2022) VAMP724 and VAMP726 are involved in autophagosome formation in Arabidopsis thaliana. Autophagy 19:1406–1423. https://doi.org/10.1080/15548627.2022.2127240
Hepler PK, Winship LJ (2015) The pollen tube clear zone: clues to the mechanism of polarized growth. J Integr Plant Biol 57:79–92. https://doi.org/10.1111/jipb.12315
Hepler PK, Vidali L, Cheung AY (2001) Polarized cell growth in higher plants. Annu Rev Cell Dev Biol 17:159–187. https://doi.org/10.1146/annurev.cellbio.17.1.159
Higashiyama T, Takeuchi H (2015) The mechanism and key molecules involved in pollen tube guidance. Annu Rev Plant Biol 66:393–413. https://doi.org/10.1146/annurev-arplant-043014-115635
Hong W (2005) SNAREs and traffic. Biochim Biophys Acta 1744:493–517. https://doi.org/10.1016/j.bbamcr.2005.03.014
Ichikawa M, Iwano M, Sato MH (2015) Nuclear membrane localization during pollen development and apex-focused polarity establishment of SYP124/125 during pollen germination in Arabidopsis thaliana. Plant Reprod 28:143–151. https://doi.org/10.1007/s00497-015-0265-3
Ishiguro S, Kawai-Oda A, Ueda J, Nishida I, Okada K (2001) The DEFECTIVE IN ANTHER DEHISCENCE1 gene encodes a novel phospholipase A1 catalyzing the initial step of jasmonic acid biosynthesis which synchronizes pollen maturation anther dehiscence and flower opening in Arabidopsis. Plant Cell 13:2191–2209. https://doi.org/10.1105/tpc.13.10.2191
Jahn R, Scheller RH (2006) SNAREs–engines for membrane fusion. Nat Rev Mol Cell Biol 7:631–643. https://doi.org/10.1038/nrm2002
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907. https://doi.org/10.1002/j.1460-2075.1987.tb02730.x
Johnson MA, Harper JF, Palanivelu R (2019) A fruitful journey: pollen tube navigation from germination to fertilization. Annu Rev Plant Biol 70:809–837. https://doi.org/10.1146/annurev-arplant-050718-100133
Kapuscinski J (1995) DAPI: a DNA-specific fluorescent probe. Biotech Histochem 70:220–233. https://doi.org/10.3109/10520299509108199
Kato N, He HY, Steger AP (2010) A systems model of vesicle trafficking in Arabidopsis pollen tubes. Plant Physiol 152:590–601. https://doi.org/10.1104/pp.109.148700
Ketelaar T, Galway ME, Mulder BM, Emons AM (2008) Rates of exocytosis and endocytosis in Arabidopsis root hairs and pollen tubes. J Microsc 231:265–273. https://doi.org/10.1111/j.1365-2818.2008.02031.x
Kiessling V, Kreutzberger AJB, Liang B, Nyenhuis SB, Seelheim P, Castle JD, Cafiso DS, Tamm LK (2018) A molecular mechanism for calcium-mediated synaptotagmin-triggered exocytosis. Nat Struct Mol Biol 25:911–917. https://doi.org/10.1038/s41594-018-0130-9
Kim SJ, Brandizzi F (2012) News and views into the SNARE complexity in Arabidopsis. Front Plant Sci. https://doi.org/10.3389/fpls.2012.00028
Kost B (2008) Spatial control of Rho (Rac-Rop) signaling in tip growing plant cells. Trends Cell Biol 18:119–127. https://doi.org/10.1016/j.tcb.2008.01.003
Kwon C, Neu C, Pajonk S, Yun HS, Lipka U, Humphry M et al (2008) Co-option of a default secretory pathway for plant immune responses. Nature 451:835-U810. https://doi.org/10.1038/nature06545
Laloux T, Matyjaszczyk I, Beaudelot S, Hachez C, Chaumont F (2020) Interaction between the SNARE SYP121 and the plasma membrane aquaporin PIP2;7 involves different protein domains. Front Plant Sci 11:631643. https://doi.org/10.3389/fpls.2020.631643
Lancelle SA, Hepler PK (1992) Ultrastructure of freeze-substituted pollen tubes of lilium-longiflorum. Protoplasma 167:215–230. https://doi.org/10.1007/Bf01403385
Lee YJ, Szumlanski A, Nielsen E, Yang ZB (2008) Rho-GTPase-dependent filamentous actin dynamics coordinate vesicle targeting and exocytosis during tip growth. J Cell Biol 181:1155–1168. https://doi.org/10.1083/jcb.200801086
Lefoulon C, Waghmare S, Karnik R, Blatt MR (2018) Gating control and K+ uptake by the KAT1 K+ channel leaveraged through membrane anchoring of the trafficking protein SYP121. Plant Cell Environ 41:2668–2677. https://doi.org/10.1111/pce.13392
Lian N, Liu X, Wang X, Zhou Y, Li H, Li J, Mao T (2017) COP1 mediates dark-specific degradation of microtubule-associated protein WDL3 in regulating Arabidopsis hypocotyl elongation. Proc Natl Acad Sci U S A 114:12321–12326. https://doi.org/10.1073/pnas.1708087114
Lipka V, Kwon C, Panstruga R (2007) SNARE-Ware: the role of SNARE-domain proteins in plant biology. Annu Rev Cell Dev Biol 23:147–174. https://doi.org/10.1146/annurev.cellbio.23.090506.123529
Luo C, Shi Y, Xiang Y (2022) SNAREs regulate vesicle trafficking during root growth and development. Front Plant Sci 13:853251. https://doi.org/10.3389/fpls.2022.853251
Meng JG, Liang L, Jia PF, Wang YC, Li HJ, Yang WC (2020) Integration of ovular signals and exocytosis of a Ca(2+) channel by MLOs in pollen tube guidance. Nat Plants 6:143–153. https://doi.org/10.1038/s41477-020-0599-1
Nebenführ A, Ritzenthaler C, Robinson DG (2002) Brefeldin A: deciphering an enigmatic inhibitor of secretion. Plant Physiol 130:1102–1108. https://doi.org/10.1104/pp.011569
Pang L, Ma Z, Zhang X, Huang Y, Li R, Miao Y, Li R (2022) The small GTPase RABA2a recruits SNARE proteins to regulate the secretory pathway in parallel with the exocyst complex in Arabidopsis. Mol Plant 15:398–418. https://doi.org/10.1016/j.molp.2021.11.008
Park M, Krause C, Karnahl M, Reichardt I, El Kasmi F, Mayer U, Stierhof YD, Hiller U, Strompen G, Bayer M, Kientz M, Sato MH, Nishimura MT, Dangl JL, Sanderfoot AA, Jurgens G (2018) Concerted action of evolutionarily ancient and novel SNARE complexes in flowering-plant cytokinesis. Dev Cell 44:500-511.e504. https://doi.org/10.1016/j.devcel.2017.12.027
Parton RM, Fischer-Parton S, Watahiki MK, Trewavas AJ (2001) Dynamics of the apical vesicle accumulation and the rate of growth are related in individual pollen tubes. J Cell Sci 114:2685–2695. https://doi.org/10.1242/jcs.114.14.2685
Qin Y, Yang ZBA (2011) Rapid tip growth: Insights from pollen tubes. Semin Cell Dev Biol 22:816–824. https://doi.org/10.1016/j.semcdb.2011.06.004
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F (2013) Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8:2281–2308. https://doi.org/10.1038/nprot.2013.143
Reim K, Mansour M, Varoqueaux F, McMahon HT, Sudhof TC, Brose N, Rosenmund C (2001) Complexins regulate a late step in Ca2+-dependent neurotransmitter release. Cell 104:71–81. https://doi.org/10.1016/s0092-8674(01)00192-1
Rounds CM, Bezanilla M (2013) Growth mechanisms in tip-growing plant cells. Annu Rev Plant Biol 64:243–265. https://doi.org/10.1146/annurev-arplant-050312-120150
Ruan H, Li J, Wang T, Ren H (2020) Secretory vesicles targeted to plasma membrane during pollen germination and tube growth. Front Cell Dev Biol 8:615447. https://doi.org/10.3389/fcell.2020.615447
Rubiato HM, Liu M, O’Connell RJ, Nielsen ME (2022) Plant SYP12 syntaxins mediate an evolutionarily conserved general immunity to filamentous pathogens. Elife. https://doi.org/10.7554/eLife.73487
Saito C, Uedat T (2009) Functions of Rab and Snare Proteins in Plant Life. Int Rev Cel Mol Bio 274:183–233. https://doi.org/10.1016/S1937-6448(08)02004-2
Sanderfoot A (2007) Increases in the number of SNARE genes parallels the rise of multicellularity among the green plants. Plant Physiol 144:6–17. https://doi.org/10.1104/pp.106.092973
Sanderfoot AA, Kovaleva V, Bassham DC, Raikhel NV (2001) Interactions between syntaxins identify at least five SNARE complexes within the golgi/prevacuolar system of the arabidopsis cell. Mol Biol Cell 12:3733–3743. https://doi.org/10.1091/mbc.12.12.3733
Schmid M, Davison TS, Henz SR, Pape UJ, Demar M, VingronM SB, Weigel D, Lohmann JU (2005) A gene expression map of Arabidopsis thaliana development. Nat Genet 37:501–506. https://doi.org/10.1038/ng1543
Shorter J, Beard MB, Seemann J, Dirac-Svejstrup AB, Warren G (2002) Sequential tethering of Golgins and catalysis of SNAREpin assembly by the vesicle-tethering protein p115. J Cell Biol 157:45–62. https://doi.org/10.1083/jcb.200112127
Slane D, Reichardt I, El Kasmi F, Bayer M, Jurgens G (2017) Evolutionarily diverse SYP1 Qa-SNAREs jointly sustain pollen tube growth in Arabidopsis. Plant J 92:375–385. https://doi.org/10.1111/tpj.13659
Sudhof TC, Rothman JE (2009) Membrane fusion: grappling with SNARE and SM proteins. Science 323:474–477. https://doi.org/10.1126/science.1161748
Sutton RB, Fasshauer D, Jahn R, Brunger AT (1998) Crystal structure of a SNARE complex involved in synaptic exocytosis at 2.4 A resolution. Nature 395:347–353. https://doi.org/10.1038/26412
Tang Y, Yin ZN, Zeng YJ, Zhang QX, Chen LQ, He Y, Lu PL, Ye D, Zhang XQ (2017) MTOPVIB interacts with AtPRD1 and plays important roles in formation of meiotic DNA double-strand breaks in Arabidopsis. Sci Rep 7:10007. https://doi.org/10.1038/s41598-017-10270-9
Towbin H, Staehelin T, Gordon J (1979) Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets - procedure and some applications. Proc Natl Acad Sci U S A 76:4350–4354.
Uemura T, Ueda T, Ohniwa RL, Nakano A, Takeyasu K, Sato MH (2004) Systematic analysis of SNARE molecules in Arabidopsis: dissection of the post-Golgi network in plant cells. Cell Struct Funct 29:49–65. https://doi.org/10.1247/csf.29.49
Wang ZP, Xing HL, Dong L, Zhang HY, Han CY, Wang XC et al (2015) Egg cell-specific promoter-controlled CRISPR/Cas9 efficiently generates homozygous mutants for multiple target genes in Arabidopsis in a single generation. Genome Biol 16:144. https://doi.org/10.1186/s13059-015-0715-0
Xia LF, Marques-Bueno MM, Bruce CG, Karnik R (2019) Unusual roles of secretory SNARE SYP132 in plasma membrane H+-ATPase traffic and vegetative plant growth. Plant Physiol 180:837–858. https://doi.org/10.1104/pp.19.00266
Xue Y, Yang YQ, Yang ZJ, Wang XF, Guo Y (2018) VAMP711 is required for abscisic acid-mediated inhibition of plasma membrane H+-ATPase activity. Plant Physiol 178:1332–1343. https://doi.org/10.1104/pp.18.00499
Zonia L, Munnik T (2008) Vesicle trafficking dynamics and visualization of zones of exocytosis and endocytosis in tobacco pollen tubes. J Exp Bot 59:861–873. https://doi.org/10.1093/jxb/ern007
Zonia L, Munnik T (2009) Uncovering hidden treasures in pollen tube growth mechanics. Trends Plant Sci 14:318–327. https://doi.org/10.1016/j.tplants.2009.03.008
Zhang B, Karnik R, Wang YZ, Wallmeroth N, Blatt MR, Grefen C (2015) The Arabidopsis R-SNARE VAMP721 interacts with KAT1 and KC1 K+ channels to moderate K+ current at the plasma membrane. Plant Cell 27:1697–1717. https://doi.org/10.1105/tpc.15.00305
Zhang B, Karnik R, Waghmare S, Donald N, Blatt MR (2017) VAMP721 conformations unmask an extended motif for K+ channel binding and gating control. Plant Physiol 173:536–551. https://doi.org/10.1104/pp.16.01549
Zhang L, Ma J, Liu H, Yi Q, Wang Y, Xing J, Zhang P, Ji S, Li M, Li J, Shen J, Lin J (2021) SNARE proteins VAMP721 and VAMP722 mediate the post-Golgi trafficking required for auxin-mediated development in Arabidopsis. Plant J 108:426–440. https://doi.org/10.1111/tpj.15450
Acknowledgements
We thank the Arabidopsis Biological Resource Center at Ohio State University for providing the T-DNA insertion lines. We thank Dr. Zhenbiao Yang (University of California, Riverside) for the gift of the plasmid contained AtPRK1; Dr. Qijun Chen (China Agricultural University) for the gift of CRISPR-Cas9 vector; Dr. Yingzhang Li (China Agricultural University) for the gift of the plasmids pCAMBIA1300-nLUC-GUS and pCAMBIA1300-GUS-cLUC.
Funding
This study was supported by the National Natural Science Foundation of China (31270230 and 32270363).
Author information
Authors and Affiliations
Contributions
YL, XL and DZ conceived and designed the experiments. XL, DZ, FZ, YG, JL and YL performed the experiments. XL, DZ, FZ and YL analyzed the data. XL and YL wrote the paper. All authors contributed to the discussion of the results and edited and approved the submitted version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Communicated by Xian Sheng Zhang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
299_2023_3075_MOESM1_ESM.zip
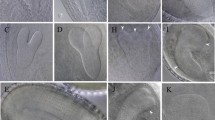
Supplementary file 1: Fig. S1. Promoter–GUS assay of VAMP726 expression in Arabidopsis. GUS histochemical activity assay showed that VAMP726 is abundantly expressed in Arabidopsis pollen and pollen tubes. A Seedling. B Inflorescence. C Flower. D Pistil. E Pollen. F Pollen tube. Fig. S2. Identification of the vamp726 mutant. A Schematic view of the genomic structure of VAMP726, and sites of the vamp726 T-DNA insertion. The solid boxes represent exons and the intervening lines represent introns. The location and orientation of T-DNA insertion are indicated. B Identification of T-DNA insertion homozygous of VAMP726 by PCR. C RT-PCR analysis of the vamp726 homozygous mutant line, indicated that there was no VAMP726 transcripts in the mutant. Actin8 was amplified as the positive internal control. D There was no significant difference in the length of pollen tubes of WT and vamp726 grown in vivo for 8 h, and the white arrow indicated the position of pollen tube growth. Bar = 500 μm. E Statistical analysis of the pollen tube relative length in vivo. To reduce the impact of individual differences, we calculated the relative length of the pollen tubes to the female tissues. 7 < n < 15, mean ± SD, P > 0.05 (Student’s t-test). At least 3 independent replicates were performed with consistent results. F WT and vamp726 pollen tubes grew in vitro for 5 h, n > 300, P > 0.05 (Student’s t-test). At least 3 independent replicates were performed with consistent results. Fig. S3. Identification of the vamp725 mutant. A CRISPR/Cas9-mediated gene editing knocked out VAMP725 in background of WT to generate a mutant vamp725. Sequence analysis showed 6-base deletion of AGAATT and a subsequent insertion of a base a occur at that target site, resulting in a frame shift and a stop codon at the 20th amino acid after the target site. B There was no significant difference in the length of pollen tubes of WT and vamp725 grown in vivo for 8 h, and the white arrow indicated the position of pollen tube growth. Bar = 500 μm. C Statistical analysis of the pollen tube relative length in vivo. To reduce the impact of individual differences, we calculated the relative length of the pollen tubes to the female tissues. 7 < n < 15, P > 0.05 (Student’s t-test). At least 3 independent replicates were performed with consistent results. D WT and vamp725 pollen tubes grew in vitro for 5 h, n > 300, P > 0.05 (Student’s t-test). At least 3 independent replicates were performed with consistent results. Fig. S4. Identification of the vamp725 vamp726 double mutant. A The sequence similarity of VAMP726 and VAMP725 is 69.82%. B CRISPR/Cas9-mediated gene editing knocked out VAMP725 in background of vamp726 to generate a double mutant vamp725 vamp726. Sequence analysis showed four bases AATT deletion at the target sequence. C RT-PCR analysis of the vamp725 vamp726 homozygous mutant line indicated that there was no VAMP726 transcripts in the double mutant. In the two homozygous lines which was complemented with GFP-VAMP726 driven by LAT52 could detect the expression of VAMP726. Actin8 was amplified as the positive internal control. D The pollen germination rates for 1 h, 2 h, 3 h, 4 h and 5 h respectively. This image showed that there was no significant difference in the germination rate between the WT and double mutant in vitro. n=5, P>0.05 (Student’s t-test). Fig. S5. Alexander staining for the mature pollen of vamp725 vamp726 mutant and WT. The pollen of WT and vamp725 vamp726 mutant were stained with Alexander, and they were uniformly red, indicating normal activity of the mutant pollen grains. Bar= 20 μm. Fig. S6. DAPI staining for the mature pollen of vamp725 vamp726 mutant and WT. DAPI staining showed that the mutant had normal karyotype during pollen development. These results indicated that pollen development was not affected obviously in the mutant. Bar= 20 μm. Fig. S7. The statistical analysis for Alexander and DAPI staining. (50 < n < 140), mean±SD, P > 0.05 (Student’s t-test). Fig. S8. The statistical analysis for the seed sets of the vamp725 vamp726 mutant and WT. (n=30), mean±SD, P > 0.05 (Student’s t-test). The average seed numbers of a mature silique in the vamp725 vamp726 mutant were 53.53 ±4.54. The average seed numbers of a mature silique in WT were 53.8 ±3.51. These results indicated that the seed sets was not obviously affected in the mutant. Fig. S9. VAMP725-GFP could rescue the phenotype of vamp725 vamp726 mutant. A The pollen tube growth patterns of WT, vamp725 vamp726 and two complemented lines which expressed VAMP725-GFP driven by LAT52 in vivo for 8 h. The pollen tube length of the mutant was short than that of WT and complemented lines. The result showed that VAMP725-GFP could rescue the phenotype of vamp725 vamp726. Bar = 500 μm. B Statistic analysis for the relative lengths of the pollen tube growth in vivo for 8 h. 10<n<15, **P < 0.01, Student’s t-test. C The subcellular localization of VAMP725-GFP in the pollen tubes of Arabidopsis. Top was the subcellular localization of VAMP725-GFP in WT and down was in vamp725 vamp726. VAMP725-GFP mainly localized on the inverted-cone-shaped region in the pollen tube tip and some also resided on the pollen tube plasma membrane. Bar= 5 μm. D Colocalization of VAMP725-GFP and FM4-64 in the pollen tube of Arabidopsis. The fluorescence distributions of VAMP725-GFP and FM4-64 staining were largely colocalization in the pollen tube tip. Bar = 10 μm. Fig. S10. The subcellular localization of GFP-VAMP726 in tobacco pollen tube. GFP-VAMP726 which was transformed through bombardment transformation mainly located on the inverted-cone-shaped region in the tobacco pollen tube tip. GFP-VAMP726 also resided on the apical plasma membrane of the tobacco pollen tube. Bar = 10 μm. Fig. S11. The localization of GFP-VAMP726 or mCherry-VAMP726 in lower expressing level in the pollen tubes. A The subcellular localization of GFP-VAMP726 driven by a native promoter of VAMP726 in the pollen tube of Arabidopsis. GFP-VAMP726 localized at the pollen tube tip and the apical plasma membrane. Bar=10 μm. B The subcellular localization of mCherry-VAMP726 in lower expressing level in the tobacco pollen tube. mCherry-VAMP726 also localized in the apical plasma membrane and pollen tube tip. Bar=5 μm. Fig. S12. Brefeldin A treatments resulted in GFP-VAMP726 to diffuse in pollen tube tip of tobacco. The growing tobacco pollen tube expressing GFP-VAMP726 was treated with 14 μM BFA, and observed by using a confocal microscope. After Brefeldin A treatments, the inverted-cone of GFP-VAMP726 in the pollen tube tip was gradually disappeared. The localization of GFP-VAMP726 was normal with pollen tube growth when methanol was used as the control. Bar = 10 μm. Fig. S13. VAMP726 and SYP131 colocalized at the apical plasma membrane in the tobacco pollen tube. A mCherry-VAMP726 mainly localized on the inverted-cone-shaped region and apical plasma membrane of the tobacco pollen tube. The SYP131-GFP wasresided on the whole plasma membrane of the tobacco pollen tube. Merged the two channels showed that mCherry-VAMP726 and SYP131-GFP colocalized at the apical plasma membrane of the pollen tube. Bar = 10 μm. B Quantitative analysis for the fluorescence intensity of mCherry-VAMP726 and SYP131-GFP along the white line in the pollen tube of tobacco. The image showed that both mCherry-VAMP726 and SYP131-GFP had a sharp fluorescence intensity increase on the apical plasma membrane of the pollen tube. The data indicated that mCherry-VAMP726 and SYP131-GFP colocalized at the apical plasma membrane of the tobacco pollen tube. Fig. S14. VAMP725 can interact with SYP131. A Firefly luciferase complementation assay showed that VAMP725 interacted with SYP131 in the tobacco leaf. nLUC+cLUC-SYP131 and nLUC +cLUC-VAMP725 were transformed as negative controls. SGT1-Nluc+Cluc-RAR1 were transformed as a positive control. The interaction between cLUC-VAMP725 and nLUC-SYP131 had an obvious fluorescent signal while there was no fluorescent signal in the negative controls. At least 10 tobacco leaves were collected and analyzed, which had similar results. B Co-IP assays showed the interaction between VAMP725 and SYP131. Protein extracted from the tobacco leaf which expressed Flag-SYP131+Myc-VAMP725 as the experimental group, and Flag+Myc-VAMP725 as control were subjected to immunoprecipitation with anti-Flag beads. Then Western blotting was performed with the anti-Myc and anti-Flag monoclonal antibodies. The results indicated that Flag-SYP131 could bind with Myc-VAMP725, no Myc-VAMP725 band was detected in the control group. The Flag-tag was too small and run out in the SDS-PAGE, so the Flag band was not shown in the results. The full-length blot is presented in Fig. S15. The experiments were repeated for three times with similar results. Fig. S15. The original immunoblotting images. A The original immunoblotting image of Figure 7C (Input, anti-Flag). B The original immunoblotting image of Figure 7C (Input, anti-Myc). C The original immunoblotting image of Figure 7C (IP, anti-Flag). D The original immunoblotting image of Figure 7C (IP, anti-Myc). E The original immunoblotting image of Figure 7D (Input and pull-down, anti-His). F The original immunoblotting image of Figure 7D (Input and pull-down, anti-GST). G The original immunoblotting image of Figure S14B (Input, anti-Flag). H The original immunoblotting image of Figure S14B (Input, anti-Myc). I The original immunoblotting image of Figure S14B (IP, anti-Flag). J The original immunoblotting image of Figure S14B (IP, anti-Myc).
Supplementary file 2: Video 1. FRAP of AtPRK1-GFP in the apical plasma membrane of Arabidopsis pollen tube.
Supplementary file 3: Video 2. FRAP of AtPRK1-GFP in the apical plasma membrane of vamp725 vamp726 pollen tube.
Supplementary file 4: Video 3. GFP-VAMP726 in the Arabidopsis growing pollen tube.
Supplementary file 5: Video 4. Brefeldin A treatments resulted in mCherry-VAMP726 to diffuse in the Arabidopsis pollen tube tip.
Supplementary file 6: Video 5. The localization of mCherry-VAMP726 was normal with pollen tube growth when methanol was used as the control.
Supplementary file 7: Video 6. Brefeldin A treatments resulted in GFP-VAMP726 to diffuse in the tobacco pollen tube tip.
Supplementary file 8: Video 7. The localization of GFP-VAMP726 in the tobacco pollen tube tip was normal with rapid pollen tube growth when methanol was used as the control.
Supplementary file 9: Video 8. FRAP assay showed that mCherry-VAMP726 first recovered at the central region of apical plasma membrane in the pollen tube.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, X., Zhu, D., Zhao, F. et al. VAMP726 and VAMP725 regulate vesicle secretion and pollen tube growth in Arabidopsis. Plant Cell Rep 42, 1951–1965 (2023). https://doi.org/10.1007/s00299-023-03075-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-023-03075-w