Abstract
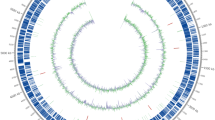
Genome sequencing was performed by the PacBio RS II platform and Illumina HiSeq 4000 platform to discover the metabolic profile of the Deinococcus wulumuqiensis R12, which was isolated from radiation-contaminated soils in Xinjiang Uygur Autonomous Region of northwest China. The genome of 3.5 Mbp comprises one circular chromosome and four circular plasmids with 3679 genes and a GC content of 66.97%. A total of 41 new transcriptional factors were identified using the DeepTFactor tool. Genomic analysis revealed the presence of genes for homologous recombination repair, which suggested high recombination efficiency in R12. Three Type I and one Type II RM systems, two CRISPR arrays, and one Cas-Type IC protein were found, allowing the development of endogenous CRISPR-Cas gene-editing tools. Additionally, we found that R12 has a broad spectrum of substrate utilization, which was validated by physiological experiments. Genes involved in the carotenoid biosynthesis pathway and the antioxidative system were also identified. Overall, the comprehensive description of the genome of R12 will facilitate the additional exploitation of this strain as a versatile cell factory for biotechnological applications.





Similar content being viewed by others

Availability of Data and Material
Not applicable.
Code Availability
Not applicable.
References
Cox MM, Battista JR (2005) Deinococcus radiodurans—the consummate survivor. Nat Rev Microbiol 3(11):882–892. https://doi.org/10.1038/nrmicro1264
Tian B, Hua YJ (2010) Carotenoid biosynthesis in extremophilic Deinococcus-Thermus bacteria. Trends Microbiol 18(11):512–520. https://doi.org/10.1016/j.tim.2010.07.007
Blasius M, Sommer S, Hubscher U (2008) Deinococcus radiodurans: what belongs to the survival kit? Crit Rev Biochem Mol Biol 43(3):221–238. https://doi.org/10.1080/10409230802122274
Makarova KS, Aravind L, Wolf YI, Tatusov RL, Minton KW, Koonin EV, Daly MJ (2001) Genome of the extremely radiation-resistant bacterium Deinococcus radiodurans viewed from the perspective of comparative genomics. Microbiol Mol Biol Rev 65(1):44–79. https://doi.org/10.1128/MMBR.65.1.44-79.2001
Slade D, Radman M (2011) Oxidative stress resistance in Deinococcus radiodurans. Microbiol Mol Biol Rev 75(1):133–191. https://doi.org/10.1128/MMBR.00015-10
Qi H-z, Wang W-z, He J-y, Ma Y, Xiao F-z, He S-y (2020) Antioxidative system of Deinococcus radiodurans. Res Microbiol 171(2):45–54. https://doi.org/10.1016/j.resmic.2019.11.002
Choi JY, Lee K, Lee PC (2019) Characterization of carotenoid biosynthesis in newly Isolated Deinococcus sp. AJ005 and investigation of the effects of environmental conditions on cell growth and carotenoid biosynthesis. Mar Drugs 17(12):10. https://doi.org/10.3390/md17120705
Jeong, SW, Kang, CK, Choi, YJ (2018) Metabolic engineering of Deinococcus radiodurans for the production of phytoene. J Microbiol Biotechnol. 28(10): 1691–1699. https://doi.org/10.4014/jmb.1808.08019
Johnston CD, Skeete CA, Fomenkov A, Roberts RJ, Rittling SR (2017) Restriction-modification mediated barriers to exogenous DNA uptake and incorporation employed by Prevotella intermedia. PLoS One 12(9):30. https://doi.org/10.1371/journal.pone.0185234
Zhao D, Zhu X, Zhou H, Sun N, Wang T, Bi C, Zhang X (2020) CRISPR-based metabolic pathway engineering. Metab Eng. https://doi.org/10.1016/j.ymben.2020.10.004
Wang W, Mao J, Zhang Z, Tang Q, Xie Y, Zhu J, Zhang L, Liu Z, Shi Y, Goodfellow M (2010) Deinococcus wulumuqiensis sp. nov., and Deinococcus xibeiensis sp. nov., isolated from radiation-polluted soil. Int J Syst Evol Microbiol 60 (9):2006–2010. https://doi.org/10.1099/ijs.0.015917-0
Xu X, Jiang L, Zhang Z, Shi Y, Huang H (2013) Genome sequence of a gamma- and UV-Ray-resistant strain, Deinococcus wulumuqiensis R12. Genome Announc 1(3):e00206-00213. https://doi.org/10.1128/genomeA.00206-13
Delcher AL, Bratke KA, Powers EC, Salzberg SL (2007) Identifying bacterial genes and endosymbiont DNA with Glimmer. Bioinformatics 23(6):673–679. https://doi.org/10.1093/bioinformatics/btm009
Chan PP, Lowe TM (2019) tRNAscan-SE: searching for tRNA genes in genomic sequences. Methods Mol Biol 1962:1–14. https://doi.org/10.1007/978-1-4939-9173-0_1
Lagesen K, Hallin P, Rødland EA, Staerfeldt H-H, Rognes T, Ussery DW (2007) RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucl Acids Res 35(9):3100–3108. https://doi.org/10.1093/nar/gkm160
Griffiths-Jones S, Bateman A, Marshall M, Khanna A, Eddy SR (2003) Rfam: an RNA family database. Nucl Acids Res 31(1):439–441. https://doi.org/10.1093/nar/gkg006
Benson G (1999) Tandem repeats finder: a program to analyze DNA sequences. Nucl Acids Res 27(2):573–580. https://doi.org/10.1093/nar/27.2.573
Hasan MS, Liu Q, Wang H, Fazekas J, Chen B, Che D (2012) GIST: Genomic island suite of tools for predicting genomic islands in genomic sequences. Bioinformation 8(4):203–205. https://doi.org/10.6026/97320630008203
Arndt D, Grant JR, Marcu A, Sajed T, Pon A, Liang Y, Wishart DS (2016) PHASTER: a better, faster version of the PHAST phage search tool. Nucl Acids Res 44(W1):W16–W21. https://doi.org/10.1093/nar/gkw387
Roberts RJ, Vincze T, Posfai J, Macelis D (2014) REBASE—a database for DNA restriction and modification: enzymes, genes and genomes. Nucl Acids Res 43(D1):D298–D299. https://doi.org/10.1093/nar/gku1046
Grissa I, Vergnaud G, Pourcel C (2007) CRISPRFinder: a web tool to identify clustered regularly interspaced short palindromic repeats. Nucl Acids Res 35 (suppl_2):W52–W57. https://doi.org/10.1093/nar/gkm360
Kim GB, Gao Y, Palsson BO, Lee SY (2021) DeepTFactor: A deep learning-based tool for the prediction of transcription factors. Proc Natl Acad Sci USA 118(2):e2021171118. https://doi.org/10.1073/pnas.2021171118
Aravind L, Anantharaman V, Balaji S, Babu MM, Iyer LM (2005) The many faces of the helix-turn-helix domain: transcription regulation and beyond. FEMS Microbiol Rev 29(2):231–262. https://doi.org/10.1016/j.femsre.2004.12.008
Clark DP, Pazdernik NJ (2013) Chapter e10—cell division and DNA replication. In: Clark DP, Pazdernik NJ (eds) Molecular biology (second edition). Academic Press, Boston, pp. e150-e158. https://doi.org/10.1016/B978-0-12-378594-7.00044-5
Tanaka M, Narumi I, Funayama T, Kikuchi M, Watanabe H, Matsunaga T, Nikaido O, Yamamoto K (2005) Characterization of pathways dependent on the uvsE, uvrA1, or uvrA2 gene product for UV resistance in Deinococcus radiodurans. J Bacteriol 187(11):3693–3697. https://doi.org/10.1128/jb.187.11.3693-3697.2005
Wigley DB (2013) Bacterial DNA repair: recent insights into the mechanism of RecBCD. AddAB and AdnAB Nat Rev Microbiol 11(1):9–13. https://doi.org/10.1038/nrmicro2917
Court DL, Sawitzke JA, Thomason LC (2002) Genetic engineering using homologous recombination. Annu Rev Genet 36:361–388. https://doi.org/10.1146/annurev.genet.36.061102.093104
Luan G, Cai Z, Gong F, Dong H, Lin Z, Zhang Y, Li Y (2013) Developing controllable hypermutable Clostridium cells through manipulating its methyl-directed mismatch repair system. Protein Cell 4(11):854–862. https://doi.org/10.1007/s13238-013-3079-9
Wu C, Zhang J, Du G, Chen J (2013) Heterologous expression of Lactobacillus casei RecO improved the multiple-stress tolerance and lactic acid production in Lactococcus lactis NZ9000 during salt stress. Bioresour Technol 143:238–241. https://doi.org/10.1016/j.biortech.2013.05.050
Pleška M, Lang M, Refardt D, Levin BR, Guet CC (2018) Phage–host population dynamics promotes prophage acquisition in bacteria with innate immunity. Nat Ecol Evolut 2(2):359–366. https://doi.org/10.1038/s41559-017-0424-z
Zhang J, Hong W, Guo L, Wang Y, Wang Y (2020) Enhancing plasmid transformation efficiency and enabling CRISPR-Cas9/Cpf1-based genome editing in Clostridium tyrobutyricum. Biotechnol Bioeng. https://doi.org/10.1002/bit.27435
Dupuis ME, Villion M, Magadán AH, Moineau S (2013) CRISPR-Cas and restriction-modification systems are compatible and increase phage resistance. Nat Commun 4(1):1–7. https://doi.org/10.1038/ncomms3087
Couvin D, Bernheim A, Toffano-Nioche C, Touchon M, Michalik J, Néron B, Rocha EPC, Vergnaud G, Gautheret D, Pourcel C (2018) CRISPRCasFinder, an update of CRISRFinder, includes a portable version, enhanced performance and integrates search for Cas proteins. Nucl Acids Res 46(W1):W246-w251. https://doi.org/10.1093/nar/gky425
Lombard V, Golaconda Ramulu H, Drula E, Coutinho PM, Henrissat B (2014) The carbohydrate-active enzymes database (CAZy) in 2013. Nucl Acids Res 42(D1):D490–D495. https://doi.org/10.1093/nar/gkt1178
Takaichi S (2011) Carotenoids in algae: distributions, biosyntheses and functions. Mar Drugs 9(6):1101–1118. https://doi.org/10.3390/md9061101
Ram S, Mitra M, Shah F, Tirkey SR, Mishra S (2020) Bacteria as an alternate biofactory for carotenoid production: a review of its applications, opportunities and challenges. J Funct Foods 67:13. https://doi.org/10.1016/j.jff.2020.103867
Daly MJ (2009) A new perspective on radiation resistance based on Deinococcus radiodurans. Nat Rev Microbiol 7(3):237–245. https://doi.org/10.1038/nrmicro2073
Lu J, Holmgren A (2014) The thioredoxin antioxidant system. Free Radical Biol Med 66:75–87. https://doi.org/10.1016/j.freeradbiomed.2013.07.036
Chandrangsu P, Loi VV, Antelmann H, Helmann JD (2018) The Role of bacillithiol in Gram-positive Firmicutes. Antioxid Redox Signal 28(6):445–462. https://doi.org/10.1089/ars.2017.7057
Acknowledgements
The authors thank the Beijing Genomics Institute (BGI, Shenzhen, China) for help with genome sequencing.
Funding
This work was supported by The National Natural Science Foundation of China (31922070, 22008114), and The Natural Science Foundation of Jiangsu Province (BK20180038, BK20200684), and The Natural Science Foundation of the Jiangsu Higher Education Institutions of China (20KJB530017).
Author information
Authors and Affiliations
Contributions
The study was designed by LJ and the paper was written by ZMZ, bioinformatics analysis was performed by ZJD, the experiments were performed by ZJD, and ZDZ, and manuscript was critically revised by LJ, ZMZ, LYZ, ZJD, and ZDZ. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declared that they have no conflict of interest to this work.
Ethical Approval
Not required.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Dai, Z., Zhang, Z., Zhu, L. et al. Complete Genome Sequencing Analysis of Deinococcus wulumuqiensis R12, an Extremely Radiation-Resistant Strain. Curr Microbiol 79, 292 (2022). https://doi.org/10.1007/s00284-022-02984-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-022-02984-5