Abstract
Eukaryotic cells have developed sophisticated systems to constantly monitor changes in the extracellular environment and to orchestrate a proper cellular response. To maximize survival, cells delay cell-cycle progression in response to environmental changes. In response to extracellular insults, stress-activated protein kinases (SAPKs) modulate cell-cycle progression and gene expression. In yeast, osmostress induces activation of the p38-related SAPK Hog1, which plays a key role in reprogramming gene expression upon osmostress. Genomic analysis has revealed the existence of a large number of long non-coding RNAs (lncRNAs) with different functions in a variety of organisms, including yeast. Upon osmostress, hundreds of lncRNAs are induced by the SAPK p38/Hog1. One gene that expresses Hog1-dependent lncRNA in an antisense orientation is the CDC28 gene, which encodes CDK1 kinase that controls the cell cycle in yeast. Cdc28 lncRNA mediates the induction of CDC28 expression and this increase in the level of Cdc28 results in more efficient re-entry of the cells into the cell cycle after stress. Thus, the control of lncRNA expression as a new mechanism for the regulation of cell-cycle progression opens new avenues to understand how stress adaptation can be accomplished in response to changing environments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Cell-cycle progression upon osmostress
Role of Cdc28 in cell-cycle control: CDK1 activity and association with cyclins
All cell-cycle events in budding yeast are biochemically coordinated, mostly by a single cyclin-dependent kinase (CDK), Cdc28 (Hartwell et al. 1973). Cdc28 activity and specificity are tightly controlled in a time-dependent manner through the regulation of different phase-specific cyclins, phosphorylations and inhibitors. The combination of multiple oscillatory mechanisms that collaborate in providing alternate periods of low and high levels of the different cyclin-CDK activities ensures orderly progression through the cell cycle (Spellman et al. 1998); (Orlando et al. 2008). In S. cerevisiae, Cdc28 cyclins have historically been classified into two broad groups: G1 cyclins (Cln1–3) that regulate events during the interval between mitosis and DNA replication and B-type cyclins (Clb1–6) that are required for replication, G2 progression and passage through Mitosis (Mendenhall and Hodge 1998). All cyclins except for Cln3 experience waves of production and destruction in pairs. Although cyclins show a high level of functional redundancy, the specificity of each cyclin is assured by a variety of mechanisms (Bloom and Cross 2007). Cyclins show different sensitivity to cell-cycle-regulated inhibitors, are restricted to different subcellular locations and are controlled in a time-dependent manner by specific transcriptional activators. Moreover, cyclins are degraded by phase-specific events, bind only to specific targets and facilitate different Cdc28 inhibitory phosphorylations when bound to it. Whereas the association of Cdc28 with cyclins and cyclin-dependent kinase inhibitor (CDKi) is a key regulatory element of Cdc28 kinase activity; the levels of Cdc28 protein do not change significantly during cell-cycle progression (Spellman et al. 1998).
Transcriptional control of the cell cycle
An astonishing amount of data has been generated over decades, which were aimed at understanding the complex regulatory network that governs cell-cycle progression. In the early nineties, mRNAs whose levels oscillated as yeast cells progressed thorough the cell cycle were identified (Price et al. 1991). Subsequently, the dynamics of cell cycle driven transcriptional waves were assessed using microarrays and these analyses significantly extended the number of known cell-cycle regulated genes to up to 800 genes (Cho et al. 1998; Spellman et al. 1998). Clustering of periodic transcripts led to the identification of cell-cycle regulated gene groups involved in cell-cycle control, budding, mating, DNA replication and repair. Cyclins are prototypical genes that display and determine induction of such periodic waves of transcription. Later studies devoted their efforts to understand the timing of these transcriptional oscillations by analyzing the genome-wide targets of the transcription factors that dictate cyclic specific expression (Lee et al. 2002; Simon et al. 2001). Consequently, a fully connected transcriptional circuit was delineated (Gauthier et al. 2008).
Control of cell-cycle progression by the Hog1 SAPK
Stress-activated protein kinases (SAPKs) modulate cell-cycle progression and gene expression in response to extracellular insults. In yeast, osmostress induces activation of the p38-related SAPK Hog1 which leads to cell-cycle delay in different phases of the cell cycle (Saito and Posas 2012). Exposure to stress leads to a rapid but transient cell-cycle delay that depends on the strength of the stress and the degree of SAPK activation (Adrover et al. 2011). Long-term activation of Hog1 leads to a sustained cell-cycle arrest that, when maintained, provokes cells to enter into apoptosis (Vendrell et al. 2011). Hog1 controls cell-cycle progression in the G1, S and G2/M phases of the cell cycle, clearly indicating that, upon stress, cells need to transiently arrest cell-cycle progression for proper adaptation. Indeed, cells unable to arrest the cell cycle in the presence of stress become osmosensitive (Clotet et al. 2006; Duch et al. 2013; Escote et al. 2004).
Phosphorylation of core components of the cell-cycle machinery by Hog1
Activation of Hog1 leads to regulation of different key components of the cell-cycle machinery (Fig. 1). In G1, Hog1 directly phosphorylates the Sic1 CDKi promoting its stability. Hog1-phosphorylated Sic1 inhibits Cdc28–Clb5/6 complexes and prevents replication (Adrover et al. 2011; Escote et al. 2004). The lack of SIC1 leads to genomic instability when cells are stressed in G1. During S phase, Hog1 phosphorylates Mrc1 a component of the replication complex that links the helicase with the DNA polymerase. Phosphorylation of Mrc1 leads to inhibition of origin firing as well as lowers down replication fork progression. Therefore, by delaying replication, Hog1 plays a key role in preventing conflicts between RNA and DNA polymerases. Hence, Mrc1 is a key protein for integration of signals from the DNA-damage checkpoint as well as of environmental signals mediated by Hog1 (Duch et al. 2012, 2013). In addition, during G2/M, Hog1 targets Hsl1 to delay cell-cycle progression via the inhibitory phosphorylation of Cdc28 by Swe1 (Clotet et al. 2006; Duch et al. 2012). The combined data suggest that, in the presence of stress, Hog1 impacts on basic components of the cell-cycle machinery to delay the cell cycle and to permit proper generation of adaptive responses before progression into the next phase of the cell cycle.
Control of cell-cycle progression by Hog1. The cyclin-dependent kinase (CDK) Cdc28 associates with phase-specific cyclins (shown around the central circle) to regulate passage through the cell cycle (G1 → S → G2 → M). Upon stress, Hog1 modulates progression at all phases of the cell cycle by acting on core elements of the cell cycle machinery
Regulation of CDK activity by the down-regulation of cyclin expression by Hog1
In addition to the direct phosphorylation of several components of the cell-cycle core machinery, Hog1 also regulates cell-cycle transitions by down-regulation of cyclin expression. In G1, activation of Hog1 prevents induction of CLN1, CLN2 and CLB5 cyclin genes (Adrover et al. 2011; Escote et al. 2004). Albeit the mechanism by which Hog1 represses expression from promoters regulated by G1 specific transcription factors, MBF and SBF, remains to be elucidated, mathematical modeling supported by quantitative in vivo experiments showed that CLB5 down-regulation is a key regulator in G1 arrest whereas CLN1,2 down-regulation or stabilization of Sic1 seems to be important only during late G1, when Hog1 control over Clb5 is not sufficiently tight to prevent S phase entry (Adrover et al. 2011). Therefore, the complex and strict Hog1 control over the G1/S network clearly illustrates the necessity of cell adaptation to osmostress prior to re-entering the cell cycle. Similarly, activation of Hog1 in G2 leads to down-regulation of the CLB2 cyclin gene (Alexander et al. 2001; Clotet et al. 2006). The synergistic regulation by Hog1 of core elements of the cell cycle by direct phosphorylation, together with Hog1 down-regulation of cyclin expression, might serve to efficiently block cell-cycle progression in response to stress.
Control of transcription in response to stress
Stress-activated protein kinases regulate gene expression to maximize cellular adaptation to environmental stresses (de Nadal et al. 2011; Weake and Workman 2010). Exposure to high osmolarity causes a sudden change in water activity and ionic strength, which impacts protein interactions. Most DNA binding proteins rapidly and transiently disassociate from chromatin upon osmostress (Proft and Struhl 2004). Moreover, a transcriptional burst of osmoresponsive genes occurs within minutes after stress. Transcriptional reprogramming is not essential for the short-term adaptive response but is crucial for long-term adaptation since mutants that display impaired transcription are unable to grow under osmostress (de Nadal et al. 2004; Mas et al. 2009; Zapater et al. 2007). Osmostress regulated genes have been implicated in carbohydrate metabolism, protein biosynthesis, general stress protection and signal transduction. Hog1 plays a key role in transcriptional reprogramming (Causton et al. 2001; Gasch et al. 2000; Pokholok et al. 2006; Posas et al. 2000; Rep et al. 2000) and coordinates the induction of osmoresponsive genes by controlling the entire process of mRNA biogenesis (de Nadal et al. 2011) (Fig. 2).
Transcriptional reprogramming by Hog1 upon stress
The best characterized mechanism by which Hog1 controls transcription initiation is by direct phosphorylation of promoter-specific transcription factors (de Nadal et al. 2003; Proft and Struhl 2002; Ruiz-Roig et al. 2012). Additionally, Hog1 itself binds to chromatin through physical interaction with the transcription factors that serve as an anchoring platform for Hog1 (Alepuz et al. 2001; Pascual-Ahuir et al. 2006; Pokholok et al. 2006; Proft et al. 2006). Binding of Hog1 stimulates recruitment of the basic transcription machinery; Mediator, SAGA, SWI/SNF, Rpd3 deacetylase and Ubp3 deubiquitinase to promote transcription (Alepuz et al. 2003; de Nadal et al. 2004; Guha et al. 2007; Kobayashi et al. 2008; Proft and Struhl 2002; Sole et al. 2011; Zapater et al. 2007).
Hog1 is recruited not only to promoters but also to coding regions where it travels with elongating RNA Pol II and serves as a selective elongation factor for stress-responsive genes (Pokholok et al. 2006; Proft et al. 2006). Strong induction of gene expression in response to stress requires major changes in the chromatin structure of osmoresponsive genes (Nadal-Ribelles et al. 2012). To achieve efficient nucleosome eviction, Hog1 targets the RSC complex to code regions by directly binding to it (Mas et al. 2009). Repositioning of nucleosomes at stress-responsive genes is mediated by the INO80 complex (Klopf et al. 2009). Analysis of the signaling and transcription dynamics of the HOG pathway in single cells demonstrated that Hog1 activation increases linearly with osmostress, while transcriptional output exhibits a bimodal behavior. The cause for this stochastic expression is determined by the state and structure of chromatin (Neuert et al. 2013; Pelet et al. 2011; Zechner et al. 2012).
mRNA processing, stability and export of stress-responsive nascent transcripts are also regulated by Hog1. Following osmostress, the synthesis and half-life of osmoresponsive mRNAs increase while a broad range of other RNAs is concurrently destabilized (Miller et al. 2011; Molin et al. 2009; Romero-Santacreu et al. 2009). Moreover, in response to osmostress, Hog1 phosphorylates components of the inner nuclear basket (Nup1, Nup2 and Nup60), which associates with osmoresponsive promoters in a Hog1 activity-dependent manner (Regot et al. 2013).
Genome-wide picture of the osmostress transcriptome
Although single case studies have provided very useful data, genome-wide approaches have provided a broader picture of the role of Hog1 as a master regulator of the massive transcription reprogramming that occurs in response to osmostress. DNA microarray studies have shown that 5–7 % of the protein coding genes show significant changes in their expression levels after mild osmotic shock. Depending on the severity of the stress, induction of up to 80 % of the induced transcripts depends on Hog1 (Capaldi et al. 2008; Causton et al. 2001; Gasch et al. 2000; O’Rourke and Herskowitz 2004; Posas et al. 2000; Rep et al. 2000). Other systematic analyses have been carried out to characterize the contribution of each stress-responsive transcription factor controlled by Hog1 to the total transcriptional response. Transcription profiling of individual or multiple mutants of transcription factors used by Hog1 shed light on the complexity of the transcription factor network (Capaldi et al. 2008; Ni et al. 2009). Further studies using high density oligonucleotide or tiling arrays (see below) complemented and deepened our knowledge of the genome-wide picture of the osmostress transcriptome.
Transcription rate in response to osmostress has been measured using two different techniques: genomic run-on (GRO) and dynamic transcription analysis (DTA) (Miller et al. 2011; Romero-Santacreu et al. 2009). Both types of studies identified changes in mRNA synthesis and decay in response to osmostress and identified three phases of the stress response: shock, induction and recovery phases (Miller et al. 2011). During the initial shock, synthesis and decay rates globally decrease causing storage of mRNA in P-bodies (Romero-Santacreu et al. 2009). Later, in the induction stage, the mRNA synthesis rates of osmoresponsive genes increase together with mRNA decay rates to ensure high production and removal of stress mRNAs. In the subsequent recovery phase, mRNA decay and synthesis are restored to prestimulation levels (Miller et al. 2011).
Genome-wide localization of Hog1 and stress-responsive transcription factors was assessed by combining chromatin immunoprecipitation with microarray technology (ChIP on chip) (Capaldi et al. 2008; Proft et al. 2006). Moreover, an attempt was made to determine the role of Hog1 in the genome-wide distribution of RNA Pol II using ChIP followed by deep sequencing (ChIP-seq) analysis (Cook and O’Shea 2012; Nadal-Ribelles et al. 2012). In response to osmostress, there is a genome-wide tendency to lose association of RNA Pol II with the genome, which leads the entire genome into a repressive state. On the other hand, while the entire genome is undergoing RNA Pol II dissociation, Hog1 binds through specific transcription factors to selectively target RNA Pol II to induce stress-responsive genes. Thus, colocalization of Hog1 and RNA Pol II bypasses the osmostress-induced global transcription repression. These data are consistent with single case studies. Osmoresponsive transcription is characterized by strong induction of gene expression from almost no expression under basal conditions to maximal activation (fold changes ranging from 5 to 200) in just 10 min (Posas et al. 2000). This process involves major changes in chromatin structure, which, for osmoresponsive genes, require the presence of Hog1 (Mas et al. 2009). Genome-wide changes in chromatin structure in response to osmostress were determined using micrococcal nuclease followed by deep sequencing (MNase-seq). Hog1-dependent genes displayed massive nucleosome eviction at promoter and coding regions in response to stress that was completely dependent on the presence of Hog1 (Nadal-Ribelles et al. 2012).
The global snapshot of gene expression, protein localization and nucleosome occupancy that was thus obtained revealed a dose-dependent correlation between chromatin remodeling and both the strength and the residence time of Hog1 at target genes, making transcription of these genes more efficient. In other words, Hog1 bypasses stress-mediated down-regulation of transcription via its effects on RNA polymerase II redistribution and chromatin remodeling.
Induction of a new set of lncRNAs in response to environmental insults
Pervasive transcription and lncRNAs
Expression profiling using microarrays has been highly efficient in obtaining quantitative measurements of total mRNA (Young 2000). However, this method cannot fully unravel the complexity of the transcriptome. To overcome these limitations, several new methods with increased sensitivity and coverage have been developed (i.e. tiling arrays, RNA-seq, GRO-seq, NET-seq and TIF-seq). These unbiased strategies interrogate both strands of the genome allowing the detection of new transcripts such as non-coding RNAs (ncRNAs) and complex transcriptional architectures (Bertone et al. 2004; Nagalakshmi et al. 2008; Garcia-Martinez et al. 2004; Churchman and Weissman 2011; Pelechano et al. 2013). Of this newly described plethora of RNA species, the best characterized in structure and function are the short ncRNAs (<200 nt in length) that encompass several classes such as: small interfering RNAs (siRNAs), micro RNAs (micRNAs) and PIWI-interacting RNAs (piRNAs). The role of short ncRNAs as powerful regulators of gene expression is well accepted and established in complex organisms reviewed in Moazed (2009). On the other hand, long noncoding RNAs (lncRNAs, >200 nt in length) have also been identified in virtually all studied organisms regardless of genome size and complexity. However, our understanding of their properties is frequently descriptive (i.e. size, stability and genomic location), and in most cases their function has not been assessed (Pelechano and Steinmetz 2013).
In S. cerevisiae as much as 85 % of the genome is expressed under basal conditions, including a large number of lncRNA transcripts (David et al. 2006). Further studies have identified and characterized new lncRNA families based on their stability. Thus, cryptic unstable transcripts (CUTs) are noncoding RNAs that are only expressed in mutants of the nuclear exosome (rrp6), Xrn1-sensitive unstable transcripts (XUTs) are noncoding transcripts that are expressed in cytosolic exosome mutants (xrn1), and stable unannotated transcripts (SUTs) are a group of stable non-coding RNAs (van Dijk et al. 2011; Xu et al. 2009). Recently, other groups reported the appearance of novel ncRNA transcripts in the deacetylase Set3 deletion strain (Kim et al. 2012) and in the absence of gene looping (in a ssu72 mutant) that have been named Ssu72-restricted transcripts (SRTs) (Tan-Wong et al. 2012).
Change of the noncoding transcriptome upon environmental changes
The transcription of lncRNAs was initially interpreted as transcriptional noise or as a by-product. However, several studies in yeast have started to shed some light on the complexity of the stress-responsive transcriptome under several conditions such as growth in different fermentable carbon sources (Xu et al. 2009), diauxic shift (Kim et al. 2012) and in response to oxidative stress in fission yeast (Quintales et al. 2010). Therefore, adaptive stress responses also require regulation of the noncoding transcriptome. Upon osmostress, Hog1 induces the expression of a complete new set of lncRNAs (Nadal-Ribelles et al. 2014).
Cdc28 is regulated by a stress-induced lncRNA
A new set of lncRNAs is induced upon stress
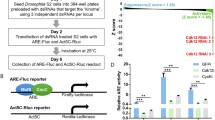
The use of tiling arrays has shown that in addition to coding genes, around 200 lncRNAs are strongly and rapidly induced upon osmostress, of which 91 are Hog1 dependent. The promoters of these stress-responsive lncRNAs show similarities with those of stress-responsive genes. Hog1 associates with these promoters and stimulates the recruitment of RNA Pol II (Nadal-Ribelles et al. 2014). Functional characterization of these stress-induced lncRNAs showed that there is a significant number of genes that display a positive correlation between expression of the sense and antisense-induced (from 8 to 41, depending on the analysis method). One of these genes that shows a clear correlation between the sense mRNA and antisense lncRNA expression is CDC28, which encodes the main CDK that drives progression of the cell cycle in yeast.
An lncRNA up regulates Cdc28 protein levels to promote cell-cycle re-entry in the presence of osmostress
Genetic and biochemical characterization of the induction of CDC28 gene expression by Hog1-induced antisense lncRNA led to the following tentative mechanistic model. Upon osmostress, Hog1 associates with the 3′ UTR region of CDC28, via its association with Sko1 (unpublished data), and induces lncRNA transcription. This induction of lncRNA permits the establishment of gene looping mediated by Ssu72. Gene looping leads to the transfer of Hog1 to the +1 nucleosome region of CDC28. Association of Hog1 with the proximal region of the gene serves to target the RSC chromatin remodeler, which remodels the +1 region, thus permitting an increase in transcription of the CDC28 gene (Fig. 3).
Schematic representation of the two-step model of lncRNA and gene looping mediated gene regulation. Hog1-mediated regulation of sense (black arrow) and antisense lncRNA (red arrow) transcripts following osmostress. Top panel Stress induces Hog1-mediated recruitment of transcriptional machinery to the 3′UTR region of an osmostress regulated gene. Bottom panel The resulting induced lncRNA, in the presence of Ssu72, causes looping of the DNA and translocation of Hog1 and RSC to the +1 nucleosome region, resulting in enhanced sense transcription in addition to antisense transcripts. Light colored nucleosomes indicate remodeled nucleosomes
The increase in CDC28 expression upon stress results in an increase in de novo synthesis of Cdc28. As described above (Sect. Control of cell cycle progression by the Hog1 SAPK), Hog1 mediates a rapid, but transient, arrest of cell-cycle progression to allow adaptation. Mechanisms under the control of Hog1 include the down-regulation of cyclins and the direct inhibitory phosphorylation of Swe1 to reduce Cdc28 activity. These down-regulatory effects of Hog1 on Cdc28 appear to be in contradiction to an increase in Cdc28 protein. However, the actual increase in Cdc28 levels does not occur until about 30 min after the cells are subjected to 0.4 M NaCl. This increase in Cdc28 protein coincides with the restart of the cell cycle after stress-induced arrest, which suggests a role for the increase in Cdc28 levels during the recovery phase. Of note, although cells deficient in CDC28 lncRNA underwent arrest similar to wild-type cells upon stress, they were less efficient in re-entering the cell cycle, suggesting that the increase in Cdc28 protein permits a faster recovery from the cell cycle delay that is caused by stress. Therefore, Hog1 is able both to induce a cell-cycle delay and to promote cell recovery by controlling Cdc28 by different mechanisms with a different temporal outcome.
Control of the cell cycle by lncRNAs
Cell-cycle oscillation of coding and lncRNAs
The establishment of a critical role for a lncRNA in CDC28 and cell-cycle control has prompted us to analyze other lncRNAs. Deep transcriptome analyses have highlighted the complexity of gene expression networks. Even in scenarios such as the cell cycle that had been previously studied in depth, tiling array analyses have extended the number of periodically expressed protein coding genes and have provided evidence of an additional layer of regulation based on lncRNAs (Granovskaia et al. 2010).
Depending on whether the sense or the antisense RNA is induced, there are four possible scenarios that can describe their transcriptional relationship (Fig. 4). The first scenario, which is the most common one for cell-cycle regulated pairs, is where sense RNA is induced but lncRNA remains constant. In the second scenario, sense RNA remains constant but lncRNA expression is induced. This scenario is the most common one for osmoresponsive lncRNAs. The third category is exemplified by CDC28 that displays the most interesting pattern in which both sense and antisense RNA expression increase with the same kinetics and times. The final scenario is where sense and antisense RNA expression increase in an oscillatory anti-correlation pattern.
Types of dynamics of sense-antisense RNA pairs. Schematic representation of expression profiles of sense-antisense RNAs during cell-cycle progression (left panels) and under osmostress conditions (right panels). Numbers in the upper corner of each graph indicate the number of cases described in the literature for the mitotic cell cycle (Granovskaia et al. 2010) and for stress conditions (Nadal-Ribelles et al. 2014, unpublished data)
Regulation of some key cell-cycle regulatory proteins falls into this last behavioral category and probably the most striking case is that of the CDK inhibitor FAR1. Expression of FAR1 lncRNA peaks during the G1/S transition, when Far1 levels should be very low to ensure entry into replication. Of note, in response to osmostress the HOG pathway controls the expression of FAR1 lncRNA that anti-correlates with the sense transcript (Nadal-Ribelles et al. 2014). The mechanism of action of this sense-antisense RNA pair remains unknown although the potential of regulating CDK inhibitors by a number of non-post-translational mechanisms might offer a higher degree of flexibility or contribute to the robustness of the response. Interestingly, in response to stress and cell-cycle progression, there is an additional class of regulated intergenic non-coding RNAs, although the physiological role, if any, of these RNAs remain obscure (Granovskaia et al. 2010; Nadal-Ribelles et al. 2014).
The presence of lncRNAs in tightly coordinated and complex networks such as the mitotic cell cycle and the response to osmostress suggests a regulatory action of antisense transcripts. Examples such as the ones described above illustrate the potential extent of lncRNA function.
Different examples of lncRNAs that target cell-cycle genes (cyclins, CDKs and CDK inhibitors)
Phenotypic variation cannot be satisfactorily explained by protein coding genes alone. Interestingly, analysis of fully sequenced genomes has shown that the number of non-protein coding sequences consistently increases with organism complexity (Taft et al. 2007). Robustness of the cell-cycle network is essential for the survival of all organisms; indeed, lncRNAs that affect the cell-cycle have been identified in a variety of organisms from prokaryotes to higher eukaryotes.
For example, replication initiation in bacteria is conducted by the replication initiation factor (DnaA) that recognizes replication origin (ori) and triggers the assembly of the replisome. The mechanism of replication (DnaA and its binding sites) is conserved throughout the prokaryotic kingdom. Recently, the existence of an RNA that was encoded in cis to the DnaA gene was shown to regulate the stability of the sense mRNA in response to environmental insults including osmostress in Salmonella typhi. Expression of the antisense RNA increases upon osmotic shock and causes a concomitant increase in DnaA mRNA stability, thereby enhancing total protein DnaA levels that ultimately help cells to recover from such stresses (Dadzie et al. 2013). This scenario resembles the CDC28 case in yeast.
In higher eukaryotes lncRNAs also target master regulators of the cell cycle by acting through several mechanisms that operate as regulators of a variety of features such as epigenetic state, transcription factors and post-transcriptional events reviewed in Kitagawa et al. (2013). An interesting example is the antisense RNA ANRIL that is induced by the DNA-damage signaling pathway (ATM-E2F1) (Wan et al. 2013). Expression of ANRIL causes an epigenetic repression of the INK locus where the p15 and p16 CDK inhibitors are located and ANRIL is critical for cell-cycle regulation and tumor progression (Kotake et al. 2011; Yap et al. 2010).
Perspective
The presence of lncRNA transcription provides an exciting landscape of new possibilities in the field of transcription in eukaryotes and challenges the classical view of gene unit. The mechanisms of lncRNA regulation of the transcription of protein coding genes are still far from being understood, mainly because detailed biochemical studies are scarce. The furthering of our knowledge in the direction of understanding of these mechanisms will allow assessment of whether specialized lncRNA transcription machinery exists (or of the requirements for lncRNA transcription). Knowledge of the transcriptional regulation, the structure and the stability of lncRNAs may indicate a new level of transcriptional or post-transcriptional control for the modulation of cell physiology. The examples described above illustrate the potential extent of the regulatory actions of lncRNA transcripts, which range from their role in cellular adaptation to stress to their role in the control of cell-cycle progression. Analysis at the single cell level will lead to an understanding of the complex dynamic interplay between sense and antisense RNA that may be masked by population-based approaches. Determination of whether sense-antisense RNA pairs coexist or are mutually exclusive will help to dissect the impact of lncRNA transcription on the transcription of protein coding genes. Major physiological roles have already been assigned to lncRNAs (Kitagawa et al. 2013; Ng et al. 2013; Bergmann and Spector 2014; Rossi and Antonangeli 2014) and the enormous variety of mechanisms by which they modulate those phenotypes are only beginning to be understood (Pelechano and Steinmetz 2013; Vance and Ponting 2014).
References
Adrover MA, Zi Z, Duch A, Schaber J, Gonzalez-Novo A, Jimenez J, Nadal-Ribelles M, Clotet J, Klipp E, Posas F (2011) Time-dependent quantitative multicomponent control of the G-S network by the stress-activated protein kinase Hog1 upon osmostress. Sci Signal, 4, ra63
Alepuz PM, Jovanovic A, Reiser V, Ammerer G (2001) Stress-induced map kinase Hog1 is part of transcription activation complexes. Mol Cell 7:767–777
Alepuz PM, de Nadal E, Zapater M, Ammerer G, Posas F (2003) Osmostress-induced transcription by Hot1 depends on a Hog1-mediated recruitment of the RNA Pol II. EMBO J 22:2433–2442
Alexander MR, Tyers M, Perret M, Craig BM, Fang KS, Gustin MC (2001) Regulation of cell cycle progression by Swe1p and Hog1p following hypertonic stress. Mol Biol Cell 12:53–62
Bergmann JH, Spector DL (2014) Long non-coding RNAs: modulators of nuclear structure and function. Curr Opin Cell Biol 26:10–18
Bertone P, Stolc V, Royce TE, Rozowsky JS, Urban AE, Zhu X, Rinn JL, Tongprasit W, Samanta M, Weissman S, Gerstein M, Snyder M (2004) Global identification of human transcribed sequences with genome tiling arrays. Science 306:2242–2246
Bloom J, Cross FR (2007) Multiple levels of cyclin specificity in cell-cycle control. Nat Rev Mol Cell Biol 8:149–160
Capaldi AP, Kaplan T, Liu Y, Habib N, Regev A, Friedman N, O’Shea EK (2008) Structure and function of a transcriptional network activated by the MAPK Hog1. Nat Genet 40:1300–1306
Causton HC, Ren B, Koh SS, Harbison CT, Kanin E, Jennings EG, Lee TI, True HL, Lander ES, Young RA (2001) Remodeling of yeast genome expression in response to environmental changes. Mol Biol Cell 12:323–337
Cho RJ, Campbell MJ, Winzeler EA, Steinmetz L, Conway A, Wodicka L, Wolfsberg TG, Gabrielian AE, Landsman D, Lockhart DJ, Davis RW (1998) A genome-wide transcriptional analysis of the mitotic cell cycle. Mol Cell 2:65–73
Churchman LS, Weissman JS (2011) Nascent transcript sequencing visualizes transcription at nucleotide resolution. Nature 469:368–373
Clotet J, Escote X, Adrover MA, Yaakov G, Gari E, Aldea M, de Nadal E, Posas F (2006) Phosphorylation of Hsl1 by Hog1 leads to a G2 arrest essential for cell survival at high osmolarity. EMBO J 25:2338–2346
Cook KE, O’Shea EK (2012) Hog1 controls global reallocation of RNA Pol II upon osmotic shock in Saccharomyces cerevisiae. G3 (Bethesda) 2:1129–1136
Dadzie I, Xu S, Ni B, Zhang X, Zhang H, Sheng X, Xu H, Huang X (2013) Identification and characterization of a cis-encoded antisense RNA associated with the replication process of Salmonella enterica serovar Typhi. PLoS One 8:e61308
David L, Huber W, Granovskaia M, Toedling J, Palm CJ, Bofkin L, Jones T, Davis RW, Steinmetz LM (2006) A high-resolution map of transcription in the yeast genome. Proc Natl Acad Sci USA 103:5320–5325
de Nadal E, Casadome L, Posas F (2003) Targeting the MEF2-like transcription factor Smp1 by the stress-activated Hog1 mitogen-activated protein kinase. Mol Cell Biol 23:229–237
de Nadal E, Zapater M, Alepuz PM, Sumoy L, Mas G, Posas F (2004) The MAPK Hog1 recruits Rpd3 histone deacetylase to activate osmoresponsive genes. Nature 427:370–374
de Nadal E, Ammerer G, Posas F (2011) Controlling gene expression in response to stress. Nat Rev Genet 12:833–845
Duch A, de Nadal E, Posas F (2012) The p38 and Hog1 SAPKs control cell cycle progression in response to environmental stresses. FEBS Lett 586:2925–2931
Duch A, Felipe-Abrio I, Barroso S, Yaakov G, Garcia-Rubio M, Aguilera A, de Nadal E, Posas F (2013) Coordinated control of replication and transcription by a SAPK protects genomic integrity. Nature 493:116–119
Escote X, Zapater M, Clotet J, Posas F (2004) Hog1 mediates cell-cycle arrest in G1 phase by the dual targeting of Sic1. Nat Cell Biol 6:997–1002
Garcia-Martinez J, Aranda A, Perez-Ortin JE (2004) Genomic run-on evaluates transcription rates for all yeast genes and identifies gene regulatory mechanisms. Mol Cell 15:303–313
Gasch AP, Spellman PT, Kao CM, Carmel-Harel O, Eisen MB, Storz G, Botstein D, Brown PO (2000) Genomic expression programs in the response of yeast cells to environmental changes. Mol Biol Cell 11:4241–4257
Gauthier NP, Larsen ME, Wernersson R, de Lichtenberg U, Jensen LJ, Brunak S, Jensen TS (2008) Cyclebase.org—a comprehensive multi-organism online database of cell-cycle experiments. Nucleic Acids Res 36:D854–D859
Granovskaia MV, Jensen LJ, Ritchie ME, Toedling J, Ning Y, Bork P, Huber W, Steinmetz LM (2010) High-resolution transcription atlas of the mitotic cell cycle in budding yeast. Genom Biol 11:R24
Guha N, Desai P, Vancura A (2007) Plc1p is required for SAGA recruitment and derepression of Sko1p-regulated genes. Mol Biol Cell 18:2419–2428
Hartwell LH, Mortimer RK, Culotti J, Culotti M (1973) Genetic control of the cell division cycle in yeast: V. Genet Anal cdc Mut. Genet 74:267–286
Kim T, Xu Z, Clauder-Munster S, Steinmetz LM, Buratowski S (2012) Set3 HDAC mediates effects of overlapping noncoding transcription on gene induction kinetics. Cell 150:1158–1169
Kitagawa M, Kitagawa K, Kotake Y, Niida H, Ohhata T (2013) Cell cycle regulation by long non-coding RNAs. Cell Mol Life Sci 70:4785–4794
Klopf E, Paskova L, Sole C, Mas G, Petryshyn A, Posas F, Wintersberger U, Ammerer G, Schuller C (2009) Cooperation between the INO80 complex and histone chaperones determines adaptation of stress gene transcription in the yeast Saccharomyces cerevisiae. Mol Cell Biol 29:4994–5007
Kobayashi Y, Inai T, Mizunuma M, Okada I, Shitamukai A, Hirata D, Miyakawa T (2008) Identification of Tup1 and Cyc8 mutations defective in the responses to osmotic stress. Biochem Biophys Res Commun 368:50–55
Kotake Y, Nakagawa T, Kitagawa K, Suzuki S, Liu N, Kitagawa M, Xiong Y (2011) Long non-coding RNA ANRIL is required for the PRC2 recruitment to and silencing of p15(INK4B) tumor suppressor gene. Oncogene 30:1956–1962
Lee TI, Rinaldi NJ, Robert F, Odom DT, Bar-Joseph Z, Gerber GK, Hannett NM, Harbison CT, Thompson CM, Simon I, Zeitlinger J, Jennings EG, Murray HL, Gordon DB, Ren B, Wyrick JJ, Tagne JB, Volkert TL, Fraenkel E, Gifford DK, Young RA (2002) Transcriptional regulatory networks in Saccharomyces cerevisiae. Science 298:799–804
Mas G, de Nadal E, Dechant R, de la Rodriguez Concepcion ML, Logie C, Jimeno-Gonzalez S, Chavez S, Ammerer G, Posas F (2009) Recruitment of a chromatin remodelling complex by the Hog1 MAP kinase to stress genes. EMBO J 28:326–336
Mendenhall MD, Hodge AE (1998) Regulation of Cdc28 cyclin-dependent protein kinase activity during the cell cycle of the yeast Saccharomyces cerevisiae. Microbiol Mol Biol Rev 62:1191–1243
Miller C, Schwalb B, Maier K, Schulz D, Dumcke S, Zacher B, Mayer A, Sydow J, Marcinowski L, Dolken L, Martin DE, Tresch A, Cramer P (2011) Dynamic transcriptome analysis measures rates of mRNA synthesis and decay in yeast. Mol Syst Biol 7:458
Moazed D (2009) Small RNAs in transcriptional gene silencing and genome defence. Nature 457:413–420
Molin C, Jauhiainen A, Warringer J, Nerman O, Sunnerhagen P (2009) mRNA stability changes precede changes in steady-state mRNA amounts during hyperosmotic stress. RNA 15:600–614
Nadal-Ribelles M, Conde N, Flores O, Gonzalez-Vallinas J, Eyras E, Orozco M, de Nadal E, Posas F (2012) Hog1 bypasses stress-mediated down-regulation of transcription by RNA polymerase II redistribution and chromatin remodeling. Genom Biol 13:R106
Nadal-Ribelles M, Sole C, Xu Z, Steinmetz LM, de Nadal E, Posas F (2014) Control of Cdc28 CDK1 by a stress-induced lncRNA. Mol Cell 53:549–561
Nagalakshmi U, Wang Z, Waern K, Shou C, Raha D, Gerstein M, Snyder M (2008) The transcriptional landscape of the yeast genome defined by RNA sequencing. Science 320:1344–1349
Neuert G, Munsky B, Tan RZ, Teytelman L, Khammash M, van Oudenaarden A (2013) Systematic identification of signal-activated stochastic gene regulation. Science 339:584–587
Ng SY, Lin L, Soh BS, Stanton LW (2013) Long noncoding RNAs in development and disease of the central nervous system. Tren Genet 29:461–468
Ni L, Bruce C, Hart C, Leigh-Bell J, Gelperin D, Umansky L, Gerstein MB, Snyder M (2009) Dynamic and complex transcription factor binding during an inducible response in yeast. Genes Dev 23:1351–1363
Orlando DA, Lin CY, Bernard A, Wang JY, Socolar JE, Iversen ES, Hartemink AJ, Haase SB (2008) Global control of cell-cycle transcription by coupled CDK and network oscillators. Nature 453:944–947
O’Rourke SM, Herskowitz I (2004) Unique and redundant roles for HOG MAPK pathway components as revealed by whole-genome expression analysis. Mol Biol Cell 15:532–542
Pascual-Ahuir A, Struhl K, Proft M (2006) Genome-wide location analysis of the stress-activated MAP kinase Hog1 in yeast. Methods 40:272–278
Pelechano V, Steinmetz LM (2013) Gene regulation by antisense transcription. Nat Rev Genet 14:880–893
Pelechano V, Wei W, Steinmetz LM (2013) Extensive transcriptional heterogeneity revealed by isoform profiling. Nature 497:127–131
Pelet S, Rudolf F, Nadal-Ribelles M, de Nadal E, Posas F, Peter M (2011) Transient activation of the HOG MAPK pathway regulates bimodal gene expression. Science 332:732–735
Pokholok DK, Zeitlinger J, Hannett NM, Reynolds DB, Young RA (2006) Activated signal transduction kinases frequently occupy target genes. Science 313:533–536
Posas F, Chambers JR, Heyman JA, Hoeffler JP, de Nadal E, Arino J (2000) The transcriptional response of yeast to saline stress. J Biol Chem 275:17249–17255
Price C, Nasmyth K, Schuster T (1991) A general approach to the isolation of cell cycle-regulated genes in the budding yeast, Saccharomyces cerevisiae. J Mol Biol 218:543–556
Proft M, Struhl K (2002) Hog1 kinase converts the Sko1-Cyc8-Tup1 repressor complex into an activator that recruits SAGA and SWI/SNF in response to osmotic stress. Mol Cell 9:1307–1317
Proft M, Struhl K (2004) MAP kinase-mediated stress relief that precedes and regulates the timing of transcriptional induction. Cell 118:351–361
Proft M, Mas G, de Nadal E, Vendrell A, Noriega N, Struhl K, Posas F (2006) The stress-activated Hog1 kinase is a selective transcriptional elongation factor for genes responding to osmotic stress. Mol Cell 23:241–250
Quintales L, Sanchez M, Antequera F (2010) Analysis of DNA strand-specific differential expression with high density tiling microarrays. BMC Bioinform 11:136
Regot S, de Nadal E, Rodriguez-Navarro S, Gonzalez-Novo A, Perez-Fernandez J, Gadal O, Seisenbacher G, Ammerer G, Posas F (2013) The Hog1 Stress-activated protein kinase targets nucleoporins to Control mRNA export upon stress. J Biol Chem 288:17384–17398
Rep M, Krantz M, Thevelein JM, Hohmann S (2000) The transcriptional response of Saccharomyces cerevisiae to osmotic shock. Hot1p and Msn2p/Msn4p are required for the induction of subsets of high osmolarity glycerol pathway-dependent genes. J Biol Chem 275:8290–8300
Romero-Santacreu L, Moreno J, Perez-Ortin JE, Alepuz P (2009) Specific and global regulation of mRNA stability during osmotic stress in Saccharomyces cerevisiae. RNA 15:1110–1120
Rossi MN, Antonangeli F (2014) LncRNAs: new players in apoptosis control. Int J Cell Biol 2014:473857
Ruiz-Roig C, Noriega N, Duch A, Posas F, de Nadal E (2012) The Hog1 SAPK controls the Rtg1/Rtg3 transcriptional complex activity by multiple regulatory mechanisms. Mol Biol Cell 23:4286–4296
Saito H, Posas F (2012) Response to hyperosmotic stress. Genetics 192:289–318
Simon I, Barnett J, Hannett N, Harbison CT, Rinaldi NJ, Volkert TL, Wyrick JJ, Zeitlinger J, Gifford DK, Jaakkola TS, Young RA (2001) Serial regulation of transcriptional regulators in the yeast cell cycle. Cell 106:697–708
Sole C, Nadal-Ribelles M, Kraft C, Peter M, Posas F, de Nadal E (2011) Control of Ubp3 ubiquitin protease activity by the Hog1 SAPK modulates transcription upon osmostress. EMBO J 30:3274–3284
Spellman PT, Sherlock G, Zhang MQ, Iyer VR, Anders K, Eisen MB, Brown PO, Botstein D, Futcher B (1998) Comprehensive identification of cell cycle-regulated genes of the yeast Saccharomyces cerevisiae by microarray hybridization. Mol Biol Cell 9:3273–3297
Taft RJ, Pheasant M, Mattick JS (2007) The relationship between non-protein-coding DNA and eukaryotic complexity. BioEssays 29:288–299
Tan-Wong SM, Zaugg JB, Camblong J, Xu Z, Zhang DW, Mischo HE, Ansari AZ, Luscombe NM, Steinmetz LM, Proudfoot NJ (2012) Gene loops enhance transcriptional directionality. Science 338:671–675
van Dijk EL, Chen CL, d’Aubenton-Carafa Y, Gourvennec S, Kwapisz M, Roche V, Bertrand C, Silvain M, Legoix-Ne P, Loeillet S, Nicolas A, Thermes C, Morillon A (2011) XUTs are a class of Xrn1-sensitive antisense regulatory non-coding RNA in yeast. Nature 475:114–117
Vance KW, Ponting CP (2014) Transcriptional regulatory functions of nuclear long noncoding RNAs. Trends Genet 30:348–355
Vendrell A, Martinez-Pastor M, Gonzalez-Novo A, Pascual-Ahuir A, Sinclair DA, Proft M, Posas F (2011) Sir2 histone deacetylase prevents programmed cell death caused by sustained activation of the Hog1 stress-activated protein kinase. EMBO Rep 12:1062–1068
Wan G, Mathur R, Hu X, Liu Y, Zhang X, Peng G, Lu X (2013) Long non-coding RNA ANRIL (CDKN2B-AS) is induced by the ATM-E2F1 signaling pathway. Cell Signal 25:1086–1095
Weake VM, Workman JL (2010) Inducible gene expression: diverse regulatory mechanisms. Nat Rev Genet 11:426–437
Xu Z, Wei W, Gagneur J, Perocchi F, Clauder-Munster S, Camblong J, Guffanti E, Stutz F, Huber W, Steinmetz LM (2009) Bidirectional promoters generate pervasive transcription in yeast. Nature 457:1033–1037
Yap KL, Li S, Munoz-Cabello AM, Raguz S, Zeng L, Mujtaba S, Gil J, Walsh MJ, Zhou MM (2010) Molecular interplay of the noncoding RNA ANRIL and methylated histone H3 lysine 27 by polycomb CBX7 in transcriptional silencing of INK4a. Mol Cell 38:662–674
Young RA (2000) Biomedical discovery with DNA arrays. Cell 102:9–15
Zapater M, Sohrmann M, Peter M, Posas F, de Nadal E (2007) Selective requirement for SAGA in Hog1-mediated gene expression depending on the severity of the external osmostress conditions. Mol Cell Biol 27:3900–3910
Zechner C, Ruess J, Krenn P, Pelet S, Peter M, Lygeros J, Koeppl H (2012) Moment-based inference predicts bimodality in transient gene expression. Proc Natl Acad Sci U S A 109:8340–8345
Acknowledgments
We apologize to colleagues in the field for not citing all relevant papers owing to space constraints. The laboratory of FP and EN is supported by grants from the Spanish Government (BFU2012-33503 and FEDER to FP, BFU2011-26722 to EN), an ERC Advanced Grant Number 294294 from the EU seventh framework program (SYNCOM) and the Fundación Marcelino Botín (FMB) to FP. FP and EN are recipients of an ICREA Acadèmia (Generalitat de Catalunya). The authors declare no competing financial interest. This review article was supported in part by a grant from the São Paulo Research Foundation (FAPESP) of Brazil # 2014/01229-4.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by D. E. N. Rangel.
Carme Solé and Mariona Nadal-Ribelles contributed equally to this review.
This article is part of the Special Issue “Fungal Stress Responses”.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Solé, C., Nadal-Ribelles, M., de Nadal, E. et al. A novel role for lncRNAs in cell cycle control during stress adaptation. Curr Genet 61, 299–308 (2015). https://doi.org/10.1007/s00294-014-0453-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-014-0453-y