Abstract
Key message
ZAT11, a Zinc Finger of Arabidopsis Thaliana 11, is a dual-function transcriptional regulator that positively regulates primary root growth but negatively regulates Ni 2+ tolerance.
Abstract
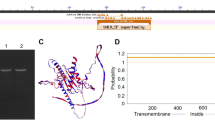
Zinc Finger of Arabidopsis Thaliana 11 (ZAT11) is a C2H2-type zinc finger protein that has been reported to function as an active transcriptional repressor. However, the biological function of ZAT11 remains unknown. Here we show that GFP-tagged ZAT11 is targeted to the nucleus. Analysis of plants expressing ZAT11 promoter-GUS showed that ZAT11 is highly expressed in roots and particularly in root tips. To identify the biological function of ZAT11, we constructed three independent lines of ZAT11 overexpressing transgenic plant (ZAT11 OE). ZAT11 OE enhanced the elongation of primary root but reduced the metal tolerance against nickel ion (Ni2+). The reduced Ni2+ tolerance of ZAT11 OE was correlated with decreased accumulation of Ni2+ in plants. The decreased accumulation of Ni2+ in ZAT11 OE was caused by the reduced transcription of a vacuolar Ni2+ transporter gene. Taken together, our results suggest that ZAT11 is a dual function transcriptional regulator that positively regulates primary root growth but negatively regulates Ni2+ tolerance.






Similar content being viewed by others
Abbreviations
- GFP:
-
Green fluorescent protein
- GSH:
-
Glutathione
- His:
-
Histidine
- NA:
-
Nicotianamine
- Ni2+ :
-
Nickel ion
References
Arazi T, Sunkar R, Kaplan B, Fromm H (1999) A tobacco plasma membrane calmodulin-binding transporter confers Ni2+ tolerance and Pb2+ hypersensitivity in transgenic plants. Plant J 20:171–182
Ciftci-Yilmaz S, Mittler R (2008) The zinc finger network of plants. Cell Mol Life Sci 65:1150–1160
Ciftci-Yilmaz S, Morsy MR, Song L, Coutu A, Krizek BA, Lewis MW, Warren D, Cushman J, Connolly EL, Mittler R (2007) The EAR-motif of the Cys2/His2-type zinc finger protein Zat7 plays a key role in the defense response of Arabidopsis to salinity stress. J Biol Chem 282:9260–9268
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Cobbett CS (2000) Phytochelatin biosynthesis and function in heavy-metal detoxification. Curr Opin Plant Biol 3:211–216
Devaiah BN, Nagarajan VK, Raghothama KG (2007) Phosphate homeostasis and root development in Arabidopsis are synchronized by the zinc finger transcription factor ZAT6. Plant Physiol 145:147–159
Gechev TS, Minkov IN, Hille J (2005) Hydrogen peroxide-induced cell death in Arabidopsis: transcriptional and mutant analysis reveals a role of an oxoglutarate-dependent dioxygenase gene in the cell death process. IUBMB Life 57:181–188
Hall JL (2002) Cellular mechanisms for heavy metal detoxification and tolerance. J Exp Bot 53:1–11
Han HJ, Park HC, Byun HJ, Lee SM, Kim HS, Yun DJ, Cho MJ, Chung WS (2012) The transcriptional repressor activity of ASYMMETRIC LEAVES1 is inhibited by direct interaction with calmodulin in Arabidopsis. Plant Cell Environ 35:1969–1982
Ingle RA, Mugford ST, Rees JD, Campbell MM, Smith JA (2005) Constitutively high expression of the histidine biosynthetic pathway contributes to nickel tolerance in hyperaccumulator plants. Plant Cell 17:2089–2106
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Kerkeb L, Kramer U (2003) The role of free histidine in xylem loading of nickel in Alyssum lesbiacum and Brassica juncea. Plant Physiol 131:716–724
Kim S, Takahashi M, Higuchi K, Tsunoda K, Nakanishi H, Yoshimura E, Mori S, Nishizawa NK (2005) Increased nicotianamine biosynthesis confers enhanced tolerance of high levels of metals, in particular nickel, to plants. Plant Cell Physiol 46:1809–1818
Krämer U, Cotter-Howells JD, Charnock JM, Baker AJM, Smith JAC (1996) Free histidine as a metal chelator in plants that accumulate nickel. Nature 379:635–638
Laity JH, Lee BM, Wright PE (2001) Zinc finger proteins: new insights into structural and functional diversity. Curr Opin Struct Biol 11:39–46
Lee SM, Kim HS, Han HJ, Moon BC, Kim CY, Harper JF, Chung WS (2007) Identification of a calmodulin-regulated autoinhibited Ca2+-ATPase (ACA11) that is localized to vacuole membranes in Arabidopsis. FEBS Lett 581:3943–3949
Liu XM, Kim KE, Kim KC, Nguyen XC, Han HJ, Jung MS, Kim HS, Kim SH, Park HC, Yun DJ, Chung WS (2010) Cadmium activates Arabidopsis MPK3 and MPK6 via accumulation of reactive oxygen species. Phytochemistry 71:614–618
Maksymiec W (2007) Signaling responses in plants to heavy metal stress. Acta Physiologiae Plantarum 29:177–187
Mittler R, Kim Y, Song L, Coutu J, Coutu A, Ciftci-Yilmaz S, Lee H, Stevenson B, Zhu JK (2006) Gain- and loss-of-function mutations in Zat10 enhance the tolerance of plants to abiotic stress. FEBS Lett 580:6537–6542
Nishida S, Tsuzuki C, Kato A, Aisu A, Yoshida J, Mizuno T (2011) AtIRT1, the primary iron uptake transporter in the root, mediates excess nickel accumulation in Arabidopsis thaliana. Plant Cell Physiol 52:1433–1442
Ohta M, Matsui K, Hiratsu K, Shinshi H, Ohme-Takagi M (2001) Repression domains of class II ERF transcriptional repressors share an essential motif for active repression. Plant Cell 13:1959–1968
Pandey N, Sharma CP (2002) Effect of heavy metals Co2+, Ni2+ and Cd2+ on growth and metabolism of cabbage. Plant Sci 163:753–758
Pianelli K, Mari S, Marques L, Lebrun M, Czernic P (2005) Nicotianamine over-accumulation confers resistance to nickel in Arabidopsis thaliana. Transgenic Res 14:739–748
Rizhsky L, Davletova S, Liang H, Mittler R (2004) The zinc finger protein Zat12 is required for cytosolic ascorbate peroxidase 1 expression during oxidative stress in Arabidopsis. J Biol Chem 279:11736–11743
Qureshi MK, Sujeeth N, Gechev TS, Hille J (2013) The zinc finger protein ZAT11 modulates paraquat-induced programmed cell death in Arabidopsis thaliana. Acta Physiol Plant 35:1863–1871
Sakamoto H, Maruyama K, Sakuma Y, Meshi T, Iwabuchi M, Shinozaki K, Yamaguchi-Shinozaki K (2004) Arabidopsis Cys2/His2-type zinc-finger proteins function as transcription repressors under drought, cold, and high-salinity stress conditions. Plant Physiol 136:2734–2746
Samarakoon AB, Rauser WE (1979) Carbohydrate levels and photoassimiilate export from leaves of phaseolus vulgaris exposed to excess cobalt, nickel, and zinc. Plant Physiol 63:1165–1169
Schaaf G, Honsbein A, Meda AR, Kirchner S, Wipf D, von Wiren N (2006) AtIREG2 encodes a tonoplast transport protein involved in iron-dependent nickel detoxification in Arabidopsis thaliana roots. J Biol Chem 281:25532–25540
Takatsuji H (1999) Zinc-finger proteins: the classical zinc finger emerges in contemporary plant science. Plant Mol Biol 39:1073–1078
Varagona MJ, Schmidt RJ, Raikhelai NV (1992) Nuclear localization signal(s) required for nuclear targeting of the maize regulatory protein opaque-2. Plant Cell 4:1213–1227
Wycisk K, Kim EJ, Schroeder JI, Kramer U (2004) Enhancing the first enzymatic step in the histidine biosynthesis pathway increases the free histidine pool and nickel tolerance in Arabidopsis thaliana. FEBS Lett 578:128–134
Acknowledgments
This work was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2013R1A1A2A10010567), and partly by a grant from the Next-Generation BioGreen 21 Program (#PJ00951401) funded by the Rural Development Administration, Republic of Korea. J. A. was supported by a scholarship from the BK21 plus program of the Ministry of Education in Korea.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Youn-Il Park.
X. -M. Liu and J. An contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, XM., An, J., Han, H.J. et al. ZAT11, a zinc finger transcription factor, is a negative regulator of nickel ion tolerance in Arabidopsis . Plant Cell Rep 33, 2015–2021 (2014). https://doi.org/10.1007/s00299-014-1675-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-014-1675-7