Abstract
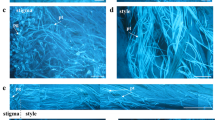
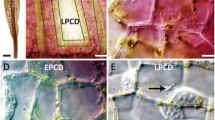
In the chestnut “replaceable bud” cultivar ‘Tima zhenzhu’, the auxiliary bud formed on the fruiting branch dies after fruiting, giving rise to a morphology more suitable than the wild type’s for intensive cultivation and heightened production. Here, we show that many of the hallmarks of programmed cell death (PCD) occur during the senescence of the replaceable bud, including DNA degradation, a high ratio of PCD cells and the breakdown of cell ultrastructure. The time course of the senescence was followed by sampling the developing bud from 20 to 40 days after flowering. In cv. ‘Tima zhenzhu’, DNA degradation was detectable prior to any visible sign of bud senescence, while it did not occur in the wild type (cv. ‘Dabanhong’). The ratio of PCD cells (as determined by flow cytometry) rose over the sampling period and was consistently higher in cv. ‘Tima zhenzhu’ than in cv. ‘Dabanhong’. After staining the bud cell nuclei with propidium iodide, it was clear that both their chromatin content and overall size fell over the sampling period in cv. ‘Tima zhenzhu’ while in cv. ‘Dabanhong’, no such decrease occurred. Other characteristics of PCD were noted in cv. ‘Tima zhenzhu’’s bud cells, including chromatin condensation, tonoplast invagination and DNA cleavage. We conclude that the replaceable bud senescence phenomenon is driven by PCD. The manipulation of this trait may have potential for remodeling the pattern of development of the fruit-bearing branches of chestnut.
Key message This paper first reported the occurrence of programmed cell death during the senescence of vegetative buds in a woody species, and the results extend the range of knowledge of PCD.





Similar content being viewed by others
References
Afanas’ev VN, Korol’ BA, Mantsygin YA, Nelipovich PA, Pechtnikov VA, Umansky SR (1986) Flow cytometry and biochemical analysis of DNA degradation characteristic of two types of cell death. Fed Exp Biol Sci Lett 194:347–350. doi:10.1016/0014-5793(86)80115-6
Belenghi B, Salomon M, Levine A (2004) Caspase-like activity in the seedlings of Pisum sativum eliminates weaker shoots during early vegetative development by induction of cell death. J Exp Bot 55:889–897. doi:101093/jxb/erh097
Bozhkov PV, Suarez MF, Filonova LH, Daniel G, Zamyatnin AA Jr, Rodriguez-Nieto S, Zhivotovsky B, Smertenko A (2005) Cysteine protease mcII-Pa executes programmed cell death during plant embryogenesis. Proc Natl Acad Sci USA 102:14463–14468. doi:10.1073_pnas0506948102
Darzynkiewicz Z, Bruno S, Del Bino G, Gorczyca W, Hots MA, Lassota P, Traganos F (1992) Features of apoptotic cells measured by flow cytometry. Cytometry 13:795–808. doi:10.1002/cyto.990130802
Donis-Gonzalez IR (2008) Microbial decay of fresh and peeled chestnuts and its control in Michigan. ProQuest, Ann Arbor, pp 9–12
Fukuda H (1996) Xylogenesis: initiation, progression, and cell death. Annu Rev Plant Physiol Plant Mol Biol 47:299–325. doi:10.1146/annurev.arplant.47.1.299
Gunawardena AHLAN (2008) Programmed cell death and tissue remodelling in plants. J Exp Bot 59:445–451. doi:10.1093/jxb/erm189
Gunawardena AHLAN, Greenwood JS, Dengler NG (2004) Programmed cell death remodels lace plant leaf shape during development. Plant Cell 16:60–73. doi:10.1105/tpc.016188
Johnson EO, Charchandi A, Babis GC, Soucacos PN (2008) Apoptosis in osteoarthritis: morphology, mechanisms, and potential means for therapeutic intervention. J Surg Orthop Adv 17:147–152
Liu QX, Kong DJ, Wang GP (2004) A new chestnut variety ‘Tima Zhenzhu’. Acta Hort Sin 31(5):698
Liu Y, Schiff M, Czymmek K, Talloczy Z, Levine B, Dinesh-Kumar SP (2005) Autophagy regulates programmed cell death during the plant innate immune response. Cell 121:567–577. doi:10.1016/j.cell.2005.03.007
Love AJ, Milner JJ, Sadanandom A (2008) Timing is everything: regulatory overlap in plant cell death. Trends Plant Sci 13:589–595. doi:10.1016/j.tplants.2008.08.006
Mishra S, Tyagi A, Dwivedi SP (2011) Regulation of apoptosis in living organisms: a biotechnological approach. Biotechnol Bioinf Bioeng 1(1):1–18
Pennell RI, Lamb C (1997) Programmed cell death in plants. Plant Cell 9:1157–1168
Reape TJ, Molony EM, McCabe PF (2008) Programmed cell death in plants: distinguishing between different modes. J Exp Bot 59(3):435–444. doi:10.1093/jxb/erm258
Tounekti O, Belehradek J Jr, Mir LM (1995) Relationships between DNA fragmentation, chromatin condensation, and changes in flow cytometry profiles detected during apoptosis. Exp Cell Res 217:506–516. doi:10.1006/excr.1995.1116
van Doorn WG (2011) Classes of programmed cell death in plants, compared to those in animals. J Exp Bot 62(14):4749–4761. doi:10.1093/jxb/err196
van Doorn WG, Woltering EJ (2004) Senescence and programmed cell death: substance or semantics? J Exp Bot 55:2147–2153. doi:10.1093/jxb/erh264
van Doorn WG, Woltering EJ (2005) Many ways to exit? Cell death categories in plants. Trends Plant Sci 10:117–122. doi:10.1016/j.tplants.2005.01.006
van Doorn WG, Beers EP, Dangl JL, Franklin-Tong VE, Gallois P, Hara-Nishimura I, Jones AM, Kawai-Yamada M, Lam E, Mundy J, Mur LAJ, Petersen M, Smertenko A, Taliansky M, Van Breusegem F, Wolpert T, Woltering E, Zhivotovsky B, Bozhkov PV (2011) Morphological classification of plant cell deaths. Cell Death Differ 18:1241–1246. doi:10.1038/cdd.2011.36
Walker PR, Sikorska M (1997) New aspects of the mechanism of DNA fragmentation in apoptosis. Biochem Cell Biol 75:287–299. doi:10.1139/o97-053
Walker NI, Harmon BV, Gobé GC, Kerr JFR (1988) Patterns of cell death. Methods Achiev Exp Pathol 13:18–54
Xiong HY, Li YS, Li LJ (2006) A unique form of cell death occurring in meristematic root tips of completely submerged maize seedlings. Plant Sci 171(5):624–631. doi:10.1016/j.plantsci.2006.06.007
Yamada T, Ichimura K, van Doorn WG (2006) DNA degradation and nuclear degeneration during programmed cell death in petals of antirrhinum, argyranthemum and petunia. J Exp Bot 57:3543–3552. doi:10.1093/jxb/erl100
Zhao GL, Zhang ZH, Sun HY, Li H, Dai HY (2007) Isolation of Ty1-copia-like retrotransposon sequences from the apple genome by chromosome walking based on modified SiteFinding-polymerase chain reaction. Acta Biochim Biophys Sin 39:675–683. doi:10.1111/j.1745-7270.2007.00328.x
Acknowledgments
This work was supported by National Natural Science Foundation of China (No. 30972394), Natural Science Foundation of Liaoning Province (No. 20102193) and Natural Science Foundation of Hebei Province (No. C2010001575).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Q. Zhao.
Rights and permissions
About this article
Cite this article
Wang, G., Zhang, Z., Kong, D. et al. Programmed cell death is responsible for replaceable bud senescence in chestnut (Castanea mollissima BL.). Plant Cell Rep 31, 1603–1610 (2012). https://doi.org/10.1007/s00299-012-1274-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-012-1274-4