Abstract
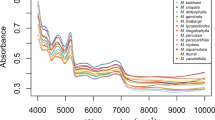
Pyrolysis mass spectrometry (PyMS) is a rapid, simple, high-resolution analytical method based on thermal degradation of complex material in a vacuum and has been widely applied to the discrimination of closely related microbial strains. Leaf samples of six species and one variety of higher plants (Rosa multiflora, R. multiflora var. platyphylla, Sedum kamtschaticum, S. takesimense, S. sarmentosum, Hepatica insularis, and H. asiatica) were subjected to PyMS for spectral fingerprinting. Principal component analysis of PyMS data was not able to discriminate these plants in discrete clusters. However, canonical variate analysis of PyMS data separated these plants from one another. A hierarchical dendrogram based on canonical variate analysis was in agreement with the known taxonomy of the plants at the variety level. These results indicate that PyMS is able to discriminate higher plants based on taxonomic classification at the family, genus, species, and variety level.



Similar content being viewed by others
Abbreviations
- ANNs :
-
Artificial neural networks
- CVA :
-
Canonical variate analysis
- GC/MS :
-
Gas chromatography/mass spectrometry
- PCA :
-
Principal component analysis
- PyMS :
-
Pyrolysis mass spectrometry
- UPGMA :
-
Unweighted pair group method with arithmetic mean
References
Fiehn O, Kopka J, Dormann P, Altmann T, Trethewey RN, Willmitzer L (2000) Metabolite profiling for plant functional genomics. Nat Biotechnol 18:1157–1161
Freeman R, Goodacre R, Sisson PR, Magee JG, Ward AC, Lightfoot NF (1994) Rapid identification of species within the Mycobacterium tuberculosis complex by artificial neural network analysis of PyMS data. J Med Microbiol 40:170–173
Goodacre R, Neal MJ, Kell DB, Greenham LW, Noble WC, Harvey RG (1994) Rapid identification using pyrolysis mass spectrometry and artificial neural networks of Propionibacterium acnes isolated from dogs. J Appl Bacteriol 76:124–134
Goodacre R, Hiom SJ, Cheeseman SL, Murdoch D, Weightman AJ, Wade WG (1996a) Identification and discrimination of oral asaccharolytic Eubacterium spp. by pyrolysis mass spectroscopy and artificial neural networks. Curr Microbiol 32:77–84
Goodacre R, Pygall J, Kell DB (1996b) Plant seed classification using pyrolysis mass spectrometry with unsupervised learning; the application of auto-associative and Kohonen artificial neural networks. Chemometr Intell Lab Sys 34:69–83
Irwin WJ (1982) Analytical pyrolysis: a comprehensive guide. Dekker, New York
Valcarce RV, Smith GG, Stevenson DN, Asay KH (1990) Chemometric analyses of Curie-point pyrolysis—mass spectral data for differentiating seeds of Triticeae grass species and hybrids. Chemometr Intell Lab Sys 9:95–105
Acknowledgements
This work was supported by a grant (no. M10104000234-01J000-10710) to JRL from the National Research Laboratory Program, a grant (no. BDM 0100211) to JRL from the Strategic National R&D Program through the Genetic Resources and Information Network Center, and a grant to JRL from the Korea Science and Engineering Foundation through the Plant Metabolism Research Center of the Kyung Hee University funded by the Korean Ministry of Science and Technology.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by I.S. Chung
Rights and permissions
About this article
Cite this article
Kim, S.W., Ban, S.H., Chung, H.J. et al. Taxonomic discrimination of higher plants by pyrolysis mass spectrometry. Plant Cell Rep 22, 519–522 (2004). https://doi.org/10.1007/s00299-003-0714-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-003-0714-6