Abstract
Key message
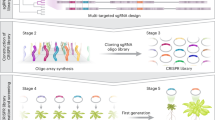
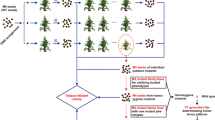
4382 available sgRNAs targeting 1060 tobacco genes were obtained, and 10,682 targeted mutants were created using high-throughput methods. Four optimization experiments were established to solve problems encountered during genetic transformation.

Similar content being viewed by others
Data availability
Data from this research that were also reviewed are included in the supplementary materials.
References
Bai M, Yuan J, Kuang H, Gong P, Li S, Zhang Z, Liu B, Sun J, Yang M, Yang L, Wang D, Song S, Guan Y (2020) Generation of a multiplex mutagenesis population via pooled CRISPR-Cas9 in soya bean. Plant Biotechnol J 18:721–731. https://doi.org/10.1111/pbi.13239
Chang Y, Long T, Wu C (2012) Effort and contribution of T-DNA Insertion mutant library for rice functional genomics research in China: review and perspective. J Integr Plant Biol 54:953–966. https://doi.org/10.1111/j.1744-7909.2012.01171.x
Jacobs TB, Zhang N, Patel D, Martin GB (2017) Generation of a collection of mutant tomato lines using pooled CRISPR libraries. Plant Physiol 174:2023–2037. https://doi.org/10.1104/pp.17.00489
Lu Y, Ye X, Guo R, Huang J, Wang W, Tang J, Tan L, Zhu JK, Chu C, Qian Y (2017) Genome-wide targeted mutagenesis in Rice using the CRISPR/Cas9 system. Mol Plant 10:1242–1245. https://doi.org/10.1016/j.molp.2017.06.007
Meng X, Yu H, Zhang Y, Zhuang F, Song X, Gao S, Gao C, Li J (2017) Construction of a genome-wide mutant library in Rice using CRISPR/Cas9. Mol Plant 10:1238–1241. https://doi.org/10.1016/j.molp.2017.06.006
Zhang J, Xing J, Mi Q, Yang W, Xiang H, Xu L, Zeng W, Wang J, Deng L, Jiang J, Yang G, Gao Q, Li X (2023) Highly efficient transgene-free genome editing in tobacco using an optimized CRISPR/Cas9 system, pOREU3TR. Plant Sci 326:111523. https://doi.org/10.1016/j.plantsci.2022.111523
Funding
This work was supported by [Creation and Identification of Editing Material Library of Tobacco Transporter and Enzyme-Related Genes in Red flower Mammoth Gold] (Grant number [110202101034] to H.X.) and [Development of Tagless Gene Editing Technology and Breeding Application of Low Nicotine Editing Materials] (Grant number [2022JY03] to H.X.).
Author information
Authors and Affiliations
Contributions
YC: Conceptualization, visualization, writing- original draft preparation, writing—review and editing. HX: Conceptualization, validation, funding acquisition, writing—original draft preparation, writing—review and editing. LJ: Investigation, writing—review and editing. QY: Writing—review and editing. JZ: Visualization, writing—review and editing. JJ: Investigation, writing—review and editing. WZ: Supervision, writing—review and editing. LD: Investigation. JJ: Investigation, writing—review and editing. QG: Conceptualization, validation, writing—review and editing. XL: Conceptualization, validation, supervision, writing—review and editing. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Communicated by Neal Stewart.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
299_2023_3050_MOESM1_ESM.rar
Supplementary file1 (RAR 207 kb) Supplementary material 1 Supplemental Materials and Methods. Supplementary material 2 sgRNA Sites and Sequences. Supplementary material 3 Statistical Data of NGS. Supplementary material 4 Results of Tissue Culture Optimization.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, Y., Xiang, H., Jia, L. et al. High-throughput creation of Nicotiana tabacum gene-targeted mutants based on CRISPR/Cas9. Plant Cell Rep 42, 2039–2042 (2023). https://doi.org/10.1007/s00299-023-03050-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-023-03050-5