Abstract
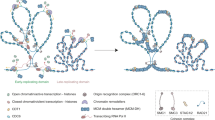
Multiple exogenous and endogenous genotoxic agents threaten the integrity of the genome, but one major source of spontaneous DNA damage is the formation of unscheduled DNA–RNA hybrids. These can be genetically detected by their ability to induce recombination. The origin of spontaneous hybrids has been mainly attributed to the nascent RNA formed co-transcriptionally in cis invading its own DNA template. However, it was unclear whether hybrids could also be spontaneously generated by RNA produced in a different locus (in trans). Using new genetic systems in the yeast Saccharomyces cerevisiae, we recently tested whether hybrids could be formed in trans and compromise genome integrity. Whereas we detected recombinogenic DNA–RNA hybrids in cis and in a Rad51-independent manner, we found no evidence for recombinogenic DNA–RNA hybrids to be formed with RNAs produced in trans. Here, we further discuss the implications in the field for the origin of genetic instability and the threats coming from RNAs.

Similar content being viewed by others
References
Aguilera A, Gomez-Gonzalez B (2017) DNA–RNA hybrids: the risks of DNA breakage during transcription. Nat Struct Mol Biol 24:439–443. https://doi.org/10.1038/nsmb.3395
Aguilera A, Klein HL (1988) Genetic control of intrachromosomal recombination in Saccharomyces cerevisiae. I. Isolation and genetic aracterization of hyper-recombination mutations. Genetics 119:779–790
Arab K, Karaulanov E, Musheev M, Trnka P, Schafer A, Grummt I, Niehrs C (2019) GADD45A binds R-loops and recruits TET1 to CpG island promoters. Nat Genet 51:217–223. https://doi.org/10.1038/s41588-018-0306-6
Ariel F, Lucero L, Christ A, Mammarella MF, Jegu T, Veluchamy A, Mariappan K, Latrasse D, Blein T, Liu C, Benhamed M, Crespi M (2020) R-loop mediated trans action of the APOLO long noncoding RNA. Mol Cell 77:1055–1065. https://doi.org/10.1016/j.molcel.2019.12.015
Britton S, Dernoncourt E, Delteil C, Froment C, Schiltz O, Salles B, Frit P, Calsou P (2014) DNA damage triggers SAF-A and RNA biogenesis factors exclusion from chromatin coupled to R-loops removal. Nucleic Acids Res 42:9047–9062. https://doi.org/10.1093/nar/gku601
Cloutier SC, Wang S, Ma WK, Al Husini N, Dhoondia Z, Ansari A, Pascuzzi PE, Tran EJ (2016) Regulated formation of lncRNA–DNA hybrids enables faster transcriptional induction and environmental adaptation. Mol Cell 61:393–404. https://doi.org/10.1016/j.molcel.2015.12.024
Cohen S, Puget N, Lin YL, Clouaire T, Aguirrebengoa M, Rocher V, Pasero P, Canitrot Y, Legube G (2018) Senataxin resolves RNA:DNA hybrids forming at DNA double-strand breaks to prevent translocations. Nat Commun 9:533. https://doi.org/10.1038/s41467-018-02894-w
Collins K (2000) Mammalian telomeres and telomerase. Curr Opin Cell Biol 12:378–383. https://doi.org/10.1016/s0955-0674(00)00103-4
Crossley MP, Bocek M, Cimprich KA (2019) R-loops as cellular regulators and genomic threats. Mol Cell 73:398–411. https://doi.org/10.1016/j.molcel.2019.01.024
D’Alessandro G, Whelan DR, Howard SM, Vitelli V, Renaudin X, Adamowicz M, Iannelli F, Jones-Weinert CW, Lee M, Matti V, Lee WTC, Morten MJ, Venkitaraman AR, Cejka P, Rothenberg E, d’Adda di Fagagna F (2018) BRCA2 controls DNA:RNA hybrid level at DSBs by mediating RNase H2 recruitment. Nat Commun 9:5376. https://doi.org/10.1038/s41467-018-07799-2
Duquette ML, Handa P, Vincent JA, Taylor AF, Maizels N (2004) Intracellular transcription of G-rich DNAs induces formation of G-loops, novel structures containing G4 DNA. Genes Dev 18:1618–1629. https://doi.org/10.1101/gad.1200804
Francia S, Michelini F, Saxena A, Tang D, de Hoon M, Anelli V, Mione M, Carninci P, d’Adda di Fagagna F (2012) Site-specific DICER and DROSHA RNA products control the DNA-damage response. Nature 488:231–235. https://doi.org/10.1038/nature11179
García-Muse T, Aguilera A (2019) R loops: from physiological to pathological roles. Cell 179:604–618. https://doi.org/10.1016/j.cell.2019.08.055
Huang D, Koshland D (2003) Chromosome integrity in Saccharomyces cerevisiae: the interplay of DNA replication initiation factors, elongation factors, and origins. Genes Dev 17:1741–1754. https://doi.org/10.1101/gad.1089203
Huertas P, Aguilera A (2003) Cotranscriptionally formed DNA:RNA hybrids mediate transcription elongation impairment and transcription-associated recombination. Mol Cell 12:711–721. https://doi.org/10.1016/j.molcel.2003.08.010
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–821. https://doi.org/10.1126/science.1225829
Kasahara M, Clikeman JA, Bates DB, Kogoma T (2000) RecA protein-dependent R-loop formation in vitro. Genes Dev 14:360–365
Keskin H, Shen Y, Huang F, Patel M, Yang T, Ashley K, Mazin AV, Storici F (2014) Transcript-RNA-templated DNA recombination and repair. Nature 515:436–439. https://doi.org/10.1038/nature13682
Lafuente-Barquero J, Garcia-Rubio ML, San Martin-Alonso M, Gómez-González B, Aguilera A (2020) Harmful DNA:RNA hybrids are formed in cis and in a Rad51-independent manner. eLife 9:e56674. https://doi.org/10.7554/eLife.56674
Li X, Manley JL (2005) Inactivation of the SR protein splicing factor ASF/SF2 results in genomic instability. Cell 122:365–378. https://doi.org/10.1016/j.cell.2005.06.008
Li L, Germain DR, Poon HY, Hildebrandt MR, Monckton EA, McDonald D, Hendzel MJ, Godbout R (2016) DEAD Box 1 facilitates removal of RNA and homologous recombination at DNA double-strand breaks. Mol Cell Biol 36:2794–2810. https://doi.org/10.1128/MCB.00415-16
Lu WT, Hawley BR, Skalka GL, Baldock RA, Smith EM, Bader AS, Malewicz M, Watts F, Wilczynska A, Bushell M (2018) Drosha drives the formation of DNA:RNA hybrids around DNA break sites to facilitate DNA repair. Nat Commun 9:532. https://doi.org/10.1038/s41467-018-02893-x
Luna R, Rondon AG, Perez-Calero C, Salas-Armenteros I, Aguilera A (2019) The THO complex as a paradigm for the prevention of cotranscriptional R-Loops. Cold Spring Harb Symp Quant Biol 84:105–114. https://doi.org/10.1101/sqb.2019.84.039594
Mizuta R, Iwai K, Shigeno M, Mizuta M, Uemura T, Ushiki T, Kitamura D (2003) Molecular visualization of immunoglobulin switch region RNA/DNA complex by atomic force microscope. J Biol Chem 278:4431–4434
Ohle C, Tesorero R, Schermann G, Dobrev N, Sinning I, Fischer T (2016) Transient RNA–DNA hybrids are required for efficient double-strand break repair. Cell 167:1001–1013. https://doi.org/10.1016/j.cell.2016.10.001
Pardo B, Gomez-Gonzalez B, Aguilera A (2009) DNA repair in mammalian cells: DNA double-strand break repair: how to fix a broken relationship. Cell Mol Life Sci 66:1039–1056. https://doi.org/10.1007/s00018-009-8740-3
Piazza A, Heyer WD (2019) Moving forward one step back at a time: reversibility during homologous recombination. Curr Genet 65:1333–1340. https://doi.org/10.1007/s00294-019-00995-7
Puget N, Miller KM, Legube G (2019) Non-canonical DNA/RNA structures during transcription-coupled double-strand break repair: roadblocks or bona fide repair intermediates? DNA Repair 81:102661. https://doi.org/10.1016/j.dnarep.2019.102661
Ray A, Machin N, Stahl FW (1989) A DNA double chain break stimulates triparental recombination in Saccharomyces cerevisiae. Proc Natl Acad Sci USA 86(16):6225–6229. https://doi.org/10.1073/pnas.86.16.6225
Reaban ME, Griffin JA (1990) Induction of RNA-stabilized DNA conformers by transcription of an immunoglobulin switch region. Nature 348:342–344. https://doi.org/10.1038/348342a0
Rich A (1960) A hybrid helix containing both deoxyribose and ribose polynucleotides and its relation to the transfer of information between the nucleic acids. Proc Natl Acad Sci USA 46:1044–1053
Rich A, Davies DR (1956) A new two-stranded helical structure: polyadenylic acid and polyuridylic acid. J Am Chem Soc 78:3548–3549. https://doi.org/10.1021/ja01595a086
Roy D, Zhang Z, Lu Z, Hsieh CL (2010) Lieber MR (2010) Competition between the RNA transcript and the nontemplate DNA strand during R-loop formation in vitro: a nick can serve as a strong R-loop initiation site. Mol Cell Biol 30:146–159. https://doi.org/10.1128/MCB.00897-09
Ruiz JF, Gómez-González B, Aguilera A (2009) Chromosomal translocations caused by either pol32-dependent or pol32-independent triparental break-induced replication. Mol Cell Biol 29:5441–5454. https://doi.org/10.1128/MCB.00256-09
Wahba L, Amon JD, Koshland D, Vuica-Ross M (2011) (2011) RNase H and multiple RNA biogenesis factors cooperate to prevent RNA:DNA hybrids from generating genome instability. Mol Cell 44:978–988. https://doi.org/10.1016/j.molcel.2011.10.017
Wahba L, Gore SK, Koshland D (2013) The homologous recombination machinery modulates the formation of RNA-DNA hybrids and associated chromosome instability. eLife 2:e00505. https://doi.org/10.7554/eLife.00505
Yasuhara T, Kato R, Hagiwara Y, Shiotani B, Yamauchi M, Nakada S, Shibata A, Miyagawa K (2018) Human Rad52 promotes XPG-mediated R-loop processing to initiate transcription-associated homologous recombination repair. Cell 175:558–570. https://doi.org/10.1016/j.cell.2018.08.056
Yu K, Chedin F, Hsieh CL, Wilson TE, Lieber MR (2003) R-loops at immunoglobulin class switch regions in the chromosomes of stimulated B cells. Nat Immunol 4:442–451. https://doi.org/10.1038/ni919
Zaitsev EN, Kowalczykowski SC (2000) A novel pairing process promoted by Escherichia coli RecA protein: inverse DNA and RNA strand exchange. Genes Dev 14:740–749
Zhang C, Chen L, Peng D, Jiang A, He Y, Zeng Y, Xie C, Zhou H, Luo X, Liu H, Chen L, Ren J, Wang W, Zhao Y (2020) METTL3 and N6-methyladenosine promote homologous recombination-mediated repair of DSBs by modulating DNA–RNA hybrid accumulation. Mol Cell 79:425–442. https://doi.org/10.1016/j.molcel.2020.06.017
Funding
This study was funded by Fundación Científica Asociación Española Contra el Cáncer (Dr. Belén Gómez-González, Grant No. AIO2015) and Ministerio de Ciencia e Innovación, Gobierno de España (Andrés Aguilera, Grant No. PID2019-104270GB-I00).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gómez-González, B., Aguilera, A. Origin matters: spontaneous DNA–RNA hybrids do not form in trans as a source of genome instability. Curr Genet 67, 93–97 (2021). https://doi.org/10.1007/s00294-020-01117-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-020-01117-4